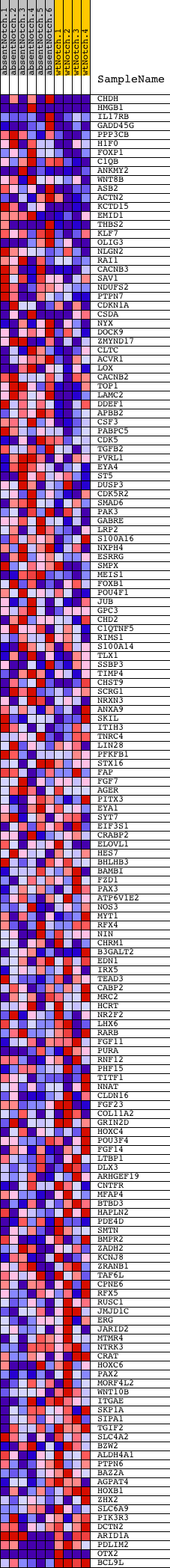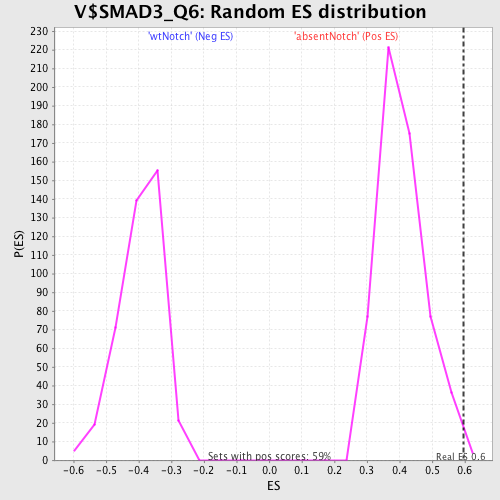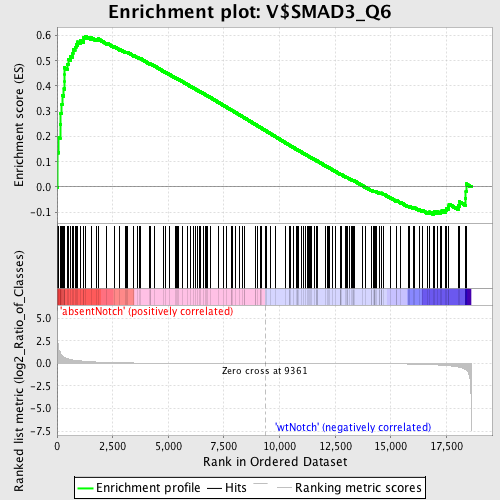
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_wtNotch.phenotype_absentNotch_versus_wtNotch.cls #absentNotch_versus_wtNotch.phenotype_absentNotch_versus_wtNotch.cls #absentNotch_versus_wtNotch_repos |
| Phenotype | phenotype_absentNotch_versus_wtNotch.cls#absentNotch_versus_wtNotch_repos |
| Upregulated in class | absentNotch |
| GeneSet | V$SMAD3_Q6 |
| Enrichment Score (ES) | 0.597551 |
| Normalized Enrichment Score (NES) | 1.4707763 |
| Nominal p-value | 0.006779661 |
| FDR q-value | 0.35890836 |
| FWER p-Value | 0.998 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CHDH | 2570711 6200338 | 11 | 3.272 | 0.1382 | Yes | ||
| 2 | HMGB1 | 2120670 2350044 | 80 | 1.465 | 0.1966 | Yes | ||
| 3 | IL17RB | 6590215 | 134 | 1.302 | 0.2490 | Yes | ||
| 4 | GADD45G | 2510142 | 166 | 1.064 | 0.2925 | Yes | ||
| 5 | PPP3CB | 6020156 | 211 | 0.891 | 0.3279 | Yes | ||
| 6 | H1F0 | 1400408 | 230 | 0.849 | 0.3629 | Yes | ||
| 7 | FOXP1 | 6510433 | 308 | 0.705 | 0.3886 | Yes | ||
| 8 | C1QB | 5910292 | 328 | 0.670 | 0.4160 | Yes | ||
| 9 | ANKMY2 | 780600 2230440 | 332 | 0.666 | 0.4441 | Yes | ||
| 10 | WNT8B | 4560301 | 336 | 0.663 | 0.4721 | Yes | ||
| 11 | ASB2 | 4760168 | 473 | 0.515 | 0.4866 | Yes | ||
| 12 | ACTN2 | 4200435 | 500 | 0.495 | 0.5062 | Yes | ||
| 13 | KCTD15 | 2810735 | 604 | 0.426 | 0.5187 | Yes | ||
| 14 | EMID1 | 3190079 5130152 | 706 | 0.376 | 0.5292 | Yes | ||
| 15 | THBS2 | 2850136 | 717 | 0.371 | 0.5444 | Yes | ||
| 16 | KLF7 | 50390 | 806 | 0.338 | 0.5539 | Yes | ||
| 17 | OLIG3 | 1050603 | 852 | 0.326 | 0.5653 | Yes | ||
| 18 | NLGN2 | 5900592 | 925 | 0.305 | 0.5744 | Yes | ||
| 19 | RAI1 | 110377 5340670 | 1039 | 0.275 | 0.5799 | Yes | ||
| 20 | CACNB3 | 2480035 4540184 | 1191 | 0.243 | 0.5820 | Yes | ||
| 21 | SAV1 | 5420541 | 1198 | 0.241 | 0.5919 | Yes | ||
| 22 | NDUFS2 | 4850020 6200402 | 1272 | 0.226 | 0.5976 | Yes | ||
| 23 | PTPN7 | 3450110 | 1539 | 0.180 | 0.5908 | No | ||
| 24 | CDKN1A | 4050088 6400706 | 1749 | 0.153 | 0.5859 | No | ||
| 25 | CSDA | 2470092 3060053 4780056 | 1843 | 0.142 | 0.5869 | No | ||
