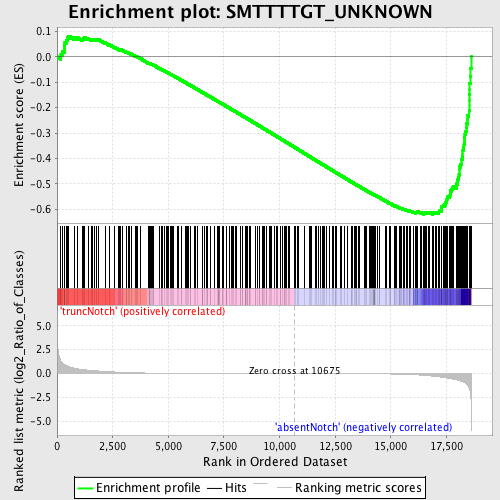
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #truncNotch_versus_absentNotch |
| Phenotype | phenotype_absentNotch_versus_truncNotch.cls#truncNotch_versus_absentNotch |
| Upregulated in class | absentNotch |
| GeneSet | SMTTTTGT_UNKNOWN |
| Enrichment Score (ES) | -0.6210938 |
| Normalized Enrichment Score (NES) | -1.7141966 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0074734064 |
| FWER p-Value | 0.034 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SFRS2 | 50707 380593 | 172 | 1.303 | 0.0094 | No | ||
| 2 | DUSP4 | 2690044 2850593 | 223 | 1.141 | 0.0232 | No | ||
| 3 | MAPKAPK2 | 2850500 | 323 | 0.956 | 0.0316 | No | ||
| 4 | ITPK1 | 3360056 | 329 | 0.946 | 0.0449 | No | ||
| 5 | SFRS10 | 6130528 6400438 7040112 | 351 | 0.925 | 0.0571 | No | ||
| 6 | SLC12A8 | 4810092 | 405 | 0.847 | 0.0665 | No | ||
| 7 | CCBL1 | 2350082 | 462 | 0.793 | 0.0748 | No | ||
| 8 | BLOC1S1 | 130341 3940717 | 522 | 0.724 | 0.0821 | No | ||
| 9 | DDX6 | 2630025 6770408 | 761 | 0.565 | 0.0772 | No | ||
| 10 | TARDBP | 870390 2350093 | 901 | 0.494 | 0.0768 | No | ||
| 11 | RAB33A | 6590195 | 1156 | 0.414 | 0.0689 | No | ||
| 12 | PRKAG1 | 6450091 | 1177 | 0.406 | 0.0737 | No | ||
| 13 | CCR7 | 2060687 | 1240 | 0.388 | 0.0759 | No | ||
| 14 | LCP1 | 4280397 | 1393 | 0.353 | 0.0727 | No | ||
| 15 | MAP4K2 | 4200037 | 1530 | 0.324 | 0.0700 | No | ||
| 16 | ATP5A1 | 1990722 | 1603 | 0.311 | 0.0705 | No | ||
| 17 | EHD1 | 3840731 | 1685 | 0.295 | 0.0704 | No | ||
| 18 | ESRRA | 6550242 | 1784 | 0.278 | 0.0690 | No | ||
| 19 | SLC3A2 | 5270358 | 1878 | 0.262 | 0.0677 | No | ||
| 20 | EPN1 | 4060725 | 2163 | 0.221 | 0.0554 | No | ||
| 21 | BTBD9 | 2450025 | 2346 | 0.194 | 0.0483 | No | ||
| 22 | LMO6 | 4210072 | 2587 | 0.163 | 0.0376 | No | ||
| 23 | XKRX | 3800192 | 2777 | 0.136 | 0.0292 | No | ||
| 24 | EPC1 | 1570050 5290095 6900193 | 2790 | 0.135 | 0.0305 | No | ||
| 25 | CAMK2G | 630097 | 2870 | 0.127 | 0.0280 | No | ||
| 26 | UBE2S | 1850110 | 2922 | 0.122 | 0.0270 | No | ||
| 27 | POU2F1 | 70577 430373 4850324 5910056 | 3115 | 0.103 | 0.0180 | No | ||
| 28 | MKRN1 | 1410097 | 3118 | 0.102 | 0.0194 | No | ||
| 29 | SFRS8 | 5130162 5700164 | 3206 | 0.096 | 0.0160 | No | ||
| 30 | NEUROG1 | 380341 | 3275 | 0.091 | 0.0136 | No | ||
| 31 | TNFSF10 | 3170300 | 3325 | 0.088 | 0.0122 | No | ||
