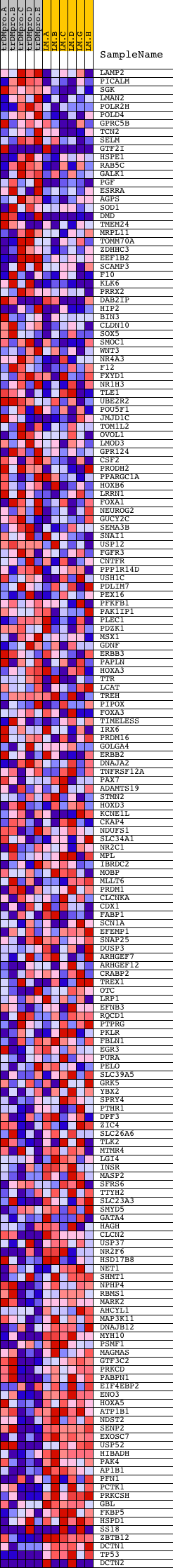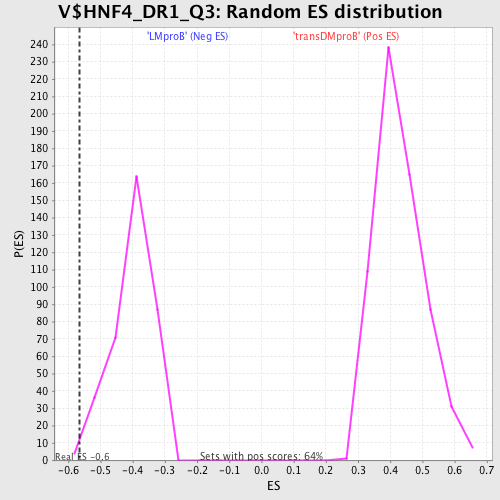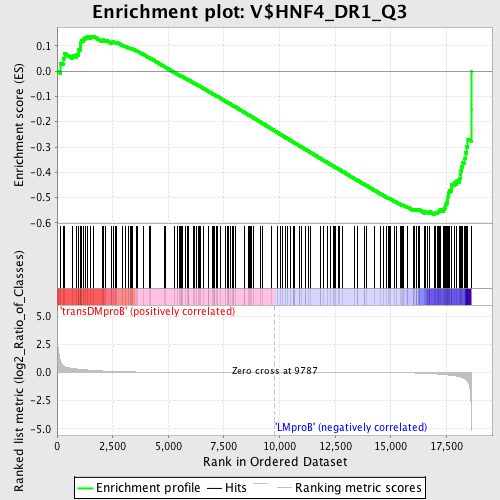
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_LMproB.phenotype_transDMproB_versus_LMproB.cls #transDMproB_versus_LMproB |
| Phenotype | phenotype_transDMproB_versus_LMproB.cls#transDMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | V$HNF4_DR1_Q3 |
| Enrichment Score (ES) | -0.56655896 |
| Normalized Enrichment Score (NES) | -1.4130038 |
| Nominal p-value | 0.005524862 |
| FDR q-value | 0.526531 |
| FWER p-Value | 0.999 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | LAMP2 | 9267 2653 | 162 | 0.982 | 0.0318 | No | ||
| 2 | PICALM | 18191 | 284 | 0.591 | 0.0496 | No | ||
| 3 | SGK | 9809 3419 | 321 | 0.544 | 0.0702 | No | ||
| 4 | LMAN2 | 21454 | 712 | 0.340 | 0.0630 | No | ||
| 5 | POLR2H | 10888 | 865 | 0.302 | 0.0673 | No | ||
| 6 | POLD4 | 12822 | 950 | 0.288 | 0.0746 | No | ||
| 7 | GPRC5B | 17662 | 979 | 0.282 | 0.0847 | No | ||
| 8 | TCN2 | 20550 | 1035 | 0.271 | 0.0930 | No | ||
| 9 | SELM | 20967 | 1053 | 0.268 | 0.1031 | No | ||
| 10 | GTF2I | 16353 5491 9863 3280 3184 | 1057 | 0.267 | 0.1140 | No | ||
| 11 | HSPE1 | 9133 | 1100 | 0.259 | 0.1224 | No | ||
| 12 | RAB5C | 20226 | 1200 | 0.240 | 0.1269 | No | ||
| 13 | GALK1 | 20147 | 1260 | 0.230 | 0.1332 | No | ||
| 14 | PGF | 21020 | 1354 | 0.217 | 0.1371 | No | ||
| 15 | ESRRA | 23802 | 1519 | 0.193 | 0.1362 | No | ||
| 16 | AGPS | 14979 | 1619 | 0.178 | 0.1382 | No | ||
| 17 | SOD1 | 9846 | 2048 | 0.128 | 0.1202 | No | ||
| 18 | DMD | 24295 2647 | 2070 | 0.125 | 0.1242 | No | ||
| 19 | TMEM24 | 19150 | 2185 | 0.115 | 0.1228 | No | ||
| 20 | MRPL11 | 23774 | 2464 | 0.090 | 0.1114 | No | ||
| 21 | TOMM70A | 6608 | 2466 | 0.090 | 0.1151 | No | ||
| 22 | ZDHHC3 | 7598 12763 | 2532 | 0.087 | 0.1152 | No | ||
| 23 | EEF1B2 | 4131 12063 | 2617 | 0.081 | 0.1140 | No | ||
| 24 | SCAMP3 | 15543 | 2662 | 0.077 | 0.1148 | No | ||
| 25 | F10 | 18682 | 2931 | 0.062 | 0.1028 | No | ||
| 26 | KLK6 | 9231 9626 | 3057 | 0.055 | 0.0983 | No | ||
| 27 | PRRX2 | 15047 | 3222 | 0.049 | 0.0914 | No | ||
| 28 | DAB2IP | 12808 2692 | 3276 | 0.047 | 0.0905 | No | ||
| 29 | HIP2 | 3626 3485 16838 | 3338 | 0.044 | 0.0890 | No | ||
