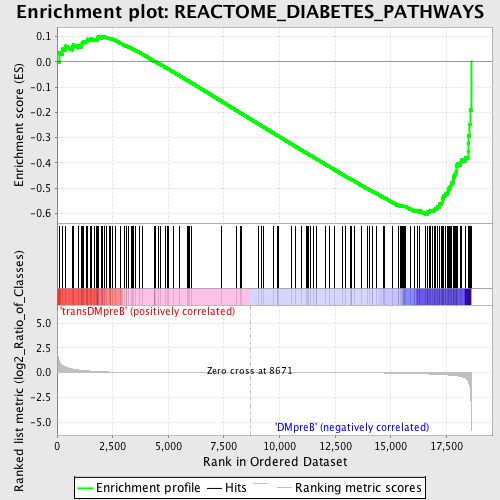
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB.phenotype_transDMpreB_versus_DMpreB.cls #transDMpreB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMpreB_versus_DMpreB.cls#transDMpreB_versus_DMpreB_repos |
| Upregulated in class | DMpreB |
| GeneSet | REACTOME_DIABETES_PATHWAYS |
| Enrichment Score (ES) | -0.60364497 |
| Normalized Enrichment Score (NES) | -1.5130213 |
| Nominal p-value | 0.00456621 |
| FDR q-value | 0.7665755 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PKM2 | 3642 9573 | 88 | 1.222 | 0.0358 | No | ||
| 2 | IQGAP1 | 6619 | 249 | 0.711 | 0.0507 | No | ||
| 3 | GNAS | 9025 2963 2752 | 376 | 0.559 | 0.0624 | No | ||
| 4 | RPL4 | 7499 19411 | 673 | 0.376 | 0.0589 | No | ||
| 5 | RPS15A | 6476 | 740 | 0.344 | 0.0667 | No | ||
| 6 | RPL38 | 12562 20606 7475 | 969 | 0.271 | 0.0633 | No | ||
| 7 | VAMP2 | 20826 | 1082 | 0.237 | 0.0651 | No | ||
| 8 | RPS28 | 12009 | 1116 | 0.228 | 0.0709 | No | ||
| 9 | OGDH | 9502 1412 | 1133 | 0.225 | 0.0775 | No | ||
| 10 | EXOC8 | 4161 | 1198 | 0.212 | 0.0811 | No | ||
| 11 | RPL39 | 12496 | 1302 | 0.190 | 0.0818 | No | ||
| 12 | RPL18A | 13358 | 1366 | 0.178 | 0.0843 | No | ||
| 13 | RPS25 | 13270 | 1370 | 0.178 | 0.0900 | No | ||
| 14 | HSPA5 | 9045 | 1512 | 0.157 | 0.0876 | No | ||
| 15 | RPS4X | 9756 | 1542 | 0.153 | 0.0911 | No | ||
| 16 | SRP72 | 16819 | 1681 | 0.132 | 0.0880 | No | ||
| 17 | RPL28 | 5392 | 1750 | 0.124 | 0.0884 | No | ||
| 18 | RPL5 | 9746 | 1811 | 0.116 | 0.0890 | No | ||
| 19 | RPS29 | 9754 | 1814 | 0.115 | 0.0927 | No | ||
| 20 | ATP5A1 | 23505 | 1815 | 0.115 | 0.0965 | No | ||
| 21 | NFYA | 5172 | 1855 | 0.110 | 0.0981 | No | ||
| 22 | COX8A | 8777 | 1862 | 0.108 | 0.1013 | No | ||
| 23 | NDUFA10 | 7420 | 2014 | 0.091 | 0.0962 | No | ||
| 24 | RPS3A | 9755 | 2021 | 0.090 | 0.0988 | No | ||
| 25 | RPL14 | 19267 | 2030 | 0.089 | 0.1014 | No | ||
| 26 | RPS21 | 12356 | 2137 | 0.080 | 0.0983 | No | ||
| 27 | RPLP1 | 3010 | 2198 | 0.076 | 0.0975 | No | ||
| 28 | RPL11 | 12450 | 2334 | 0.066 | 0.0924 | No | ||
| 29 | RPL8 | 22437 | 2387 | 0.063 | 0.0917 | No | ||
| 30 | MDH2 | 9399 | 2406 | 0.062 | 0.0927 | No | ||
| 31 | RPL37A | 9744 | 2504 | 0.056 | 0.0893 | No | ||
| 32 | RPL27 | 9740 | 2619 | 0.050 | 0.0848 | No | ||
| 33 | RPS5 | 18391 | 2849 | 0.042 | 0.0738 | No | ||
| 34 | PRKAR2B | 5288 2107 | 3033 | 0.036 | 0.0651 | No | ||
| 35 | RPS17 | 9753 | 3113 | 0.034 | 0.0619 | No | ||