| 32 | KCNH7 | 7510068 | 3514 | 0.077 | 0.0031 | No | ||
| 33 | NIN | 3450576 5080048 | 3547 | 0.075 | 0.0024 | No | ||
| 34 | FGF8 | 6350504 | 3631 | 0.071 | -0.0011 | No | ||
| 35 | DAAM1 | 1500273 | 3739 | 0.065 | -0.0060 | No | ||
| 36 | BCL11A | 6860369 | 4121 | 0.052 | -0.0260 | No | ||
| 37 | DPYSL5 | 4900333 | 4136 | 0.052 | -0.0260 | No | ||
| 38 | SP8 | 4060576 | 4155 | 0.051 | -0.0263 | No | ||
| 39 | RRAS2 | 6290170 | 4167 | 0.051 | -0.0261 | No | ||
| 40 | PTGS1 | 730193 1780273 6450195 | 4189 | 0.050 | -0.0266 | No | ||
| 41 | DPF3 | 2650128 2970168 | 4231 | 0.049 | -0.0281 | No | ||
| 42 | BMPR2 | 5340066 6960328 | 4265 | 0.048 | -0.0292 | No | ||
| 43 | NEUROD1 | 3060619 | 4321 | 0.047 | -0.0315 | No | ||
| 44 | FOXP2 | 3520561 4150372 4760524 | 4343 | 0.046 | -0.0320 | No | ||
| 45 | ITGB4 | 1740021 3840482 | 4602 | 0.039 | -0.0455 | No | ||
| 46 | TRPM8 | 2340706 | 4683 | 0.038 | -0.0493 | No | ||
| 47 | IRAK1 | 4120593 | 4692 | 0.037 | -0.0492 | No | ||
| 48 | HOXC6 | 3850133 4920736 | 4742 | 0.036 | -0.0514 | No | ||
| 49 | HOXB3 | 520292 | 4838 | 0.035 | -0.0561 | No | ||
| 50 | BACH2 | 6760131 | 4924 | 0.033 | -0.0602 | No | ||
| 51 | PDGFB | 3060440 6370008 | 4977 | 0.032 | -0.0626 | No | ||
| 52 | PCDHA11 | 1510253 | 5005 | 0.032 | -0.0636 | No | ||
| 53 | PROX1 | 4210193 | 5083 | 0.031 | -0.0674 | No | ||
| 54 | FZD5 | 4070452 | 5131 | 0.029 | -0.0695 | No | ||
| 55 | SOX5 | 2370576 2900167 3190128 5050528 | 5174 | 0.029 | -0.0714 | No | ||
| 56 | CNKSR2 | 110348 1770372 | 5226 | 0.028 | -0.0738 | No | ||
| 57 | ONECUT1 | 6450136 | 5413 | 0.025 | -0.0835 | No | ||
| 58 | NHLH2 | 6200021 | 5458 | 0.025 | -0.0856 | No | ||
| 59 | A2BP1 | 2370390 4590593 5550014 | 5581 | 0.023 | -0.0919 | No | ||
| 60 | CLK1 | 1780551 2100102 2340347 2480347 | 5604 | 0.023 | -0.0928 | No | ||
| 61 | GGN | 1410528 4210082 5290170 | 5777 | 0.021 | -0.1018 | No | ||
| 62 | ATCAY | 4070494 | 5796 | 0.021 | -0.1025 | No | ||
| 63 | SEMA4C | 1980315 | 5881 | 0.020 | -0.1068 | No | ||
| 64 | ARID4A | 520156 1050707 | 5909 | 0.020 | -0.1080 | No | ||
| 65 | SCG3 | 1690050 | 5973 | 0.019 | -0.1112 | No | ||
| 66 | KIF1B | 1240494 2370139 4570270 6510102 | 6007 | 0.019 | -0.1127 | No | ||
| 67 | ATRX | 2340039 3610132 4210373 | 6152 | 0.018 | -0.1203 | No | ||