| 26 | NYX | 4120239 | 2234 | 0.106 | 0.5703 | No | ||
| 27 | DOCK9 | 5080037 | 2559 | 0.084 | 0.5563 | No | ||
| 28 | ZMYND17 | 840500 | 2824 | 0.069 | 0.5449 | No | ||
| 29 | CLTC | 6590458 | 3069 | 0.057 | 0.5341 | No | ||
| 30 | ACVR1 | 6840671 | 3108 | 0.056 | 0.5344 | No | ||
| 31 | LOX | 5340372 | 3169 | 0.053 | 0.5334 | No | ||
| 32 | CACNB2 | 1500095 7330707 | 3454 | 0.043 | 0.5199 | No | ||
| 33 | TOP1 | 770471 4060632 6650324 | 3620 | 0.039 | 0.5126 | No | ||
| 34 | LAMC2 | 450692 2680041 4200059 4670148 | 3696 | 0.037 | 0.5101 | No | ||
| 35 | DDEF1 | 1170411 4070465 | 3755 | 0.036 | 0.5084 | No | ||
| 36 | APBB2 | 5130403 | 4167 | 0.028 | 0.4874 | No | ||
| 37 | CSF3 | 2230193 6660707 | 4194 | 0.027 | 0.4871 | No | ||
| 38 | PABPC5 | 2320408 | 4209 | 0.027 | 0.4875 | No | ||
| 39 | CDK5 | 940348 | 4384 | 0.025 | 0.4791 | No | ||
| 40 | TGFB2 | 4920292 | 4792 | 0.020 | 0.4579 | No | ||
| 41 | PVRL1 | 2570167 5220519 5890500 3190309 | 4868 | 0.019 | 0.4546 | No | ||
| 42 | EYA4 | 5360286 | 5062 | 0.017 | 0.4449 | No | ||
| 43 | ST5 | 3780204 | 5299 | 0.015 | 0.4328 | No | ||
| 44 | DUSP3 | 1240129 6840215 | 5357 | 0.015 | 0.4303 | No | ||
| 45 | CDK5R2 | 3800110 | 5409 | 0.015 | 0.4282 | No | ||
| 46 | SMAD6 | 870504 | 5477 | 0.014 | 0.4251 | No | ||
| 47 | PAK3 | 4210136 | 5641 | 0.013 | 0.4169 | No | ||
| 48 | GABRE | 4780020 | 5844 | 0.012 | 0.4064 | No | ||
| 49 | LRP2 | 1400204 3290706 | 6016 | 0.011 | 0.3976 | No | ||
| 50 | S100A16 | 2510546 | 6135 | 0.010 | 0.3917 | No | ||
| 51 | NXPH4 | 3830500 | 6241 | 0.010 | 0.3864 | No | ||
| 52 | ESRRG | 4010181 | 6290 | 0.010 | 0.3842 | No | ||
| 53 | SMPX | 6590440 | 6400 | 0.009 | 0.3787 | No | ||
| 54 | MEIS1 | 1400575 | 6447 | 0.009 | 0.3766 | No | ||
| 55 | FOXB1 | 4920270 5290463 | 6562 | 0.008 | 0.3708 | No | ||
| 56 | POU4F1 | 2260315 | 6650 | 0.008 | 0.3664 | No | ||
| 57 | JUB | 5860711 5900500 | 6732 | 0.008 | 0.3623 | No | ||
| 58 | GPC3 | 6180735 | 6767 | 0.008 | 0.3608 | No | ||
| 59 | CHD2 | 4070039 | 6907 | 0.007 | 0.3536 | No | ||
| 60 | C1QTNF5 | 1690402 | 7258 | 0.006 | 0.3349 | No | ||
| 61 | RIMS1 | 5670022 | 7498 | 0.005 | 0.3221 | No | ||