| 30 | BIN3 | 21969 | 3402 | 0.042 | 0.0873 | No | ||
| 31 | CLDN10 | 21935 2785 2601 2720 7188 | 3560 | 0.038 | 0.0804 | No | ||
| 32 | SOX5 | 9850 1044 5480 16937 | 3612 | 0.036 | 0.0791 | No | ||
| 33 | SMOC1 | 21221 | 3883 | 0.030 | 0.0657 | No | ||
| 34 | WNT3 | 20635 | 4150 | 0.025 | 0.0523 | No | ||
| 35 | NR4A3 | 9473 16212 5183 | 4183 | 0.024 | 0.0515 | No | ||
| 36 | F12 | 3260 21453 | 4808 | 0.017 | 0.0184 | No | ||
| 37 | FXYD1 | 17874 85 | 4888 | 0.016 | 0.0147 | No | ||
| 38 | NR1H3 | 10258 2787 | 5284 | 0.013 | -0.0061 | No | ||
| 39 | TLE1 | 5765 10185 15859 | 5397 | 0.013 | -0.0117 | No | ||
| 40 | UBE2R2 | 16240 | 5486 | 0.012 | -0.0160 | No | ||
| 41 | POU5F1 | 1547 23255 | 5531 | 0.012 | -0.0179 | No | ||
| 42 | JMJD1C | 11531 19996 | 5574 | 0.012 | -0.0197 | No | ||
| 43 | TOM1L2 | 10115 | 5577 | 0.012 | -0.0193 | No | ||
| 44 | OVOL1 | 23983 | 5620 | 0.011 | -0.0211 | No | ||
| 45 | LMOD3 | 17059 | 5772 | 0.011 | -0.0289 | No | ||
| 46 | GPR124 | 18643 918 | 5877 | 0.010 | -0.0341 | No | ||
| 47 | CSF2 | 20454 | 5902 | 0.010 | -0.0350 | No | ||
| 48 | PRODH2 | 1854 18300 | 5903 | 0.010 | -0.0346 | No | ||
| 49 | PPARGC1A | 16533 | 6148 | 0.009 | -0.0474 | No | ||
| 50 | HOXB6 | 20688 | 6175 | 0.009 | -0.0485 | No | ||
| 51 | LRRN1 | 17343 | 6285 | 0.008 | -0.0540 | No | ||
| 52 | FOXA1 | 21060 | 6372 | 0.008 | -0.0584 | No | ||
| 53 | NEUROG2 | 15426 | 6392 | 0.008 | -0.0591 | No | ||
| 54 | GUCY2C | 1140 16954 | 6427 | 0.008 | -0.0606 | No | ||
| 55 | SEMA3B | 5422 | 6580 | 0.007 | -0.0685 | No | ||
| 56 | SNAI1 | 14726 | 6823 | 0.007 | -0.0814 | No | ||
| 57 | USP12 | 5825 5824 | 6824 | 0.007 | -0.0811 | No | ||
| 58 | FGFR3 | 8969 3566 | 7001 | 0.006 | -0.0904 | No | ||
| 59 | CNTFR | 2515 15906 | 7043 | 0.006 | -0.0924 | No | ||
| 60 | PPP1R14D | 14471 | 7091 | 0.006 | -0.0947 | No | ||
| 61 | USH1C | 1471 17821 | 7093 | 0.006 | -0.0945 | No | ||
| 62 | PDLIM7 | 7434 | 7185 | 0.006 | -0.0992 | No | ||
| 63 | PEX16 | 2681 | 7196 | 0.006 | -0.0995 | No | ||
| 64 | PFKFB1 | 9552 | 7348 | 0.005 | -0.1075 | No | ||
| 65 | PAK1IP1 | 21660 | 7570 | 0.005 | -0.1193 | No | ||
| 66 | PLEC1 | 2273 2198 2205 2178 2230 22247 2213 2231 2295 2172 5263 2266 | 7640 | 0.004 | -0.1229 | No | ||