| 36 | RPS13 | 12633 | 3119 | 0.034 | 0.0628 | No | ||
| 37 | RPS20 | 7438 | 3209 | 0.031 | 0.0590 | No | ||
| 38 | SLC2A2 | 15623 1880 1855 | 3357 | 0.028 | 0.0519 | No | ||
| 39 | ATP5B | 19846 | 3368 | 0.027 | 0.0523 | No | ||
| 40 | FFAR1 | 10583 | 3419 | 0.026 | 0.0505 | No | ||
| 41 | NDUFB10 | 23084 | 3532 | 0.024 | 0.0452 | No | ||
| 42 | UQCRB | 12545 | 3696 | 0.022 | 0.0371 | No | ||
| 43 | ATP5D | 19949 | 3826 | 0.020 | 0.0308 | No | ||
| 44 | IGFALS | 9158 | 4361 | 0.014 | 0.0023 | No | ||
| 45 | RPSA | 19270 4984 | 4369 | 0.014 | 0.0024 | No | ||
| 46 | UQCRFS1 | 21499 | 4386 | 0.014 | 0.0020 | No | ||
| 47 | IDH3G | 24132 | 4400 | 0.014 | 0.0018 | No | ||
| 48 | CTSL1 | 4574 | 4559 | 0.013 | -0.0063 | No | ||
| 49 | UQCRC1 | 10259 | 4648 | 0.012 | -0.0107 | No | ||
| 50 | RPL41 | 12611 | 4883 | 0.011 | -0.0230 | No | ||
| 51 | KLF4 | 9230 4962 4963 | 4951 | 0.011 | -0.0263 | No | ||
| 52 | RPL18 | 450 5390 | 5026 | 0.010 | -0.0300 | No | ||
| 53 | SDHD | 19125 | 5222 | 0.009 | -0.0402 | No | ||
| 54 | EIF2S1 | 4658 | 5504 | 0.008 | -0.0552 | No | ||
| 55 | RPS2 | 9279 | 5882 | 0.007 | -0.0754 | No | ||
| 56 | RPL37 | 12502 7421 22521 | 5891 | 0.007 | -0.0756 | No | ||
| 57 | MBTPS1 | 18730 3809 3770 | 5956 | 0.006 | -0.0788 | No | ||
| 58 | LEP | 17505 | 6042 | 0.006 | -0.0832 | No | ||
| 59 | KLK3 | 16337 | 7366 | 0.003 | -0.1548 | No | ||
| 60 | RPS16 | 9752 | 8046 | 0.001 | -0.1916 | No | ||
| 61 | PLA2G7 | 23233 | 8254 | 0.001 | -0.2028 | No | ||
| 62 | SDHA | 12436 | 8267 | 0.001 | -0.2034 | No | ||
| 63 | NKX2-2 | 5176 | 9059 | -0.001 | -0.2462 | No | ||
| 64 | SRP19 | 2004 12341 | 9190 | -0.001 | -0.2532 | No | ||
| 65 | RPL17 | 11429 6653 | 9297 | -0.001 | -0.2589 | No | ||
| 66 | PDX1 | 16621 | 9715 | -0.002 | -0.2814 | No | ||
| 67 | PFKFB1 | 9552 | 9907 | -0.003 | -0.2917 | No | ||
| 68 | CHRM3 | 21547 | 9911 | -0.003 | -0.2918 | No | ||
| 69 | KIF5B | 4955 2031 | 9960 | -0.003 | -0.2943 | No | ||
| 70 | SRP54 | 6282 | 10515 | -0.004 | -0.3241 | No | ||
| 71 | IGFBP1 | 20948 | 10732 | -0.005 | -0.3357 | No | ||
| 72 | NDUFS7 | 19946 | 10993 | -0.005 | -0.3496 | No | ||
| 73 | RAP1A | 8467 | 11212 | -0.006 | -0.3612 | No | ||
| 74 | IGFBP5 | 4899 | 11258 | -0.006 | -0.3634 | No | ||
| 75 | NDUFB4 | 22596 7541 | 11294 | -0.006 | -0.3651 | No | ||
| 76 | RPS3 | 6549 11295 | 11388 | -0.006 | -0.3700 | No | ||
| 77 | EXOC5 | 8370 | 11507 | -0.007 | -0.3761 | No | ||
| 78 | SYT5 | 17986 | 11663 | -0.007 | -0.3843 | No | ||
