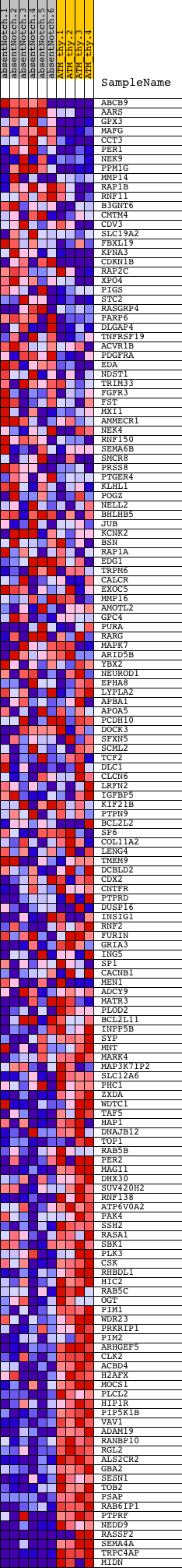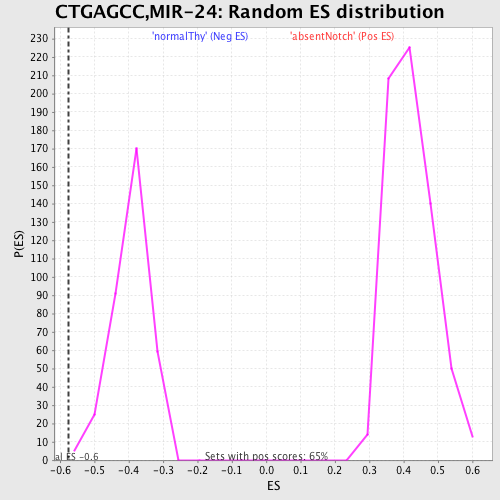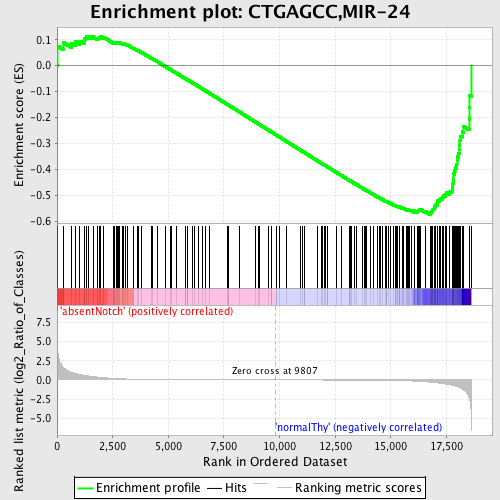
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_normalThy.phenotype_absentNotch_versus_normalThy.cls #absentNotch_versus_normalThy.phenotype_absentNotch_versus_normalThy.cls #absentNotch_versus_normalThy_repos |
| Phenotype | phenotype_absentNotch_versus_normalThy.cls#absentNotch_versus_normalThy_repos |
| Upregulated in class | normalThy |
| GeneSet | CTGAGCC,MIR-24 |
| Enrichment Score (ES) | -0.57462674 |
| Normalized Enrichment Score (NES) | -1.460563 |
| Nominal p-value | 0.0057142857 |
| FDR q-value | 0.24642874 |
| FWER p-Value | 0.932 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ABCB9 | 110603 4560592 6110338 | 18 | 4.236 | 0.0758 | No | ||
| 2 | AARS | 1570279 2370136 | 302 | 1.539 | 0.0884 | No | ||
| 3 | GPX3 | 1340450 | 662 | 0.961 | 0.0864 | No | ||
| 4 | MAFG | 4120440 | 814 | 0.833 | 0.0933 | No | ||
| 5 | CCT3 | 7100403 | 1021 | 0.701 | 0.0948 | No | ||
| 6 | PER1 | 3290315 | 1217 | 0.596 | 0.0951 | No | ||
| 7 | NEK9 | 3440402 | 1243 | 0.584 | 0.1043 | No | ||
| 8 | PPM1G | 610725 | 1298 | 0.558 | 0.1115 | No | ||
| 9 | MMP14 | 670079 | 1422 | 0.504 | 0.1140 | No | ||
| 10 | RAP1B | 3710138 | 1621 | 0.429 | 0.1110 | No | ||
| 11 | RNF11 | 3990068 5080458 | 1825 | 0.365 | 0.1067 | No | ||
| 12 | B3GNT6 | 1340736 | 1893 | 0.348 | 0.1093 | No | ||
| 13 | CMTM4 | 360600 3520332 | 1936 | 0.333 | 0.1131 | No | ||
| 14 | CDV3 | 1940577 | 2104 | 0.289 | 0.1093 | No | ||
| 15 | SLC19A2 | 6200750 | 2514 | 0.200 | 0.0908 | No | ||
| 16 | FBXL19 | 4570039 | 2587 | 0.188 | 0.0903 | No | ||
| 17 | KPNA3 | 7040088 | 2647 | 0.178 | 0.0903 | No | ||
| 18 | CDKN1B | 3800025 6450044 | 2706 | 0.167 | 0.0902 | No | ||
| 19 | RAP2C | 1690132 | 2779 | 0.157 | 0.0891 | No | ||
| 20 | XPO4 | 3840551 | 2818 | 0.152 | 0.0898 | No | ||
| 21 | PIGS | 3610142 | 2953 | 0.133 | 0.0850 | No | ||
| 22 | STC2 | 4920601 | 2999 | 0.128 | 0.0849 | No | ||
| 23 | RASGRP4 | 540102 | 3091 | 0.119 | 0.0821 | No | ||
| 24 | PARP6 | 3290332 4760167 6900181 | 3148 | 0.113 | 0.0811 | No | ||
| 25 | DLGAP4 | 4570672 5690161 | 3435 | 0.087 | 0.0672 | No | ||
| 26 | TNFRSF19 | 3870731 6940730 | 3626 | 0.074 | 0.0582 | No | ||
| 27 | ACVR1B | 3610446 5570195 | 3665 | 0.072 | 0.0575 | No | ||
| 28 | PDGFRA | 2940332 | 3806 | 0.063 | 0.0510 | No | ||
| 29 | EDA | 6420441 | 4230 | 0.045 | 0.0289 | No | ||
| 30 | NDST1 | 1500427 5340121 | 4271 | 0.043 | 0.0275 | No | ||