| 67 | PDZK1 | 15490 | 7694 | 0.004 | -0.1256 | No | ||
| 68 | MSX1 | 16547 | 7716 | 0.004 | -0.1266 | No | ||
| 69 | GDNF | 22514 2275 | 7813 | 0.004 | -0.1316 | No | ||
| 70 | ERBB3 | 19593 | 7885 | 0.004 | -0.1353 | No | ||
| 71 | PAPLN | 9301 | 7915 | 0.004 | -0.1367 | No | ||
| 72 | HOXA3 | 9108 1075 | 7917 | 0.004 | -0.1366 | No | ||
| 73 | TTR | 23615 | 8026 | 0.003 | -0.1423 | No | ||
| 74 | LCAT | 18762 | 8410 | 0.003 | -0.1630 | No | ||
| 75 | TREH | 19474 | 8414 | 0.003 | -0.1631 | No | ||
| 76 | PIPOX | 20340 | 8618 | 0.002 | -0.1740 | No | ||
| 77 | FOXA3 | 17949 | 8664 | 0.002 | -0.1763 | No | ||
| 78 | TIMELESS | 10177 | 8688 | 0.002 | -0.1775 | No | ||
| 79 | IRX6 | 7225 | 8740 | 0.002 | -0.1802 | No | ||
| 80 | PRDM16 | 12893 | 8812 | 0.002 | -0.1840 | No | ||
| 81 | GOLGA4 | 12022 | 9134 | 0.001 | -0.2013 | No | ||
| 82 | ERBB2 | 8913 | 9244 | 0.001 | -0.2072 | No | ||
| 83 | DNAJA2 | 12133 18806 | 9629 | 0.000 | -0.2280 | No | ||
| 84 | TNFRSF12A | 23107 | 9919 | -0.000 | -0.2437 | No | ||
| 85 | PAX7 | 15697 | 10052 | -0.001 | -0.2508 | No | ||
| 86 | ADAMTS19 | 23544 | 10133 | -0.001 | -0.2551 | No | ||
| 87 | STMN2 | 15638 | 10262 | -0.001 | -0.2620 | No | ||
| 88 | HOXD3 | 9115 | 10345 | -0.001 | -0.2664 | No | ||
| 89 | KCNE1L | 24044 | 10504 | -0.001 | -0.2749 | No | ||
| 90 | CKAP4 | 19663 | 10629 | -0.002 | -0.2816 | No | ||
| 91 | NDUFS1 | 13940 | 10638 | -0.002 | -0.2819 | No | ||
| 92 | SLC34A1 | 9824 | 10653 | -0.002 | -0.2826 | No | ||
| 93 | NR2C1 | 10215 | 10666 | -0.002 | -0.2832 | No | ||
| 94 | MPL | 15780 | 10900 | -0.002 | -0.2958 | No | ||
| 95 | IBRDC2 | 5747 | 10914 | -0.002 | -0.2964 | No | ||
| 96 | MOBP | 5112 | 10986 | -0.002 | -0.3001 | No | ||
| 97 | MLLT6 | 20678 | 11145 | -0.003 | -0.3086 | No | ||
| 98 | PRDM1 | 19775 3337 | 11162 | -0.003 | -0.3094 | No | ||
| 99 | CLCNKA | 15690 | 11299 | -0.003 | -0.3166 | No | ||
| 100 | CDX1 | 23429 | 11368 | -0.003 | -0.3202 | No | ||
| 101 | FABP1 | 17420 | 11822 | -0.004 | -0.3446 | No | ||
| 102 | SCN1A | 9784 | 11860 | -0.004 | -0.3464 | No | ||
| 103 | EFEMP1 | 20937 | 11969 | -0.004 | -0.3521 | No | ||
| 104 | SNAP25 | 5466 | 12140 | -0.005 | -0.3611 | No | ||