| 62 | S100A14 | 450400 | 7599 | 0.005 | 0.3169 | No | ||
| 63 | TLX1 | 2630162 | 7826 | 0.004 | 0.3048 | No | ||
| 64 | SSBP3 | 870717 1050377 7040551 | 7845 | 0.004 | 0.3040 | No | ||
| 65 | TIMP4 | 6110433 | 7884 | 0.004 | 0.3021 | No | ||
| 66 | CHST9 | 4560092 | 8002 | 0.004 | 0.2960 | No | ||
| 67 | SCRG1 | 2060541 | 8217 | 0.003 | 0.2845 | No | ||
| 68 | NRXN3 | 1400546 3360471 | 8345 | 0.003 | 0.2777 | No | ||
| 69 | ANXA9 | 1190195 | 8410 | 0.003 | 0.2744 | No | ||
| 70 | SKIL | 1090364 | 8417 | 0.003 | 0.2741 | No | ||
| 71 | ITIH3 | 3830551 | 8898 | 0.001 | 0.2482 | No | ||
| 72 | TNRC4 | 4050156 | 8906 | 0.001 | 0.2478 | No | ||
| 73 | LIN28 | 6590672 | 9004 | 0.001 | 0.2426 | No | ||
| 74 | PFKFB1 | 2370128 | 9121 | 0.001 | 0.2364 | No | ||
| 75 | STX16 | 70315 | 9143 | 0.001 | 0.2353 | No | ||
| 76 | FAP | 4560019 | 9179 | 0.000 | 0.2334 | No | ||
| 77 | FGF7 | 5390484 | 9359 | 0.000 | 0.2237 | No | ||
| 78 | AGER | 5860538 5910180 | 9390 | -0.000 | 0.2221 | No | ||
| 79 | PITX3 | 3130162 | 9608 | -0.001 | 0.2103 | No | ||
| 80 | EYA1 | 1450278 5220390 | 9830 | -0.001 | 0.1984 | No | ||
| 81 | SYT7 | 2450528 3390500 3610687 | 10282 | -0.002 | 0.1741 | No | ||
| 82 | EIF3S1 | 6130368 6770044 | 10450 | -0.003 | 0.1651 | No | ||
| 83 | CRABP2 | 6940725 | 10460 | -0.003 | 0.1648 | No | ||
| 84 | ELOVL1 | 2450019 4760138 5340685 | 10489 | -0.003 | 0.1634 | No | ||
| 85 | HES7 | 4610239 | 10627 | -0.003 | 0.1561 | No | ||
| 86 | BHLHB3 | 3450438 | 10762 | -0.004 | 0.1490 | No | ||
| 87 | BAMBI | 6650463 | 10787 | -0.004 | 0.1479 | No | ||
| 88 | FZD1 | 3140215 | 10800 | -0.004 | 0.1474 | No | ||
| 89 | PAX3 | 50551 | 10837 | -0.004 | 0.1456 | No | ||
| 90 | ATP6V1E2 | 6840706 | 10967 | -0.004 | 0.1388 | No | ||
| 91 | NOS3 | 630152 670465 | 11056 | -0.005 | 0.1342 | No | ||
| 92 | MYT1 | 6770524 | 11180 | -0.005 | 0.1278 | No | ||
| 93 | RFX4 | 3710100 5340215 | 11260 | -0.005 | 0.1237 | No | ||
| 94 | NIN | 3450576 5080048 | 11301 | -0.005 | 0.1218 | No | ||
| 95 | CHRM1 | 4280619 | 11355 | -0.006 | 0.1192 | No | ||
| 96 | B3GALT2 | 430136 | 11398 | -0.006 | 0.1171 | No | ||
| 97 | EDN1 | 1770047 | 11444 | -0.006 | 0.1149 | No | ||