| 68 | HOXB5 | 2900538 | 6224 | 0.017 | -0.1239 | No | ||
| 69 | SMPD3 | 520164 | 6293 | 0.016 | -0.1274 | No | ||
| 70 | LHX6 | 2360731 3360397 | 6512 | 0.015 | -0.1391 | No | ||
| 71 | LMOD1 | 2120035 | 6544 | 0.014 | -0.1406 | No | ||
| 72 | CRYGC | 1660338 | 6549 | 0.014 | -0.1406 | No | ||
| 73 | GFPT2 | 2370129 | 6616 | 0.014 | -0.1440 | No | ||
| 74 | DIO3 | 6180292 | 6697 | 0.013 | -0.1481 | No | ||
| 75 | HOXD9 | 2570181 | 6739 | 0.013 | -0.1502 | No | ||
| 76 | MID1 | 4070114 4760458 | 6745 | 0.013 | -0.1503 | No | ||
| 77 | EPN2 | 1340451 6550239 6860020 6860403 | 6897 | 0.012 | -0.1583 | No | ||
| 78 | MTUS1 | 780348 4920609 | 6901 | 0.012 | -0.1583 | No | ||
| 79 | CALD1 | 1770129 1940397 | 7055 | 0.011 | -0.1665 | No | ||
| 80 | GNAQ | 430670 4210131 5900736 | 7211 | 0.011 | -0.1748 | No | ||
| 81 | SMAD1 | 630537 1850333 | 7237 | 0.011 | -0.1760 | No | ||
| 82 | GPM6A | 1660044 2750152 | 7246 | 0.011 | -0.1763 | No | ||
| 83 | SLC30A3 | 1850451 | 7267 | 0.010 | -0.1772 | No | ||
| 84 | KLHL4 | 5080438 | 7290 | 0.010 | -0.1783 | No | ||
| 85 | APLN | 6620632 | 7292 | 0.010 | -0.1782 | No | ||
| 86 | NEUROD6 | 4670731 | 7430 | 0.010 | -0.1855 | No | ||
| 87 | ARHGEF17 | 2060008 | 7441 | 0.010 | -0.1859 | No | ||
| 88 | DHRS3 | 360609 | 7478 | 0.009 | -0.1878 | No | ||
| 89 | LMO4 | 3800746 | 7606 | 0.009 | -0.1946 | No | ||
| 90 | JARID2 | 6290538 | 7627 | 0.009 | -0.1955 | No | ||
| 91 | FGFR4 | 2650072 | 7629 | 0.009 | -0.1955 | No | ||
| 92 | FLRT3 | 70685 2760497 6040519 | 7758 | 0.008 | -0.2023 | No | ||
| 93 | DAB1 | 2060193 7000605 | 7850 | 0.008 | -0.2072 | No | ||
| 94 | FBN2 | 50435 | 7862 | 0.008 | -0.2077 | No | ||
| 95 | LUC7L2 | 1850402 6350088 | 7925 | 0.008 | -0.2109 | No | ||
| 96 | SRGAP1 | 3940253 | 8004 | 0.008 | -0.2151 | No | ||
| 97 | ALDOC | 450121 610427 | 8041 | 0.007 | -0.2169 | No | ||
| 98 | DMD | 1740041 3990332 | 8245 | 0.007 | -0.2279 | No | ||
| 99 | FRAS1 | 5080170 5670347 | 8263 | 0.007 | -0.2287 | No | ||
| 100 | HOXA5 | 6840026 | 8322 | 0.006 | -0.2318 | No | ||
| 101 | MOG | 610364 6020722 | 8329 | 0.006 | -0.2321 | No | ||
| 102 | GPR110 | 1190215 | 8452 | 0.006 | -0.2386 | No | ||
| 103 | SOX3 | 4570537 | 8474 | 0.006 | -0.2397 | No | ||
| 104 | ROR1 | 6220026 | 8478 | 0.006 | -0.2398 | No | ||
| 105 | POU6F2 | 5270451 | 8515 | 0.006 | -0.2416 | No | ||