| 105 | DUSP3 | 1358 13022 | 12288 | -0.005 | -0.3689 | No | ||
| 106 | ARHGEF7 | 3823 | 12410 | -0.005 | -0.3752 | No | ||
| 107 | ARHGEF12 | 19156 | 12482 | -0.006 | -0.3788 | No | ||
| 108 | CRABP2 | 15554 | 12493 | -0.006 | -0.3792 | No | ||
| 109 | TREX1 | 10219 3111 | 12666 | -0.006 | -0.3882 | No | ||
| 110 | OTC | 9518 | 12694 | -0.006 | -0.3894 | No | ||
| 111 | LRP1 | 9284 | 12835 | -0.007 | -0.3968 | No | ||
| 112 | EFNB3 | 20391 | 13361 | -0.008 | -0.4249 | No | ||
| 113 | RQCD1 | 12205 | 13504 | -0.009 | -0.4322 | No | ||
| 114 | PTPRG | 22097 3151 | 13509 | -0.009 | -0.4321 | No | ||
| 115 | PKLR | 1850 15545 | 13797 | -0.010 | -0.4472 | No | ||
| 116 | FBLN1 | 4716 2208 | 13816 | -0.010 | -0.4478 | No | ||
| 117 | EGR3 | 4656 | 13887 | -0.011 | -0.4511 | No | ||
| 118 | PURA | 9670 | 14263 | -0.013 | -0.4709 | No | ||
| 119 | PELO | 8360 | 14514 | -0.014 | -0.4839 | No | ||
| 120 | SLC39A5 | 3364 19596 | 14657 | -0.016 | -0.4909 | No | ||
| 121 | GRK5 | 9036 23814 | 14816 | -0.017 | -0.4988 | No | ||
| 122 | YBX2 | 20818 | 14912 | -0.018 | -0.5032 | No | ||
| 123 | SPRY4 | 23448 | 14922 | -0.018 | -0.5029 | No | ||
| 124 | PTHR1 | 5318 18985 | 14995 | -0.019 | -0.5060 | No | ||
| 125 | DPF3 | 7659 12853 | 15176 | -0.022 | -0.5149 | No | ||
| 126 | ZIC4 | 5992 | 15237 | -0.023 | -0.5172 | No | ||
| 127 | SLC26A6 | 19304 | 15453 | -0.027 | -0.5277 | No | ||
| 128 | TLK2 | 20628 | 15496 | -0.028 | -0.5289 | No | ||
| 129 | MTMR4 | 20712 | 15520 | -0.029 | -0.5289 | No | ||
| 130 | LGI4 | 18296 | 15577 | -0.030 | -0.5307 | No | ||
| 131 | INSR | 18950 | 15741 | -0.035 | -0.5381 | No | ||
| 132 | MASP2 | 2547 | 15762 | -0.036 | -0.5377 | No | ||
| 133 | SFRS6 | 14751 | 15999 | -0.046 | -0.5486 | No | ||
| 134 | TTYH2 | 8585 4382 20605 | 16017 | -0.046 | -0.5476 | No | ||
| 135 | SLC23A3 | 10367 | 16057 | -0.048 | -0.5477 | No | ||
| 136 | SMYD5 | 17386 | 16144 | -0.053 | -0.5502 | No | ||
| 137 | GATA4 | 21792 4755 | 16154 | -0.053 | -0.5485 | No | ||
| 138 | HAGH | 9020 | 16163 | -0.054 | -0.5467 | No | ||
| 139 | CLCN2 | 22637 | 16235 | -0.058 | -0.5482 | No | ||
| 140 | USP37 | 13923 | 16267 | -0.059 | -0.5474 | No | ||
| 141 | NR2F6 | 4651 8876 18848 8912 | 16528 | -0.077 | -0.5583 | No | ||