| 98 | IRX5 | 2630494 | 11568 | -0.006 | 0.1085 | No | ||
| 99 | TEAD3 | 2320142 | 11675 | -0.007 | 0.1031 | No | ||
| 100 | CABP2 | 840142 | 11677 | -0.007 | 0.1033 | No | ||
| 101 | MRC2 | 110369 2480520 | 11684 | -0.007 | 0.1033 | No | ||
| 102 | HCRT | 2970463 | 11709 | -0.007 | 0.1023 | No | ||
| 103 | NR2F2 | 3170609 3310577 | 12061 | -0.008 | 0.0836 | No | ||
| 104 | LHX6 | 2360731 3360397 | 12142 | -0.009 | 0.0797 | No | ||
| 105 | RARB | 430139 1410138 | 12215 | -0.009 | 0.0761 | No | ||
| 106 | FGF11 | 840292 | 12253 | -0.009 | 0.0745 | No | ||
| 107 | PURA | 3870156 | 12399 | -0.010 | 0.0671 | No | ||
| 108 | RNF12 | 6860717 | 12490 | -0.010 | 0.0626 | No | ||
| 109 | PHF15 | 6550066 6860673 | 12731 | -0.012 | 0.0501 | No | ||
| 110 | TITF1 | 3390554 4540292 | 12732 | -0.012 | 0.0506 | No | ||
| 111 | NNAT | 4920053 5130253 | 12739 | -0.012 | 0.0508 | No | ||
| 112 | CLDN16 | 1850286 | 12786 | -0.012 | 0.0488 | No | ||
| 113 | FGF23 | 510746 | 12971 | -0.013 | 0.0394 | No | ||
| 114 | COL11A2 | 5130341 | 13014 | -0.013 | 0.0377 | No | ||
| 115 | GRIN2D | 6620372 | 13030 | -0.014 | 0.0374 | No | ||
| 116 | HOXC4 | 1940193 | 13037 | -0.014 | 0.0377 | No | ||
| 117 | POU3F4 | 870274 | 13143 | -0.014 | 0.0326 | No | ||
| 118 | FGF14 | 2630390 3520075 6770048 | 13221 | -0.015 | 0.0290 | No | ||
| 119 | LTBP1 | 780035 2100746 2760402 3780551 6550184 6860193 | 13296 | -0.015 | 0.0257 | No | ||
| 120 | DLX3 | 3190497 | 13310 | -0.016 | 0.0256 | No | ||
| 121 | ARHGEF19 | 940102 | 13316 | -0.016 | 0.0260 | No | ||
| 122 | CNTFR | 4480092 6200064 | 13344 | -0.016 | 0.0252 | No | ||
| 123 | MFAP4 | 520181 | 13711 | -0.019 | 0.0062 | No | ||
| 124 | BTBD3 | 1980041 2510577 3190053 4200746 | 13856 | -0.021 | -0.0007 | No | ||
| 125 | HAPLN2 | 2900647 | 14129 | -0.024 | -0.0144 | No | ||
| 126 | PDE4D | 2470528 6660014 | 14130 | -0.024 | -0.0134 | No | ||
| 127 | SMTN | 670563 1190438 | 14218 | -0.025 | -0.0171 | No | ||
| 128 | BMPR2 | 5340066 6960328 | 14260 | -0.026 | -0.0182 | No | ||
| 129 | ZADH2 | 520438 | 14276 | -0.026 | -0.0179 | No | ||
| 130 | KCNJ8 | 870494 | 14277 | -0.026 | -0.0168 | No | ||
| 131 | ZRANB1 | 510373 4540278 | 14313 | -0.026 | -0.0176 | No | ||
