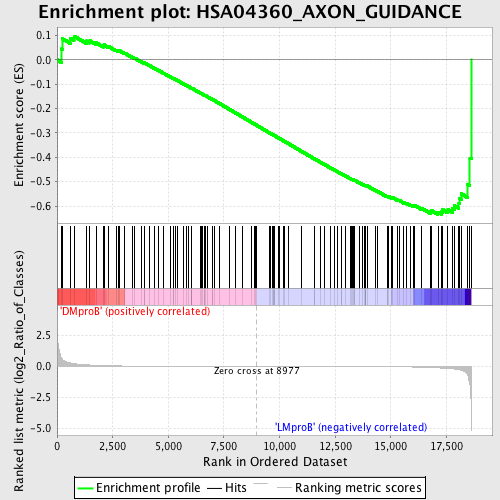
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_DMproB_versus_LMproB.phenotype_DMproB_versus_LMproB.cls #DMproB_versus_LMproB |
| Phenotype | phenotype_DMproB_versus_LMproB.cls#DMproB_versus_LMproB |
| Upregulated in class | LMproB |
| GeneSet | HSA04360_AXON_GUIDANCE |
| Enrichment Score (ES) | -0.63451946 |
| Normalized Enrichment Score (NES) | -1.4448322 |
| Nominal p-value | 0.015118791 |
| FDR q-value | 0.7891542 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ITGB1 | 3872 18411 | 188 | 0.692 | 0.0446 | No | ||
| 2 | ROBO3 | 9705 | 236 | 0.567 | 0.0869 | No | ||
| 3 | SLIT1 | 23678 | 613 | 0.268 | 0.0878 | No | ||
| 4 | RAC2 | 22217 | 760 | 0.220 | 0.0973 | No | ||
| 5 | CFL1 | 4516 | 1320 | 0.130 | 0.0774 | No | ||
| 6 | CDK5 | 16591 | 1476 | 0.113 | 0.0780 | No | ||
| 7 | PAK1 | 9527 | 1759 | 0.089 | 0.0697 | No | ||
| 8 | ROCK1 | 5386 | 2079 | 0.067 | 0.0578 | No | ||
| 9 | CXCR4 | 13844 | 2107 | 0.065 | 0.0615 | No | ||
| 10 | SRGAP2 | 13836 17045 6433 17046 | 2300 | 0.056 | 0.0555 | No | ||
| 11 | RASA1 | 10174 | 2654 | 0.041 | 0.0397 | No | ||
| 12 | GSK3B | 22761 | 2764 | 0.038 | 0.0368 | No | ||
| 13 | PTK2 | 22271 | 2806 | 0.036 | 0.0375 | No | ||
| 14 | NCK2 | 9448 | 3038 | 0.031 | 0.0274 | No | ||
| 15 | EPHA3 | 22563 1643 | 3392 | 0.024 | 0.0102 | No | ||
| 16 | SEMA4D | 9800 | 3495 | 0.022 | 0.0065 | No | ||
| 17 | FYN | 3375 3395 20052 | 3774 | 0.019 | -0.0071 | No | ||
| 18 | RND1 | 5868 | 3919 | 0.017 | -0.0135 | No | ||
| 19 | UNC5C | 10256 | 3921 | 0.017 | -0.0122 | No | ||
| 20 | NTN1 | 20400 | 4145 | 0.015 | -0.0231 | No | ||
| 21 | DPYSL5 | 92 | 4384 | 0.013 | -0.0349 | No | ||
| 22 | ROBO1 | 9731 | 4561 | 0.012 | -0.0435 | No | ||
| 23 | SEMA5B | 9801 22777 | 4781 | 0.011 | -0.0545 | No | ||
| 24 | EFNB2 | 18938 | 5075 | 0.009 | -0.0695 | No | ||
| 25 | NFATC2 | 5168 2866 | 5215 | 0.009 | -0.0764 | No | ||
| 26 | PPP3CB | 5285 | 5226 | 0.009 | -0.0762 | No | ||
| 27 | PPP3CA | 1863 5284 | 5331 | 0.008 | -0.0811 | No | ||
| 28 | SEMA3B | 5422 | 5397 | 0.008 | -0.0840 | No | ||
| 29 | NFATC3 | 5169 | 5702 | 0.007 | -0.0999 | No | ||
| 30 | SEMA6B | 1587 22932 | 5832 | 0.007 | -0.1063 | No | ||
| 31 | L1CAM | 3 | 5893 | 0.007 | -0.1090 | No | ||
| 32 | EPHB3 | 22816 | 6030 | 0.006 | -0.1159 | No | ||
| 33 | SEMA6D | 2800 14876 90 | 6051 | 0.006 | -0.1165 | No | ||
| 34 | UNC5D | 9980 | 6059 | 0.006 | -0.1164 | No | ||
| 35 | SEMA3C | 16914 | 6465 | 0.005 | -0.1379 | No | ||