| 106 | NRG1 | 1050332 | 8543 | 0.006 | -0.2430 | No | ||
| 107 | BMX | 50136 2940242 4540059 | 8563 | 0.006 | -0.2440 | No | ||
| 108 | GATM | 380671 | 8630 | 0.005 | -0.2475 | No | ||
| 109 | SLC10A1 | 2570154 | 8652 | 0.005 | -0.2486 | No | ||
| 110 | TIAL1 | 4150048 6510605 | 8678 | 0.005 | -0.2499 | No | ||
| 111 | SRGAP2 | 1050092 1230403 3830066 6130215 | 8914 | 0.005 | -0.2626 | No | ||
| 112 | HOXD12 | 580102 | 8934 | 0.005 | -0.2636 | No | ||
| 113 | SS18 | 1940440 2900072 6900129 | 8997 | 0.004 | -0.2669 | No | ||
| 114 | CHN1 | 6040441 | 9076 | 0.004 | -0.2711 | No | ||
| 115 | EHF | 4560088 | 9080 | 0.004 | -0.2712 | No | ||
| 116 | ZBTB11 | 6520092 | 9247 | 0.004 | -0.2802 | No | ||
| 117 | LMO1 | 3170528 | 9249 | 0.004 | -0.2802 | No | ||
| 118 | GABRA2 | 3290167 | 9278 | 0.004 | -0.2817 | No | ||
| 119 | TTL | 6220048 | 9332 | 0.003 | -0.2845 | No | ||
| 120 | MTSS1 | 780435 2370114 | 9421 | 0.003 | -0.2893 | No | ||
| 121 | PURA | 3870156 | 9426 | 0.003 | -0.2895 | No | ||
| 122 | NF1 | 6980433 | 9533 | 0.003 | -0.2952 | No | ||
| 123 | TRERF1 | 6370017 6940138 | 9584 | 0.003 | -0.2979 | No | ||
| 124 | CHES1 | 4570121 5700162 | 9627 | 0.003 | -0.3001 | No | ||
| 125 | CBX5 | 3830072 6290167 | 9785 | 0.002 | -0.3087 | No | ||
| 126 | SATB2 | 2470181 | 9863 | 0.002 | -0.3129 | No | ||
| 127 | NF2 | 4150735 6450139 | 9906 | 0.002 | -0.3151 | No | ||
| 128 | UNC5C | 2640056 | 10018 | 0.002 | -0.3212 | No | ||
| 129 | PCDH8 | 2360047 4200452 | 10061 | 0.002 | -0.3234 | No | ||
| 130 | GLRA2 | 7000019 | 10134 | 0.001 | -0.3273 | No | ||
| 131 | LHX3 | 670164 1770280 | 10202 | 0.001 | -0.3310 | No | ||
| 132 | CACNG3 | 520093 | 10207 | 0.001 | -0.3312 | No | ||
| 133 | DDX5 | 940021 2630338 3440148 | 10260 | 0.001 | -0.3340 | No | ||
| 134 | LHX9 | 2350112 | 10311 | 0.001 | -0.3367 | No | ||
| 135 | CCNJ | 870047 | 10416 | 0.001 | -0.3424 | No | ||
| 136 | CNGB3 | 2340408 | 10464 | 0.000 | -0.3449 | No | ||
| 137 | CRYGA | 2340309 | 10687 | -0.000 | -0.3570 | No | ||
| 138 | KIRREL3 | 2630195 | 10727 | -0.000 | -0.3592 | No | ||
| 139 | WNT9A | 3830711 | 10796 | -0.000 | -0.3629 | No | ||
| 140 | NRXN1 | 460408 5340270 6370066 | 10842 | -0.000 | -0.3653 | No | ||
| 141 | DSTN | 3140592 | 11141 | -0.001 | -0.3816 | No | ||
| 142 | SPRR4 | 5720273 | 11343 | -0.002 | -0.3925 | No | ||
| 143 | HOXA4 | 940152 | 11385 | -0.002 | -0.3947 | No | ||