| 132 | TAF6L | 1850056 | 14358 | -0.027 | -0.0188 | No | ||
| 133 | CPNE6 | 6520338 | 14480 | -0.029 | -0.0241 | No | ||
| 134 | RFX5 | 5420128 | 14500 | -0.030 | -0.0239 | No | ||
| 135 | RUSC1 | 6450093 6980575 | 14563 | -0.031 | -0.0260 | No | ||
| 136 | JMJD1C | 940575 2120025 | 14568 | -0.031 | -0.0249 | No | ||
| 137 | ERG | 50154 1770739 | 14571 | -0.031 | -0.0237 | No | ||
| 138 | JARID2 | 6290538 | 14682 | -0.033 | -0.0282 | No | ||
| 139 | MTMR4 | 4540524 | 14995 | -0.040 | -0.0434 | No | ||
| 140 | NTRK3 | 4050687 4070167 4610026 6100484 | 15238 | -0.047 | -0.0546 | No | ||
| 141 | CRAT | 540020 2060364 3520148 | 15266 | -0.048 | -0.0540 | No | ||
| 142 | HOXC6 | 3850133 4920736 | 15450 | -0.055 | -0.0616 | No | ||
| 143 | PAX2 | 6040270 7000133 | 15775 | -0.068 | -0.0763 | No | ||
| 144 | MORF4L2 | 6450133 | 15852 | -0.072 | -0.0773 | No | ||
| 145 | WNT10B | 510050 | 16005 | -0.079 | -0.0822 | No | ||
| 146 | ITGAE | 4210632 4670239 5340253 5420746 5690154 5700685 6770154 | 16044 | -0.082 | -0.0808 | No | ||
| 147 | SKP1A | 2450102 | 16293 | -0.097 | -0.0901 | No | ||
| 148 | SIPA1 | 5220687 | 16402 | -0.105 | -0.0915 | No | ||
| 149 | TGIF2 | 3840364 4050520 | 16656 | -0.123 | -0.1000 | No | ||
| 150 | SLC4A2 | 2690114 | 16738 | -0.131 | -0.0988 | No | ||
| 151 | BZW2 | 940079 | 16924 | -0.149 | -0.1026 | No | ||
| 152 | ALDH4A1 | 2450450 | 16941 | -0.150 | -0.0970 | No | ||
| 153 | PTPN6 | 4230128 | 17084 | -0.167 | -0.0976 | No | ||
| 154 | BAZ2A | 730184 | 17253 | -0.194 | -0.0985 | No | ||
| 155 | AGPAT4 | 6040079 | 17298 | -0.203 | -0.0923 | No | ||
| 156 | HOXB1 | 2120110 | 17460 | -0.230 | -0.0913 | No | ||
| 157 | ZHX2 | 2900452 | 17500 | -0.238 | -0.0833 | No | ||
| 158 | SLC6A9 | 2470520 | 17582 | -0.254 | -0.0769 | No | ||
| 159 | PIK3R3 | 1770333 | 17613 | -0.260 | -0.0675 | No | ||
| 160 | DCTN2 | 540471 3780717 | 18041 | -0.400 | -0.0736 | No | ||
| 161 | ARID1A | 2630022 1690551 4810110 | 18090 | -0.417 | -0.0586 | No | ||
| 162 | PDLIM2 | 2450603 2810551 3830040 4670164 | 18357 | -0.653 | -0.0453 | No | ||
| 163 | OTX2 | 1190471 | 18377 | -0.689 | -0.0170 | No | ||
| 164 | BCL9L | 6940053 | 18389 | -0.706 | 0.0123 | No |