| 36 | SLIT2 | 5456 | 6468 | 0.005 | -0.1376 | No | ||
| 37 | NFATC4 | 22002 | 6522 | 0.005 | -0.1401 | No | ||
| 38 | UNC5A | 3288 21631 | 6614 | 0.005 | -0.1447 | No | ||
| 39 | CFL2 | 2155 21069 | 6622 | 0.005 | -0.1447 | No | ||
| 40 | SEMA4F | 17105 | 6686 | 0.004 | -0.1478 | No | ||
| 41 | EPHA6 | 22564 | 6767 | 0.004 | -0.1518 | No | ||
| 42 | PLXNA3 | 24299 | 6973 | 0.004 | -0.1625 | No | ||
| 43 | EPHA5 | 16498 | 7006 | 0.004 | -0.1640 | No | ||
| 44 | SEMA3E | 16918 | 7064 | 0.004 | -0.1668 | No | ||
| 45 | MET | 17520 | 7293 | 0.003 | -0.1789 | No | ||
| 46 | SEMA4G | 11182 | 7745 | 0.002 | -0.2031 | No | ||
| 47 | NCK1 | 9447 5152 | 8016 | 0.002 | -0.2175 | No | ||
| 48 | ROBO2 | 6501 | 8353 | 0.001 | -0.2356 | No | ||
| 49 | SRGAP1 | 19615 | 8733 | 0.000 | -0.2561 | No | ||
| 50 | PLXNC1 | 7056 | 8873 | 0.000 | -0.2636 | No | ||
| 51 | EPHB2 | 4675 2440 8910 | 8933 | 0.000 | -0.2667 | No | ||
| 52 | SEMA3A | 16919 | 8963 | 0.000 | -0.2683 | No | ||
| 53 | FES | 17779 3949 | 9550 | -0.001 | -0.2999 | No | ||
| 54 | RHOA | 8624 4409 4410 | 9583 | -0.001 | -0.3016 | No | ||
| 55 | LIMK1 | 16350 | 9662 | -0.001 | -0.3057 | No | ||
| 56 | EFNB3 | 20391 | 9703 | -0.001 | -0.3078 | No | ||
| 57 | UNC5B | 8404 | 9718 | -0.001 | -0.3084 | No | ||
| 58 | ARHGEF12 | 19156 | 9754 | -0.001 | -0.3102 | No | ||
| 59 | NGEF | 13891 | 9789 | -0.001 | -0.3119 | No | ||
| 60 | PLXNA1 | 5268 | 9964 | -0.002 | -0.3212 | No | ||
| 61 | EPHA4 | 4673 | 9975 | -0.002 | -0.3216 | No | ||
| 62 | GNAI1 | 9024 | 10178 | -0.002 | -0.3324 | No | ||
| 63 | EPHB4 | 4676 | 10215 | -0.002 | -0.3342 | No | ||
| 64 | SEMA6A | 23439 9802 | 10413 | -0.002 | -0.3446 | No | ||
| 65 | NTN4 | 19903 | 10998 | -0.003 | -0.3759 | No | ||
| 66 | SEMA6C | 1797 5424 | 11586 | -0.005 | -0.4073 | No | ||
| 67 | EPHA7 | 8909 4674 | 11590 | -0.005 | -0.4071 | No | ||
| 68 | PAK3 | 9528 | 11832 | -0.005 | -0.4197 | No | ||
| 69 | EFNA4 | 1866 15277 | 12001 | -0.006 | -0.4283 | No | ||
| 70 | GNAI3 | 15198 | 12266 | -0.006 | -0.4421 | No | ||
| 71 | NRAS | 5191 | 12478 | -0.007 | -0.4530 | No | ||
| 72 | SEMA3D | 4240 | 12581 | -0.007 | -0.4579 | No | ||
| 73 | SLIT3 | 20925 | 12761 | -0.008 | -0.4670 | No | ||
| 74 | PPP3R2 | 9612 | 12785 | -0.008 | -0.4676 | No | ||
| 75 | EFNA2 | 8883 | 12954 | -0.008 | -0.4760 | No | ||
| 76 | PLXNB3 | 24304 | 13187 | -0.009 | -0.4878 | No | ||
| 77 | EPHB1 | 19024 | 13238 | -0.009 | -0.4898 | No | ||
| 78 | SEMA3F | 18999 3097 | 13291 | -0.010 | -0.4918 | No | ||
