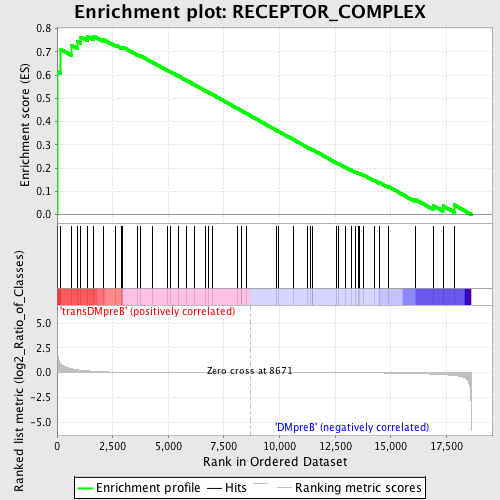
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | RECEPTOR_COMPLEX |
| Enrichment Score (ES) | 0.76784265 |
| Normalized Enrichment Score (NES) | 1.6199782 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.22532463 |
| FWER p-Value | 0.746 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MYH9 | 2252 2244 | 1 | 5.596 | 0.6149 | Yes | ||
| 2 | ITGB7 | 22112 | 146 | 0.951 | 0.7116 | Yes | ||
| 3 | CARD11 | 8436 | 636 | 0.392 | 0.7284 | Yes | ||
| 4 | SYK | 21636 | 921 | 0.288 | 0.7448 | Yes | ||
| 5 | ADRB2 | 23422 | 1067 | 0.244 | 0.7638 | Yes | ||
| 6 | SRPR | 12525 7433 | 1367 | 0.178 | 0.7672 | Yes | ||
| 7 | CD79A | 18342 | 1638 | 0.138 | 0.7678 | Yes | ||
| 8 | SMAD3 | 19084 | 2085 | 0.085 | 0.7531 | No | ||
| 9 | ITGA10 | 15493 | 2642 | 0.049 | 0.7286 | No | ||
| 10 | IL6R | 1862 4919 | 2912 | 0.040 | 0.7184 | No | ||
| 11 | ZAP70 | 14271 4042 | 2953 | 0.038 | 0.7205 | No | ||
| 12 | CHRNB4 | 19107 | 3631 | 0.023 | 0.6866 | No | ||
| 13 | ITGB3 | 20631 | 3749 | 0.021 | 0.6826 | No | ||
| 14 | CHRNE | 20368 | 4296 | 0.015 | 0.6548 | No | ||
| 15 | ITGA5 | 22105 | 4968 | 0.010 | 0.6198 | No | ||
| 16 | CHRNA3 | 4291 | 5075 | 0.010 | 0.6152 | No | ||
| 17 | ACVR1C | 14585 | 5434 | 0.008 | 0.5969 | No | ||
| 18 | ITGAX | 18058 | 5823 | 0.007 | 0.5767 | No | ||
| 19 | CHRND | 3931 4084 | 6192 | 0.006 | 0.5575 | No | ||
| 20 | CHRNA6 | 18900 | 6660 | 0.004 | 0.5329 | No | ||
| 21 | CD247 | 4498 8715 | 6805 | 0.004 | 0.5256 | No | ||
| 22 | ITGB6 | 14581 2809 | 6996 | 0.004 | 0.5157 | No | ||
| 23 | IL13RA1 | 24361 | 8122 | 0.001 | 0.4553 | No | ||
| 24 | ITGA11 | 6677 11449 | 8300 | 0.001 | 0.4458 | No | ||
| 25 | ADRB3 | 18901 | 8301 | 0.001 | 0.4459 | No | ||
| 26 | SDCBP | 2339 6998 6999 | 8509 | 0.000 | 0.4348 | No | ||
| 27 | CHRNA1 | 4319 14559 8530 | 9860 | -0.003 | 0.3624 | No | ||
| 28 | BMPR1B | 8661 15152 4451 | 9933 | -0.003 | 0.3588 | No | ||
| 29 | TGFBR1 | 16213 10165 5745 | 10618 | -0.004 | 0.3225 | No | ||
| 30 | GRIN1 | 9041 4804 | 11249 | -0.006 | 0.2892 | No | ||
| 31 | CHRNB3 | 18642 44 3879 | 11368 | -0.006 | 0.2835 | No | ||
| 32 | ITGAE | 20791 1481 1397 1380 1278 1238 1359 | 11457 | -0.007 | 0.2795 | No | ||
| 33 | ITGA3 | 20284 | 11500 | -0.007 | 0.2780 | No | ||
| 34 | ITGAM | 9188 | 12542 | -0.011 | 0.2232 | No | ||
| 35 | CHRNB2 | 8532 4321 | 12648 | -0.012 | 0.2188 | No | ||
| 36 | ITGA8 | 14685 | 12976 | -0.014 | 0.2027 | No | ||
| 37 | CHRNA2 | 21978 | 13214 | -0.015 | 0.1916 | No | ||
| 38 | TRIP6 | 16331 | 13414 | -0.017 | 0.1827 | No | ||
| 39 | OSMR | 22333 | 13561 | -0.018 | 0.1768 | No | ||
| 40 | CHRNA4 | 8531 4320 | 13570 | -0.018 | 0.1784 | No | ||
| 41 | CHRNA7 | 17808 | 13792 | -0.020 | 0.1687 | No | ||
| 42 | TGFBR2 | 2994 3001 10167 | 14248 | -0.026 | 0.1471 | No | ||
| 43 | IL28RA | 10823 16038 | 14481 | -0.030 | 0.1378 | No | ||
| 44 | ITGB4 | 9669 159 | 14901 | -0.039 | 0.1195 | No | ||
| 45 | ITGA9 | 19280 | 16121 | -0.086 | 0.0634 | No | ||
| 46 | ACVR1 | 4334 | 16905 | -0.154 | 0.0382 | No | ||
| 47 | BCL10 | 15397 | 17346 | -0.203 | 0.0368 | No | ||
| 48 | SRP9 | 11297 14019 | 17849 | -0.287 | 0.0413 | No |