| 79 | GCG | 14578 | 11670 | -0.007 | -0.3844 | No | ||
| 80 | IGF1 | 3352 9156 3409 | 12045 | -0.009 | -0.4043 | No | ||
| 81 | IGFBP6 | 9161 | 12243 | -0.010 | -0.4147 | No | ||
| 82 | ADRA2A | 4355 | 12489 | -0.011 | -0.4276 | No | ||
| 83 | PGK1 | 9557 5244 | 12818 | -0.013 | -0.4450 | No | ||
| 84 | SDHB | 2348 12566 14880 | 12963 | -0.014 | -0.4523 | No | ||
| 85 | RPS23 | 12352 | 13195 | -0.015 | -0.4643 | No | ||
| 86 | PCSK2 | 9537 5231 14824 | 13229 | -0.015 | -0.4656 | No | ||
| 87 | MMP2 | 18525 | 13239 | -0.015 | -0.4656 | No | ||
| 88 | NDUFA6 | 12473 | 13358 | -0.016 | -0.4714 | No | ||
| 89 | RPLP2 | 7401 | 13683 | -0.019 | -0.4884 | No | ||
| 90 | NDUFC2 | 18183 | 13937 | -0.022 | -0.5013 | No | ||
| 91 | PRKCA | 20174 | 14043 | -0.023 | -0.5063 | No | ||
| 92 | GLP1R | 23304 | 14184 | -0.025 | -0.5130 | No | ||
| 93 | PFKFB2 | 4116 5242 | 14189 | -0.025 | -0.5124 | No | ||
| 94 | KCNJ11 | 17822 | 14369 | -0.028 | -0.5212 | No | ||
| 95 | ERN1 | 8137 | 14659 | -0.033 | -0.5357 | No | ||
| 96 | ATP5C1 | 8635 | 14712 | -0.034 | -0.5374 | No | ||
| 97 | CYCS | 8821 | 15080 | -0.043 | -0.5559 | No | ||
| 98 | NDUFAB1 | 7667 | 15325 | -0.050 | -0.5674 | No | ||
| 99 | CS | 19839 | 15329 | -0.050 | -0.5659 | No | ||
| 100 | COX7B | 7260 | 15412 | -0.053 | -0.5686 | No | ||
| 101 | NDUFB5 | 12277 | 15499 | -0.056 | -0.5714 | No | ||
| 102 | CPE | 18868 | 15507 | -0.057 | -0.5699 | No | ||
| 103 | ABCC8 | 9941 11649 | 15547 | -0.059 | -0.5701 | No | ||
| 104 | ETFB | 18279 | 15608 | -0.061 | -0.5713 | No | ||
| 105 | RPS14 | 9751 | 15643 | -0.063 | -0.5711 | No | ||
| 106 | RPL6 | 9747 | 15882 | -0.074 | -0.5815 | No | ||
| 107 | F2 | 14524 | 16051 | -0.082 | -0.5879 | No | ||
| 108 | ATP5I | 8637 | 16085 | -0.084 | -0.5869 | No | ||
| 109 | SSR4 | 24303 5519 | 16205 | -0.092 | -0.5902 | No | ||
| 110 | ATP5O | 22539 | 16280 | -0.097 | -0.5910 | No | ||
| 111 | ETFDH | 15313 | 16292 | -0.098 | -0.5884 | No | ||
| 112 | COX6A1 | 8774 4553 | 16575 | -0.120 | -0.5997 | Yes | ||
| 113 | NDUFB7 | 12429 | 16629 | -0.125 | -0.5984 | Yes | ||
| 114 | RPL19 | 9736 | 16635 | -0.125 | -0.5945 | Yes | ||
| 115 | NDUFS8 | 23950 | 16650 | -0.127 | -0.5911 | Yes | ||
| 116 | COX5B | 4552 | 16733 | -0.134 | -0.5911 | Yes | ||
| 117 | ATP5E | 14321 | 16764 | -0.137 | -0.5881 | Yes | ||
| 118 | SEC11A | 12144 | 16854 | -0.149 | -0.5880 | Yes | ||
| 119 | ATP5F1 | 15212 | 16941 | -0.158 | -0.5874 | Yes | ||