| 144 | KLF12 | 1660095 4810288 5340546 6520286 | 11455 | -0.002 | -0.3984 | No | ||
| 145 | ENPP2 | 5860546 | 11610 | -0.002 | -0.4068 | No | ||
| 146 | PCYT1B | 1780402 | 11673 | -0.003 | -0.4102 | No | ||
| 147 | NTRK2 | 6220463 | 11740 | -0.003 | -0.4137 | No | ||
| 148 | ELF5 | 3140156 5220020 | 11823 | -0.003 | -0.4181 | No | ||
| 149 | OMG | 6760066 | 11937 | -0.003 | -0.4243 | No | ||
| 150 | GATA2 | 6590280 | 11955 | -0.004 | -0.4251 | No | ||
| 151 | ISL1 | 4810239 | 12031 | -0.004 | -0.4292 | No | ||
| 152 | ETV1 | 70735 2940603 5080463 | 12126 | -0.004 | -0.4342 | No | ||
| 153 | SET | 6650286 | 12246 | -0.004 | -0.4407 | No | ||
| 154 | DDX3Y | 1580278 4200519 | 12364 | -0.005 | -0.4470 | No | ||
| 155 | HOXD11 | 2260242 | 12402 | -0.005 | -0.4489 | No | ||
| 156 | NOX4 | 6040136 | 12517 | -0.006 | -0.4551 | No | ||
| 157 | LRRC4 | 3850041 | 12521 | -0.006 | -0.4552 | No | ||
| 158 | SPO11 | 540288 4780239 | 12540 | -0.006 | -0.4561 | No | ||
| 159 | EDN3 | 2690722 | 12755 | -0.007 | -0.4676 | No | ||
| 160 | HOXD4 | 5900301 | 12777 | -0.007 | -0.4687 | No | ||
| 161 | HPCAL1 | 1980129 2350348 | 12896 | -0.007 | -0.4750 | No | ||
| 162 | D4ST1 | 1780671 | 13030 | -0.008 | -0.4822 | No | ||
| 163 | PELI2 | 1940364 | 13216 | -0.009 | -0.4921 | No | ||
| 164 | CLDN2 | 580053 | 13245 | -0.009 | -0.4935 | No | ||
| 165 | GPC3 | 6180735 | 13268 | -0.009 | -0.4946 | No | ||
| 166 | SP3 | 3840338 | 13383 | -0.010 | -0.5007 | No | ||
| 167 | HOXA2 | 2120121 | 13430 | -0.010 | -0.5030 | No | ||
| 168 | QKI | 5220093 6130044 | 13435 | -0.010 | -0.5031 | No | ||
| 169 | GPR21 | 6520152 | 13470 | -0.010 | -0.5048 | No | ||
| 170 | SHH | 5570400 | 13531 | -0.011 | -0.5079 | No | ||
| 171 | GPR12 | 610288 | 13583 | -0.011 | -0.5106 | No | ||
| 172 | DOC2B | 2190021 2850142 | 13809 | -0.014 | -0.5226 | No | ||
| 173 | ERG | 50154 1770739 | 13880 | -0.014 | -0.5262 | No | ||
| 174 | CPNE1 | 3440131 | 13920 | -0.015 | -0.5282 | No | ||
| 175 | NR4A3 | 2900021 5860095 5910039 | 14052 | -0.017 | -0.5351 | No | ||
| 176 | CSNK1A1 | 2340427 | 14088 | -0.017 | -0.5367 | No | ||
| 177 | PNRC2 | 3360309 | 14147 | -0.018 | -0.5396 | No | ||
| 178 | NLGN3 | 2650463 | 14173 | -0.018 | -0.5407 | No | ||
| 179 | GNAO1 | 2510292 6590487 | 14218 | -0.019 | -0.5428 | No | ||
