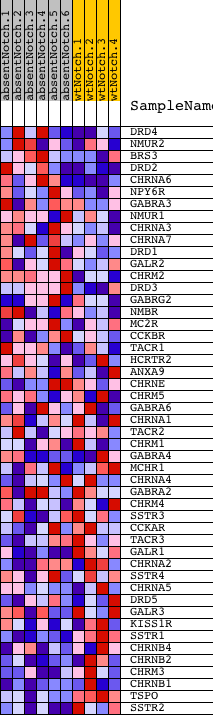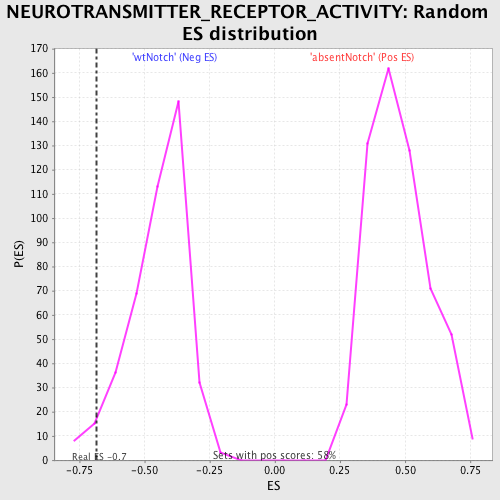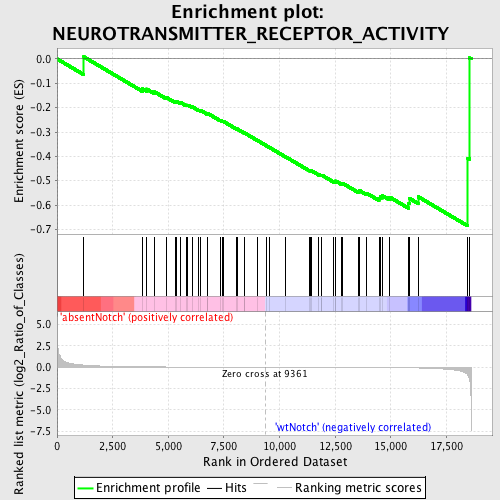
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_wtNotch.phenotype_absentNotch_versus_wtNotch.cls #absentNotch_versus_wtNotch.phenotype_absentNotch_versus_wtNotch.cls #absentNotch_versus_wtNotch_repos |
| Phenotype | phenotype_absentNotch_versus_wtNotch.cls#absentNotch_versus_wtNotch_repos |
| Upregulated in class | wtNotch |
| GeneSet | NEUROTRANSMITTER_RECEPTOR_ACTIVITY |
| Enrichment Score (ES) | -0.6858389 |
| Normalized Enrichment Score (NES) | -1.5278367 |
| Nominal p-value | 0.028301887 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DRD4 | 6350390 | 1201 | 0.241 | 0.0085 | No | ||
| 2 | NMUR2 | 1580717 | 3832 | 0.034 | -0.1227 | No | ||
| 3 | BRS3 | 1740403 | 4017 | 0.030 | -0.1234 | No | ||
| 4 | DRD2 | 5890369 | 4379 | 0.025 | -0.1353 | No | ||
| 5 | CHRNA6 | 5340092 | 4919 | 0.018 | -0.1587 | No | ||
| 6 | NPY6R | 4560136 | 5310 | 0.015 | -0.1751 | No | ||
| 7 | GABRA3 | 4810142 | 5356 | 0.015 | -0.1729 | No | ||
| 8 | NMUR1 | 1500403 | 5531 | 0.014 | -0.1781 | No | ||
| 9 | CHRNA3 | 6760100 | 5805 | 0.012 | -0.1891 | No | ||
| 10 | CHRNA7 | 2970446 | 5873 | 0.012 | -0.1891 | No | ||
| 11 | DRD1 | 430025 | 6076 | 0.011 | -0.1968 | No | ||
| 12 | GALR2 | 6450739 | 6370 | 0.009 | -0.2097 | No | ||
| 13 | CHRM2 | 870750 | 6462 | 0.009 | -0.2119 | No | ||
| 14 | DRD3 | 4780402 | 6739 | 0.008 | -0.2244 | No | ||
| 15 | GABRG2 | 2350402 4210204 6130279 6550037 | 6773 | 0.008 | -0.2239 | No | ||
| 16 | NMBR | 6180315 | 7348 | 0.006 | -0.2531 | No | ||
| 17 | MC2R | 2970504 | 7434 | 0.005 | -0.2560 | No | ||
| 18 | CCKBR | 2760128 | 7456 | 0.005 | -0.2556 | No | ||
| 19 | TACR1 | 70358 3840411 | 8066 | 0.003 | -0.2873 | No | ||