| 79 | EPHA8 | 15705 | 13300 | -0.010 | -0.4915 | No | ||
| 80 | CDC42 | 4503 8722 4504 2465 | 13354 | -0.010 | -0.4936 | No | ||
| 81 | ABLIM2 | 16866 3605 | 13376 | -0.010 | -0.4939 | No | ||
| 82 | PLXNB1 | 19301 | 13572 | -0.011 | -0.5036 | No | ||
| 83 | PAK7 | 14417 | 13716 | -0.012 | -0.5104 | No | ||
| 84 | ABL1 | 2693 4301 2794 | 13802 | -0.012 | -0.5140 | No | ||
| 85 | RAC3 | 20561 | 13871 | -0.013 | -0.5166 | No | ||
| 86 | EFNA1 | 15279 | 13875 | -0.013 | -0.5158 | No | ||
| 87 | EFNA3 | 15278 | 13880 | -0.013 | -0.5150 | No | ||
| 88 | RHOD | 23961 | 13939 | -0.013 | -0.5170 | No | ||
| 89 | PPP3R1 | 9611 489 | 14315 | -0.016 | -0.5361 | No | ||
| 90 | EPHA1 | 17168 8908 17167 | 14381 | -0.016 | -0.5383 | No | ||
| 91 | ABLIM1 | 3717 3766 3679 | 14834 | -0.021 | -0.5611 | No | ||
| 92 | SEMA4C | 9799 | 14845 | -0.021 | -0.5600 | No | ||
| 93 | DCC | 8839 | 14906 | -0.022 | -0.5615 | No | ||
| 94 | RAC1 | 16302 | 15046 | -0.024 | -0.5672 | No | ||
| 95 | RGS3 | 2434 2474 6937 11935 2374 2332 2445 2390 | 15048 | -0.024 | -0.5653 | No | ||
| 96 | SEMA5A | 22496 5423 | 15076 | -0.024 | -0.5649 | No | ||
| 97 | EFNA5 | 4655 8884 | 15300 | -0.028 | -0.5748 | No | ||
| 98 | CXCL12 | 9792 1182 1031 | 15370 | -0.029 | -0.5762 | No | ||
| 99 | ROCK2 | 21309 5387 | 15591 | -0.034 | -0.5854 | No | ||
| 100 | NTNG1 | 13484 8156 1814 | 15706 | -0.037 | -0.5886 | No | ||
| 101 | MAPK3 | 6458 11170 | 15904 | -0.043 | -0.5958 | No | ||
| 102 | SEMA7A | 19426 9803 | 16024 | -0.048 | -0.5985 | No | ||
| 103 | NFATC1 | 23398 1999 5167 9455 1985 1957 | 16084 | -0.050 | -0.5977 | No | ||
| 104 | EFNB1 | 24285 | 16392 | -0.064 | -0.6092 | No | ||
| 105 | PAK4 | 17909 | 16798 | -0.090 | -0.6240 | No | ||
| 106 | SEMA4B | 18205 | 16807 | -0.090 | -0.6173 | No | ||
| 107 | EPHB6 | 17473 | 17120 | -0.114 | -0.6251 | Yes | ||
| 108 | KRAS | 9247 | 17295 | -0.133 | -0.6240 | Yes | ||
| 109 | PAK2 | 10310 5875 | 17335 | -0.137 | -0.6152 | Yes | ||
| 110 | HRAS | 4868 | 17569 | -0.163 | -0.6149 | Yes | ||
| 111 | PLXNA2 | 5269 | 17782 | -0.192 | -0.6111 | Yes | ||
| 112 | SEMA4A | 15289 | 17861 | -0.205 | -0.5991 | Yes | ||
| 113 | EPHA2 | 16006 | 18060 | -0.262 | -0.5890 | Yes | ||
| 114 | MAPK1 | 1642 11167 | 18096 | -0.272 | -0.5694 | Yes | ||
| 115 | PPP3CC | 21763 | 18165 | -0.297 | -0.5496 | Yes | ||
| 116 | LIMK2 | 9278 1296 | 18453 | -0.686 | -0.5108 | Yes | ||
| 117 | NFAT5 | 3921 7037 12036 | 18556 | -1.432 | -0.4029 | Yes | ||
| 118 | DPYSL2 | 3122 4561 8787 | 18616 | -5.131 | -0.0000 | Yes |