| 31 | TRIM33 | 580619 2230280 3990433 6200747 | 4510 | 0.036 | 0.0153 | No | ||
| 32 | FGFR3 | 5390632 6020021 | 4881 | 0.028 | -0.0042 | No | ||
| 33 | FST | 1110600 | 5094 | 0.024 | -0.0153 | No | ||
| 34 | MXI1 | 5050064 5130484 | 5146 | 0.024 | -0.0176 | No | ||
| 35 | AMMECR1 | 3800435 | 5369 | 0.021 | -0.0293 | No | ||
| 36 | NEK4 | 430731 2060010 7000333 | 5749 | 0.017 | -0.0495 | No | ||
| 37 | RNF150 | 7000500 | 5790 | 0.016 | -0.0514 | No | ||
| 38 | SEMA6B | 6770725 7100091 | 5845 | 0.016 | -0.0540 | No | ||
| 39 | SMCR8 | 730653 | 5867 | 0.016 | -0.0549 | No | ||
| 40 | PRSS8 | 3170577 | 6107 | 0.014 | -0.0676 | No | ||
| 41 | PTGER4 | 6400670 | 6193 | 0.013 | -0.0719 | No | ||
| 42 | KLHL1 | 6350021 | 6344 | 0.012 | -0.0798 | No | ||
| 43 | POGZ | 870348 7040286 | 6544 | 0.011 | -0.0904 | No | ||
| 44 | NELL2 | 2340725 4280301 5420008 | 6681 | 0.011 | -0.0976 | No | ||
| 45 | BHLHB5 | 6510520 | 6870 | 0.010 | -0.1076 | No | ||
| 46 | JUB | 5860711 5900500 | 7669 | 0.006 | -0.1507 | No | ||
| 47 | KCNK2 | 1230592 7000497 | 7693 | 0.006 | -0.1518 | No | ||
| 48 | BSN | 5270347 | 7723 | 0.006 | -0.1533 | No | ||
| 49 | RAP1A | 1090025 | 8182 | 0.005 | -0.1780 | No | ||
| 50 | EDG1 | 4200619 | 8898 | 0.003 | -0.2167 | No | ||
| 51 | TRPM6 | 2570072 | 9055 | 0.002 | -0.2251 | No | ||
| 52 | CALCR | 1690494 | 9114 | 0.002 | -0.2282 | No | ||
| 53 | EXOC5 | 7000711 | 9495 | 0.001 | -0.2488 | No | ||
| 54 | MMP16 | 2680139 | 9619 | 0.001 | -0.2554 | No | ||
| 55 | AMOTL2 | 4730082 | 9655 | 0.000 | -0.2573 | No | ||
| 56 | GPC4 | 5910408 | 9860 | -0.000 | -0.2684 | No | ||
| 57 | PURA | 3870156 | 9881 | -0.000 | -0.2694 | No | ||
| 58 | RARG | 6760136 | 10016 | -0.001 | -0.2767 | No | ||
| 59 | MAPK7 | 5670079 6100128 | 10297 | -0.001 | -0.2918 | No | ||
| 60 | ARID5B | 110403 | 10923 | -0.003 | -0.3256 | No | ||
| 61 | YBX2 | 3520441 | 11011 | -0.004 | -0.3303 | No | ||
| 62 | NEUROD1 | 3060619 | 11107 | -0.004 | -0.3354 | No | ||
| 63 | EPHA8 | 6520242 | 11137 | -0.004 | -0.3369 | No | ||
| 64 | LYPLA2 | 1500112 | 11689 | -0.006 | -0.3666 | No | ||
| 65 | APBA1 | 3140487 | 11899 | -0.007 | -0.3778 | No | ||
| 66 | APOA5 | 6130471 | 11913 | -0.007 | -0.3784 | No | ||
| 67 | PCDH10 | 940014 1190592 3190204 | 12031 | -0.007 | -0.3846 | No | ||
| 68 | DOCK3 | 3290273 3360497 2850068 | 12070 | -0.007 | -0.3865 | No | ||
| 69 | SFXN5 | 2630152 | 12143 | -0.008 | -0.3903 | No | ||
| 70 | SCML2 | 3290603 | 12153 | -0.008 | -0.3906 | No | ||