| 20 | HCRTR2 | 2350463 | 8085 | 0.003 | -0.2873 | No | ||
| 21 | ANXA9 | 1190195 | 8410 | 0.003 | -0.3039 | No | ||
| 22 | CHRNE | 3190170 | 8421 | 0.003 | -0.3037 | No | ||
| 23 | CHRM5 | 5390333 | 9007 | 0.001 | -0.3349 | No | ||
| 24 | GABRA6 | 4120520 | 9401 | -0.000 | -0.3561 | No | ||
| 25 | CHRNA1 | 1170025 3610364 5220292 | 9556 | -0.001 | -0.3642 | No | ||
| 26 | TACR2 | 1740358 | 10274 | -0.002 | -0.4021 | No | ||
| 27 | CHRM1 | 4280619 | 11355 | -0.006 | -0.4586 | No | ||
| 28 | GABRA4 | 6290465 | 11411 | -0.006 | -0.4597 | No | ||
| 29 | MCHR1 | 1780022 | 11437 | -0.006 | -0.4593 | No | ||
| 30 | CHRNA4 | 730075 2680091 | 11768 | -0.007 | -0.4749 | No | ||
| 31 | GABRA2 | 3290167 | 11870 | -0.008 | -0.4780 | No | ||
| 32 | CHRM4 | 1770427 | 11901 | -0.008 | -0.4773 | No | ||
| 33 | SSTR3 | 5420064 | 12441 | -0.010 | -0.5033 | No | ||
| 34 | CCKAR | 4210079 | 12494 | -0.010 | -0.5029 | No | ||
| 35 | TACR3 | 4780017 | 12520 | -0.010 | -0.5011 | No | ||
| 36 | GALR1 | 6020452 | 12792 | -0.012 | -0.5121 | No | ||
| 37 | CHRNA2 | 5050315 | 12820 | -0.012 | -0.5099 | No | ||
| 38 | SSTR4 | 6100398 | 13552 | -0.018 | -0.5439 | No | ||
| 39 | CHRNA5 | 580195 | 13586 | -0.018 | -0.5402 | No | ||
| 40 | DRD5 | 2970133 | 13905 | -0.021 | -0.5509 | No | ||
| 41 | GALR3 | 4850113 | 14471 | -0.029 | -0.5724 | No | ||
| 42 | KISS1R | 2480601 | 14519 | -0.030 | -0.5658 | No | ||
| 43 | SSTR1 | 2630471 | 14617 | -0.032 | -0.5614 | No | ||
| 44 | CHRNB4 | 6370110 | 14938 | -0.039 | -0.5669 | No | ||
| 45 | CHRNB2 | 580204 3120739 | 15798 | -0.069 | -0.5922 | Yes | ||
| 46 | CHRM3 | 6110725 | 15839 | -0.071 | -0.5728 | Yes | ||
| 47 | CHRNB1 | 1190671 | 16233 | -0.092 | -0.5659 | Yes | ||
| 48 | TSPO | 110692 3390452 | 18461 | -0.915 | -0.4074 | Yes | ||
| 49 | SSTR2 | 4590687 | 18537 | -1.366 | 0.0043 | Yes |