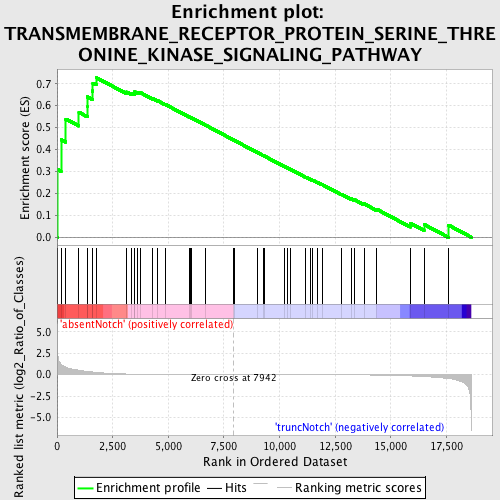
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #absentNotch_versus_truncNotch_repos |
| Phenotype | phenotype_absentNotch_versus_truncNotch.cls#absentNotch_versus_truncNotch_repos |
| Upregulated in class | absentNotch |
| GeneSet | TRANSMEMBRANE_RECEPTOR_PROTEIN_SERINE_THREONINE_KINASE_SIGNALING_PATHWAY |
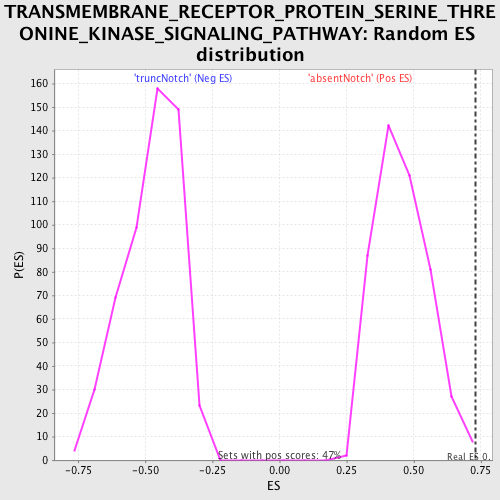
| Enrichment Score (ES) | 0.72806984 |
| Normalized Enrichment Score (NES) | 1.5972126 |
| Nominal p-value | 0.0042735045 |
| FDR q-value | 0.22729369 |
| FWER p-Value | 0.81 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CDKN1C | 6520577 | 20 | 2.517 | 0.3088 | Yes | ||
| 2 | SMAD5 | 5700497 | 179 | 1.178 | 0.4453 | Yes | ||
| 3 | SMAD4 | 5670519 | 392 | 0.852 | 0.5388 | Yes | ||
| 4 | ACVR2B | 1660546 | 981 | 0.507 | 0.5696 | Yes | ||
| 5 | LEFTY1 | 3290452 | 1361 | 0.372 | 0.5949 | Yes | ||
| 6 | MAP3K7 | 6040068 | 1368 | 0.370 | 0.6402 | Yes | ||
| 7 | SMAD2 | 4200592 | 1581 | 0.312 | 0.6672 | Yes | ||
| 8 | FNTA | 70059 | 1609 | 0.303 | 0.7031 | Yes | ||
| 9 | EID2 | 3130176 | 1757 | 0.267 | 0.7281 | Yes | ||
| 10 | ACVR1B | 3610446 5570195 | 3139 | 0.071 | 0.6624 | No | ||
| 11 | CITED2 | 5670114 5130088 | 3359 | 0.057 | 0.6576 | No | ||
| 12 | BMPR1A | 2190193 4060603 | 3492 | 0.050 | 0.6567 | No | ||
| 13 | SMURF1 | 520609 580647 | 3494 | 0.050 | 0.6628 | No | ||
| 14 | GDF9 | 2350181 | 3630 | 0.043 | 0.6608 | No | ||
| 15 | ZFYVE9 | 6980288 7100017 | 3728 | 0.038 | 0.6602 | No | ||
| 16 | CDKN2B | 6020040 | 4293 | 0.021 | 0.6325 | No | ||
| 17 | ACVR1 | 6840671 | 4525 | 0.017 | 0.6222 | No | ||
| 18 | RGMB | 670504 4570450 | 4870 | 0.013 | 0.6052 | No | ||
| 19 | TGFBR3 | 5290577 | 5937 | 0.006 | 0.5486 | No | ||
| 20 | GDF5 | 510440 | 5979 | 0.006 | 0.5471 | No | ||
| 21 | LEFTY2 | 5670364 | 6060 | 0.006 | 0.5435 | No | ||
| 22 | FMOD | 540577 | 6671 | 0.004 | 0.5111 | No | ||
| 23 | KLF10 | 4850056 | 7935 | 0.000 | 0.4431 | No | ||
| 24 | GDF10 | 4850082 | 7989 | -0.000 | 0.4403 | No | ||
| 25 | LTBP2 | 4850039 | 9022 | -0.003 | 0.3851 | No | ||
| 26 | TGFBR1 | 1400148 4280020 6550711 | 9263 | -0.003 | 0.3725 | No | ||
| 27 | SMAD7 | 430377 | 9309 | -0.003 | 0.3705 | No | ||
| 28 | ACVR2A | 6110647 | 10221 | -0.006 | 0.3223 | No | ||
| 29 | IL17F | 670577 | 10346 | -0.007 | 0.3164 | No | ||
| 30 | FOXH1 | 2060180 | 10511 | -0.007 | 0.3085 | No | ||
| 31 | SOST | 1170195 | 11164 | -0.009 | 0.2745 | No | ||
| 32 | SMAD1 | 630537 1850333 | 11379 | -0.011 | 0.2643 | No | ||
| 33 | HIPK2 | 6350647 | 11479 | -0.011 | 0.2603 | No | ||
| 34 | GDF15 | 4730017 | 11711 | -0.012 | 0.2494 | No | ||
| 35 | ACVRL1 | 3290600 | 11913 | -0.013 | 0.2402 | No | ||
| 36 | CIDEA | 4560020 | 12775 | -0.020 | 0.1964 | No | ||
| 37 | SNX6 | 6200086 | 13212 | -0.025 | 0.1760 | No | ||
| 38 | ENG | 4280270 | 13361 | -0.027 | 0.1714 | No | ||
| 39 | HPGD | 6770192 | 13799 | -0.035 | 0.1522 | No | ||
| 40 | BMPR2 | 5340066 6960328 | 14351 | -0.048 | 0.1285 | No | ||
| 41 | TGFB1 | 1940162 | 15875 | -0.140 | 0.0637 | No | ||
| 42 | STUB1 | 5860086 | 16496 | -0.227 | 0.0582 | No | ||
| 43 | SMAD3 | 6450671 | 17586 | -0.454 | 0.0555 | No |