| 142 | HSD17B8 | 9061 | 16547 | -0.079 | -0.5560 | No | ||
| 143 | NET1 | 7100 | 16631 | -0.087 | -0.5569 | No | ||
| 144 | SHMT1 | 5431 | 16719 | -0.095 | -0.5577 | No | ||
| 145 | NPHP4 | 15978 | 16748 | -0.098 | -0.5552 | No | ||
| 146 | RBMS1 | 14580 | 16958 | -0.120 | -0.5616 | Yes | ||
| 147 | MARK2 | 8899 | 17003 | -0.126 | -0.5588 | Yes | ||
| 148 | AHCYL1 | 6037 | 17099 | -0.139 | -0.5582 | Yes | ||
| 149 | MAP3K11 | 11163 | 17143 | -0.146 | -0.5545 | Yes | ||
| 150 | DNAJB12 | 7146 | 17170 | -0.149 | -0.5498 | Yes | ||
| 151 | MYH10 | 8077 | 17236 | -0.160 | -0.5467 | Yes | ||
| 152 | PSMF1 | 6009 | 17379 | -0.183 | -0.5469 | Yes | ||
| 153 | MAGMAS | 1677 22678 | 17411 | -0.189 | -0.5407 | Yes | ||
| 154 | GTF3C2 | 7749 | 17468 | -0.199 | -0.5355 | Yes | ||
| 155 | PRKCD | 21897 | 17470 | -0.200 | -0.5274 | Yes | ||
| 156 | PABPN1 | 12018 | 17501 | -0.206 | -0.5205 | Yes | ||
| 157 | EIF4EBP2 | 4662 | 17561 | -0.218 | -0.5147 | Yes | ||
| 158 | ENO3 | 8905 | 17566 | -0.219 | -0.5058 | Yes | ||
| 159 | HOXA5 | 17149 | 17569 | -0.220 | -0.4968 | Yes | ||
| 160 | ATP1B1 | 4420 | 17583 | -0.223 | -0.4883 | Yes | ||
| 161 | NDST2 | 21906 | 17596 | -0.225 | -0.4797 | Yes | ||
| 162 | SENP2 | 7990 | 17626 | -0.231 | -0.4717 | Yes | ||
| 163 | EXOSC7 | 19256 | 17713 | -0.250 | -0.4661 | Yes | ||
| 164 | USP52 | 3397 19841 | 17721 | -0.251 | -0.4561 | Yes | ||
| 165 | HIBADH | 17144 | 17732 | -0.253 | -0.4462 | Yes | ||
| 166 | PAK4 | 17909 | 17866 | -0.291 | -0.4414 | Yes | ||
| 167 | AP1B1 | 8599 | 17970 | -0.329 | -0.4334 | Yes | ||
| 168 | PFN1 | 9555 | 18097 | -0.381 | -0.4245 | Yes | ||
| 169 | PCTK1 | 24369 | 18135 | -0.399 | -0.4100 | Yes | ||
| 170 | PRKCSH | 19535 | 18146 | -0.404 | -0.3939 | Yes | ||
| 171 | GBL | 23091 1612 | 18188 | -0.433 | -0.3782 | Yes | ||
| 172 | FKBP5 | 23051 | 18239 | -0.466 | -0.3616 | Yes | ||
| 173 | HSPD1 | 4078 9129 | 18315 | -0.533 | -0.3437 | Yes | ||
| 174 | SS18 | 6506 6505 2030 | 18356 | -0.579 | -0.3219 | Yes | ||
| 175 | ZBTB12 | 23268 | 18395 | -0.637 | -0.2977 | Yes | ||
| 176 | DCTN1 | 1100 17392 | 18464 | -0.790 | -0.2687 | Yes | ||
| 177 | TP53 | 20822 | 18610 | -3.019 | -0.1519 | Yes | ||
| 178 | DCTN2 | 7635 12812 | 18612 | -3.684 | 0.0002 | Yes |