| 120 | NDUFS5 | 9296 | 16976 | -0.162 | -0.5839 | Yes | ||
| 121 | NDUFB8 | 7418 | 17006 | -0.165 | -0.5800 | Yes | ||
| 122 | FAU | 8954 | 17075 | -0.173 | -0.5779 | Yes | ||
| 123 | COX7C | 8776 | 17098 | -0.174 | -0.5733 | Yes | ||
| 124 | NDUFS2 | 4089 13760 | 17185 | -0.184 | -0.5719 | Yes | ||
| 125 | COX6B1 | 4283 8479 | 17188 | -0.184 | -0.5659 | Yes | ||
| 126 | ATP5G1 | 8636 | 17192 | -0.184 | -0.5599 | Yes | ||
| 127 | NDUFA4 | 9451 | 17294 | -0.197 | -0.5589 | Yes | ||
| 128 | ATP5J2 | 12186 | 17301 | -0.198 | -0.5526 | Yes | ||
| 129 | UQCRC2 | 18102 | 17307 | -0.198 | -0.5463 | Yes | ||
| 130 | PDHA1 | 24020 | 17331 | -0.200 | -0.5409 | Yes | ||
| 131 | IDH3A | 19447 | 17370 | -0.206 | -0.5362 | Yes | ||
| 132 | RPS19 | 5398 | 17385 | -0.208 | -0.5300 | Yes | ||
| 133 | NDUFA8 | 14605 | 17442 | -0.216 | -0.5259 | Yes | ||
| 134 | DDOST | 16021 | 17474 | -0.221 | -0.5202 | Yes | ||
| 135 | NDUFA2 | 9450 | 17556 | -0.232 | -0.5169 | Yes | ||
| 136 | NDUFS4 | 9452 5157 3173 | 17574 | -0.235 | -0.5100 | Yes | ||
| 137 | NDUFB3 | 12362 | 17582 | -0.236 | -0.5026 | Yes | ||
| 138 | UQCR | 12382 | 17638 | -0.244 | -0.4975 | Yes | ||
| 139 | RPL27A | 11181 6467 18130 | 17675 | -0.250 | -0.4911 | Yes | ||
| 140 | SLC30A7 | 15186 | 17705 | -0.254 | -0.4843 | Yes | ||
| 141 | NDUFV1 | 23953 | 17746 | -0.263 | -0.4777 | Yes | ||
| 142 | NDUFA9 | 12288 | 17800 | -0.276 | -0.4714 | Yes | ||
| 143 | SRP14 | 9894 | 17821 | -0.280 | -0.4632 | Yes | ||
| 144 | COX4I1 | 18444 | 17830 | -0.281 | -0.4543 | Yes | ||
| 145 | NDUFA11 | 12832 | 17879 | -0.294 | -0.4472 | Yes | ||
| 146 | RPS24 | 5399 | 17927 | -0.307 | -0.4395 | Yes | ||
| 147 | ATF4 | 22417 2239 | 17935 | -0.307 | -0.4297 | Yes | ||
| 148 | SSR2 | 15549 | 17937 | -0.308 | -0.4196 | Yes | ||
| 149 | NDUFA7 | 12347 | 17938 | -0.308 | -0.4094 | Yes | ||
| 150 | COX6C | 8775 | 18006 | -0.326 | -0.4022 | Yes | ||
| 151 | TPI1 | 5795 10212 | 18147 | -0.387 | -0.3969 | Yes | ||
| 152 | SEC61G | 9795 | 18160 | -0.394 | -0.3845 | Yes | ||
| 153 | DLD | 2097 21090 | 18373 | -0.571 | -0.3770 | Yes | ||
| 154 | NDUFS3 | 7553 12683 | 18471 | -0.850 | -0.3541 | Yes | ||
| 155 | IGFBP4 | 20670 9160 | 18493 | -0.936 | -0.3241 | Yes | ||
| 156 | SEC61B | 12305 7266 | 18503 | -0.999 | -0.2915 | Yes | ||
| 157 | ATP5J | 880 22555 | 18553 | -1.407 | -0.2474 | Yes | ||
| 158 | SUCLG1 | 17409 1059 | 18574 | -1.757 | -0.1902 | Yes | ||
| 159 | NDUFC1 | 7287 12339 | 18616 | -5.799 | -0.0000 | Yes |