| 180 | PTP4A1 | 130692 | 14223 | -0.019 | -0.5428 | No | ||
| 181 | THRAP1 | 2470121 | 14234 | -0.020 | -0.5430 | No | ||
| 182 | RCOR1 | 5360717 | 14257 | -0.020 | -0.5440 | No | ||
| 183 | NR6A1 | 4010347 | 14259 | -0.020 | -0.5437 | No | ||
| 184 | SLC7A1 | 2450193 2710241 | 14283 | -0.020 | -0.5447 | No | ||
| 185 | LRP1 | 6270386 | 14286 | -0.021 | -0.5445 | No | ||
| 186 | USP2 | 1190292 1240253 4850035 | 14290 | -0.021 | -0.5444 | No | ||
| 187 | MYO10 | 3830576 5570092 | 14407 | -0.023 | -0.5504 | No | ||
| 188 | P4HA1 | 2630692 | 14496 | -0.025 | -0.5548 | No | ||
| 189 | DUSP6 | 5910286 7100070 | 14508 | -0.025 | -0.5550 | No | ||
| 190 | PPP1R8 | 6290292 | 14743 | -0.033 | -0.5673 | No | ||
| 191 | LTBP1 | 780035 2100746 2760402 3780551 6550184 6860193 | 14802 | -0.035 | -0.5700 | No | ||
| 192 | PCF11 | 1240180 | 14956 | -0.042 | -0.5777 | No | ||
| 193 | EN1 | 5360168 7050025 | 14967 | -0.042 | -0.5777 | No | ||
| 194 | ZFP36L1 | 2510138 4120048 | 15003 | -0.044 | -0.5790 | No | ||
| 195 | PDE4D | 2470528 6660014 | 15170 | -0.052 | -0.5873 | No | ||
| 196 | ELAVL4 | 50735 3360086 5220167 | 15185 | -0.053 | -0.5873 | No | ||
| 197 | MAGED1 | 6650332 | 15231 | -0.055 | -0.5889 | No | ||
| 198 | CITED2 | 5670114 5130088 | 15257 | -0.057 | -0.5895 | No | ||
| 199 | SLC38A1 | 520465 | 15379 | -0.064 | -0.5951 | No | ||
| 200 | ZFP161 | 110546 4810195 | 15384 | -0.065 | -0.5944 | No | ||
| 201 | EDARADD | 840180 2360435 4560014 | 15452 | -0.069 | -0.5971 | No | ||
| 202 | IGF1 | 1990193 3130377 3290280 | 15490 | -0.071 | -0.5981 | No | ||
| 203 | NR4A2 | 60273 | 15551 | -0.075 | -0.6003 | No | ||
| 204 | PHF15 | 6550066 6860673 | 15583 | -0.078 | -0.6008 | No | ||
| 205 | GPR85 | 5130750 5910020 | 15609 | -0.080 | -0.6010 | No | ||
| 206 | STC1 | 360161 | 15685 | -0.086 | -0.6039 | No | ||
| 207 | CHD2 | 4070039 | 15749 | -0.092 | -0.6060 | No | ||
| 208 | KCTD15 | 2810735 | 15818 | -0.098 | -0.6083 | No | ||
| 209 | PABPC4 | 1990170 6760270 5390138 | 15843 | -0.099 | -0.6082 | No | ||
| 210 | PANK3 | 3060687 | 15845 | -0.099 | -0.6068 | No | ||
| 211 | RAB10 | 2360008 5890020 | 15894 | -0.105 | -0.6079 | No | ||
| 212 | CDK2AP1 | 2340156 | 16020 | -0.119 | -0.6130 | No | ||
| 213 | RBM12 | 4570239 6040411 6660129 | 16029 | -0.120 | -0.6117 | No | ||
| 214 | CSAD | 770215 | 16103 | -0.126 | -0.6139 | No | ||