| 71 | TCF2 | 870338 5050632 | 12543 | -0.010 | -0.4115 | No | ||
| 72 | DLC1 | 1090632 6450594 | 12761 | -0.011 | -0.4231 | No | ||
| 73 | CLCN6 | 3520400 | 13136 | -0.013 | -0.4431 | No | ||
| 74 | LRFN2 | 6450411 | 13164 | -0.014 | -0.4443 | No | ||
| 75 | IGFBP5 | 2360592 | 13175 | -0.014 | -0.4446 | No | ||
| 76 | KIF21B | 2630026 6200020 6980040 | 13234 | -0.014 | -0.4475 | No | ||
| 77 | PTPN9 | 3290408 | 13357 | -0.015 | -0.4538 | No | ||
| 78 | BCL2L2 | 2760692 6770739 | 13454 | -0.016 | -0.4587 | No | ||
| 79 | SP6 | 60484 510452 2690333 | 13718 | -0.019 | -0.4727 | No | ||
| 80 | COL11A2 | 5130341 | 13798 | -0.020 | -0.4766 | No | ||
| 81 | LENG4 | 2060400 5700487 | 13853 | -0.021 | -0.4791 | No | ||
| 82 | TMEM9 | 130039 | 13896 | -0.021 | -0.4810 | No | ||
| 83 | DCBLD2 | 5890333 | 14090 | -0.024 | -0.4910 | No | ||
| 84 | CDX2 | 3850349 | 14232 | -0.027 | -0.4982 | No | ||
| 85 | CNTFR | 4480092 6200064 | 14421 | -0.030 | -0.5078 | No | ||
| 86 | PTPRD | 520008 3120097 | 14507 | -0.032 | -0.5118 | No | ||
| 87 | DUSP16 | 2370082 2480450 | 14554 | -0.034 | -0.5137 | No | ||
| 88 | INSIG1 | 1500332 3450458 4540301 | 14610 | -0.035 | -0.5160 | No | ||
| 89 | RNF2 | 2340497 | 14779 | -0.041 | -0.5244 | No | ||
| 90 | FURIN | 4120168 | 14787 | -0.042 | -0.5240 | No | ||
| 91 | GRIA3 | 2900164 6130278 6860154 | 14816 | -0.043 | -0.5248 | No | ||
| 92 | ING5 | 3830440 5910239 6200471 6650576 | 14897 | -0.046 | -0.5283 | No | ||
| 93 | SP1 | 6590017 | 14989 | -0.051 | -0.5323 | No | ||
| 94 | CACNB1 | 2940427 3710487 | 15112 | -0.059 | -0.5378 | No | ||
| 95 | MEN1 | 4850300 5900671 | 15190 | -0.064 | -0.5408 | No | ||
| 96 | ADCY9 | 5690168 | 15191 | -0.065 | -0.5396 | No | ||
| 97 | MATR3 | 1940170 5340278 | 15272 | -0.070 | -0.5427 | No | ||
| 98 | PLOD2 | 3360427 | 15320 | -0.073 | -0.5439 | No | ||
| 99 | BCL2L11 | 780044 4200601 | 15372 | -0.078 | -0.5453 | No | ||
| 100 | INPP5B | 6370088 | 15399 | -0.080 | -0.5452 | No | ||
| 101 | SYP | 2690168 | 15502 | -0.089 | -0.5491 | No | ||
| 102 | MNT | 4920575 | 15554 | -0.096 | -0.5502 | No | ||
| 103 | MARK4 | 730519 | 15698 | -0.110 | -0.5559 | No | ||
| 104 | MAP3K7IP2 | 2340242 | 15743 | -0.114 | -0.5562 | No | ||
| 105 | SLC12A6 | 610278 1980035 5340671 | 15789 | -0.119 | -0.5565 | No | ||
| 106 | PHC1 | 520195 | 15860 | -0.127 | -0.5580 | No | ||
| 107 | ZXDA | 1580332 | 15945 | -0.136 | -0.5601 | No | ||
| 108 | WDTC1 | 1690541 | 16064 | -0.153 | -0.5637 | No | ||
| 109 | TAF5 | 3450288 5890193 6860435 | 16067 | -0.153 | -0.5610 | No | ||
