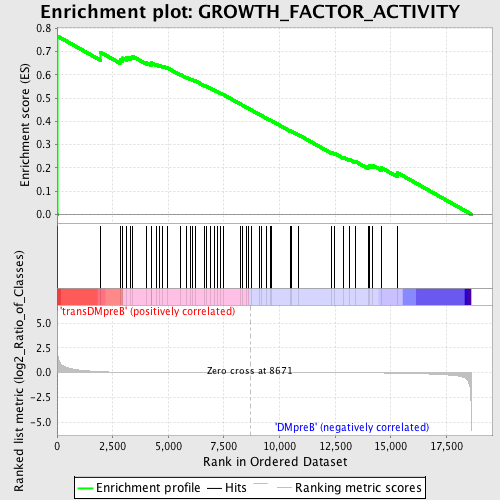
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | transDMpreB |
| GeneSet | GROWTH_FACTOR_ACTIVITY |
| Enrichment Score (ES) | 0.7666241 |
| Normalized Enrichment Score (NES) | 1.6124051 |
| Nominal p-value | 0.009881423 |
| FDR q-value | 0.14832547 |
| FWER p-Value | 0.814 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FGF13 | 4720 8963 2627 | 16 | 2.188 | 0.7666 | Yes | ||
| 2 | EGF | 15169 | 1943 | 0.098 | 0.6974 | No | ||
| 3 | JAG2 | 20975 | 2831 | 0.042 | 0.6645 | No | ||
| 4 | FGF3 | 17997 | 2935 | 0.039 | 0.6726 | No | ||
| 5 | FGF17 | 21757 | 3120 | 0.034 | 0.6745 | No | ||
| 6 | NUDT6 | 15352 | 3301 | 0.029 | 0.6749 | No | ||
| 7 | CLCF1 | 12160 3742 | 3402 | 0.027 | 0.6789 | No | ||
| 8 | IL4 | 9174 | 4010 | 0.018 | 0.6525 | No | ||
| 9 | FGF18 | 4721 1351 | 4227 | 0.016 | 0.6463 | No | ||
| 10 | IL3 | 20453 | 4232 | 0.016 | 0.6515 | No | ||
| 11 | CSF3 | 1394 20671 | 4462 | 0.014 | 0.6440 | No | ||
| 12 | FGF10 | 4719 8962 | 4612 | 0.013 | 0.6404 | No | ||
| 13 | BTC | 16483 | 4755 | 0.012 | 0.6369 | No | ||
| 14 | GRN | 20640 | 4941 | 0.011 | 0.6307 | No | ||
| 15 | FGF14 | 21716 3005 3463 | 5530 | 0.008 | 0.6018 | No | ||
| 16 | IL1F5 | 12037 | 5817 | 0.007 | 0.5888 | No | ||
| 17 | IL5 | 20884 10220 | 6004 | 0.006 | 0.5810 | No | ||
| 18 | AREG | 16796 | 6092 | 0.006 | 0.5784 | No | ||
| 19 | NTF5 | 18248 | 6210 | 0.006 | 0.5741 | No | ||
| 20 | DKK1 | 23700 | 6625 | 0.005 | 0.5534 | No | ||
| 21 | IL21 | 12241 | 6694 | 0.004 | 0.5512 | No | ||
| 22 | FGF4 | 8964 | 6886 | 0.004 | 0.5423 | No | ||
| 23 | PGF | 21020 | 7089 | 0.003 | 0.5326 | No | ||
| 24 | PDGFC | 7048 | 7199 | 0.003 | 0.5278 | No | ||
| 25 | IL12B | 20918 | 7351 | 0.003 | 0.5206 | No | ||
| 26 | IL7 | 4921 | 7473 | 0.002 | 0.5149 | No | ||
| 27 | FGF11 | 20383 | 8220 | 0.001 | 0.4751 | No | ||
| 28 | CSF2 | 20454 | 8345 | 0.001 | 0.4686 | No | ||
| 29 | SDCBP | 2339 6998 6999 | 8509 | 0.000 | 0.4600 | No | ||
| 30 | FGF7 | 8965 | 8584 | 0.000 | 0.4561 | No | ||
| 31 | FGF12 | 1723 22621 | 8735 | -0.000 | 0.4480 | No | ||
| 32 | EPGN | 16798 | 9112 | -0.001 | 0.4281 | No | ||
| 33 | FGF16 | 24270 | 9185 | -0.001 | 0.4246 | No | ||
| 34 | IL1RN | 15096 | 9398 | -0.002 | 0.4137 | No | ||
| 35 | INHA | 14214 | 9586 | -0.002 | 0.4044 | No | ||
| 36 | THPO | 22636 | 9640 | -0.002 | 0.4023 | No | ||
| 37 | BDNF | 14926 2797 | 10471 | -0.004 | 0.3590 | No | ||
| 38 | TGFA | 17381 | 10535 | -0.004 | 0.3571 | No | ||
| 39 | GDF5 | 4768 | 10839 | -0.005 | 0.3425 | No | ||
| 40 | TDGF1 | 5704 10108 | 12335 | -0.010 | 0.2655 | No | ||
| 41 | GDF10 | 4767 | 12461 | -0.011 | 0.2625 | No | ||
| 42 | IL2 | 15354 | 12851 | -0.013 | 0.2461 | No | ||
| 43 | CSF1 | 15202 | 13132 | -0.015 | 0.2361 | No | ||
| 44 | TRIP6 | 16331 | 13414 | -0.017 | 0.2268 | No | ||
| 45 | FGF1 | 1994 23447 | 13978 | -0.022 | 0.2043 | No | ||
| 46 | GDF9 | 20887 | 14041 | -0.023 | 0.2090 | No | ||
| 47 | FGF9 | 8966 | 14196 | -0.025 | 0.2095 | No | ||
| 48 | CSPG5 | 19296 | 14586 | -0.032 | 0.1997 | No | ||
| 49 | CLEC11A | 5411 | 15292 | -0.049 | 0.1790 | No |