| 215 | KCNN3 | 6520600 6420138 | 16128 | -0.129 | -0.6133 | No | ||
| 216 | CLASP1 | 6860279 | 16134 | -0.130 | -0.6117 | No | ||
| 217 | YWHAE | 5310435 | 16190 | -0.138 | -0.6127 | No | ||
| 218 | H2AFY | 4730750 | 16198 | -0.139 | -0.6111 | No | ||
| 219 | COX7B | 2340504 | 16215 | -0.142 | -0.6099 | No | ||
| 220 | PCBP2 | 5690048 | 16316 | -0.157 | -0.6131 | No | ||
| 221 | BTK | 3130044 | 16356 | -0.162 | -0.6129 | No | ||
| 222 | SFRS5 | 3450176 6350008 | 16484 | -0.186 | -0.6172 | Yes | ||
| 223 | TPP2 | 3850093 | 16519 | -0.192 | -0.6163 | Yes | ||
| 224 | NRF1 | 2650195 | 16530 | -0.194 | -0.6140 | Yes | ||
| 225 | FKBP5 | 2190048 | 16537 | -0.195 | -0.6115 | Yes | ||
| 226 | YWHAG | 3780341 | 16606 | -0.207 | -0.6122 | Yes | ||
| 227 | PHF12 | 870400 3130687 | 16672 | -0.222 | -0.6126 | Yes | ||
| 228 | FUS | 4760619 | 16757 | -0.242 | -0.6137 | Yes | ||
| 229 | AMMECR1 | 3800435 | 16894 | -0.274 | -0.6171 | Yes | ||
| 230 | YWHAQ | 6760524 | 16921 | -0.282 | -0.6145 | Yes | ||
| 231 | APP | 2510053 | 16987 | -0.298 | -0.6137 | Yes | ||
| 232 | TCF4 | 520021 | 17052 | -0.320 | -0.6126 | Yes | ||
| 233 | H1FX | 1190088 | 17141 | -0.341 | -0.6125 | Yes | ||
| 234 | YTHDF2 | 4060332 4610279 | 17165 | -0.348 | -0.6087 | Yes | ||
| 235 | OLIG3 | 1050603 | 17204 | -0.356 | -0.6057 | Yes | ||
| 236 | TCF7 | 3800736 5390181 | 17271 | -0.374 | -0.6039 | Yes | ||
| 237 | SLC38A2 | 1470242 3800026 | 17287 | -0.378 | -0.5992 | Yes | ||
| 238 | SFRS3 | 770315 4230593 | 17293 | -0.381 | -0.5940 | Yes | ||
| 239 | IDH2 | 2680397 | 17295 | -0.381 | -0.5886 | Yes | ||
| 240 | HN1 | 3360156 | 17350 | -0.401 | -0.5857 | Yes | ||
| 241 | AKAP8L | 5290452 | 17411 | -0.422 | -0.5829 | Yes | ||
| 242 | GSK3B | 5360348 | 17444 | -0.435 | -0.5784 | Yes | ||
| 243 | MSN | 4060044 | 17465 | -0.443 | -0.5731 | Yes | ||
| 244 | ICK | 1580746 3140021 | 17495 | -0.454 | -0.5681 | Yes | ||
| 245 | CRKL | 4050427 | 17522 | -0.462 | -0.5628 | Yes | ||
| 246 | CD4 | 1090010 | 17532 | -0.466 | -0.5566 | Yes | ||
| 247 | RYBP | 60746 | 17563 | -0.481 | -0.5513 | Yes | ||
| 248 | MAP3K7IP2 | 2340242 | 17639 | -0.508 | -0.5481 | Yes | ||
| 249 | ARPC2 | 6020484 | 17661 | -0.516 | -0.5418 | Yes | ||
| 250 | HNRPC | 1570133 3610131 | 17672 | -0.523 | -0.5348 | Yes | ||
| 251 | TRIM8 | 6130441 | 17683 | -0.528 | -0.5277 | Yes | ||
| 252 | STMN1 | 1990717 | 17737 | -0.551 | -0.5226 | Yes | ||
| 253 | CLK4 | 3440484 3610156 | 17773 | -0.564 | -0.5164 | Yes | ||
| 254 | KLF7 | 50390 | 17829 | -0.591 | -0.5109 | Yes | ||
| 255 | TIAM1 | 5420288 | 17957 | -0.655 | -0.5083 | Yes | ||
| 256 | SMOC2 | 1170170 | 17969 | -0.661 | -0.4994 | Yes | ||
| 257 | ETS2 | 360451 | 18011 | -0.686 | -0.4917 | Yes | ||
| 258 | WBP4 | 2230019 | 18014 | -0.688 | -0.4819 | Yes | ||
| 259 | SIN3A | 2190121 | 18047 | -0.708 | -0.4734 | Yes | ||
| 260 | CXCR4 | 4590519 | 18063 | -0.720 | -0.4639 | Yes | ||
| 261 | DDX3X | 2190020 | 18098 | -0.751 | -0.4549 | Yes | ||
| 262 | ACLY | 1740609 | 18099 | -0.751 | -0.4440 | Yes | ||
| 263 | RNF44 | 5910692 | 18104 | -0.755 | -0.4334 | Yes | ||
| 264 | CCAR1 | 3190092 | 18130 | -0.771 | -0.4236 | Yes | ||
| 265 | FOXP1 | 6510433 | 18175 | -0.810 | -0.4143 | Yes | ||
| 266 | SRPK2 | 6380341 | 18176 | -0.812 | -0.4026 | Yes | ||
| 267 | SFPQ | 4760110 | 18222 | -0.850 | -0.3928 | Yes | ||
| 268 | JUP | 2510671 | 18238 | -0.871 | -0.3810 | Yes | ||
| 269 | MAPK6 | 6760520 | 18240 | -0.875 | -0.3684 | Yes | ||
| 270 | IPO11 | 70167 | 18251 | -0.888 | -0.3562 | Yes | ||
| 271 | AKAP8 | 4210026 | 18288 | -0.926 | -0.3448 | Yes | ||
| 272 | MBNL1 | 2640762 7100048 | 18303 | -0.941 | -0.3319 | Yes | ||
| 273 | PIAS1 | 5340301 | 18308 | -0.946 | -0.3185 | Yes | ||
| 274 | HNRPA1 | 4670398 5050687 | 18332 | -0.983 | -0.3056 | Yes | ||
| 275 | NFIX | 2450152 | 18366 | -1.032 | -0.2925 | Yes | ||
| 276 | TLK1 | 7100605 | 18394 | -1.071 | -0.2785 | Yes | ||
| 277 | GADD45G | 2510142 | 18408 | -1.113 | -0.2631 | Yes | ||
| 278 | BCL6 | 940100 | 18435 | -1.170 | -0.2477 | Yes | ||
| 279 | CTCF | 5340017 | 18455 | -1.243 | -0.2308 | Yes | ||
| 280 | RBM3 | 2360739 5700167 | 18513 | -1.452 | -0.2129 | Yes | ||
| 281 | HMGB1 | 2120670 2350044 | 18518 | -1.471 | -0.1919 | Yes | ||
| 282 | CNN3 | 6110020 | 18519 | -1.472 | -0.1706 | Yes | ||
| 283 | EIF4G2 | 3800575 6860184 | 18522 | -1.497 | -0.1492 | Yes | ||
| 284 | ITPR1 | 3450519 | 18537 | -1.590 | -0.1270 | Yes | ||
| 285 | RBM5 | 3520402 5270121 | 18540 | -1.603 | -0.1039 | Yes | ||
| 286 | ILF3 | 940722 3190647 6520110 | 18576 | -2.079 | -0.0758 | Yes | ||
| 287 | RBMX | 7100162 2900541 | 18589 | -2.229 | -0.0443 | Yes | ||
| 288 | MMP14 | 670079 | 18610 | -3.170 | 0.0003 | Yes |