| 110 | HAP1 | 1240010 4230079 6980195 | 16179 | -0.169 | -0.5640 | No | ||
| 111 | DNAJB12 | 2030148 | 16191 | -0.170 | -0.5615 | No | ||
| 112 | TOP1 | 770471 4060632 6650324 | 16236 | -0.176 | -0.5607 | No | ||
| 113 | RAB5B | 130397 1980273 6900731 | 16259 | -0.179 | -0.5586 | No | ||
| 114 | PER2 | 3120204 3390593 | 16295 | -0.185 | -0.5572 | No | ||
| 115 | MAGI1 | 6510500 1990239 4670471 | 16346 | -0.194 | -0.5563 | No | ||
| 116 | DHX30 | 2480494 | 16564 | -0.240 | -0.5637 | No | ||
| 117 | SUV420H2 | 4050368 | 16766 | -0.285 | -0.5695 | Yes | ||
| 118 | RNF138 | 110142 3870129 6020441 6400193 | 16806 | -0.296 | -0.5662 | Yes | ||
| 119 | ATP6V0A2 | 4050129 | 16822 | -0.301 | -0.5616 | Yes | ||
| 120 | PAK4 | 1690152 | 16865 | -0.310 | -0.5582 | Yes | ||
| 121 | SSH2 | 360056 | 16893 | -0.318 | -0.5539 | Yes | ||
| 122 | RASA1 | 1240315 | 16954 | -0.335 | -0.5511 | Yes | ||
| 123 | SBK1 | 940452 | 16963 | -0.337 | -0.5454 | Yes | ||
| 124 | PLK3 | 2640592 | 16988 | -0.346 | -0.5405 | Yes | ||
| 125 | CSK | 6350593 | 17026 | -0.354 | -0.5360 | Yes | ||
| 126 | RHBDL1 | 1500736 | 17090 | -0.379 | -0.5326 | Yes | ||
| 127 | HIC2 | 4010433 | 17104 | -0.385 | -0.5263 | Yes | ||
| 128 | RAB5C | 1990397 | 17113 | -0.390 | -0.5197 | Yes | ||
| 129 | OGT | 2360131 4610333 | 17176 | -0.406 | -0.5157 | Yes | ||
| 130 | PIM1 | 630047 | 17249 | -0.433 | -0.5117 | Yes | ||
| 131 | WDR23 | 2030372 | 17322 | -0.463 | -0.5072 | Yes | ||
| 132 | PRKRIP1 | 4280746 6100390 | 17365 | -0.480 | -0.5008 | Yes | ||
| 133 | PIM2 | 840136 5720692 7100577 | 17467 | -0.527 | -0.4967 | Yes | ||
| 134 | ARHGEF5 | 3170605 | 17483 | -0.534 | -0.4879 | Yes | ||
| 135 | CLK2 | 510079 2630736 | 17657 | -0.627 | -0.4859 | Yes | ||
| 136 | ACBD4 | 6980079 | 17751 | -0.685 | -0.4785 | Yes | ||
| 137 | H2AFX | 3520082 | 17772 | -0.696 | -0.4670 | Yes | ||
| 138 | MOCS1 | 2120022 | 17786 | -0.704 | -0.4549 | Yes | ||
| 139 | PLCL2 | 6100575 | 17806 | -0.717 | -0.4429 | Yes | ||
| 140 | HIP1R | 3120368 | 17825 | -0.730 | -0.4307 | Yes | ||
| 141 | PIP5K1B | 3450113 6380278 | 17827 | -0.731 | -0.4175 | Yes | ||
| 142 | VAV1 | 6020487 | 17855 | -0.751 | -0.4053 | Yes | ||
| 143 | ADAM19 | 5340075 | 17886 | -0.774 | -0.3929 | Yes | ||
| 144 | RANBP10 | 2340184 | 17944 | -0.819 | -0.3811 | Yes | ||
| 145 | RGL2 | 1740017 | 17975 | -0.845 | -0.3674 | Yes | ||
| 146 | ALS2CR2 | 6450128 7100092 | 17982 | -0.853 | -0.3523 | Yes | ||
| 147 | GBA2 | 2900372 | 18044 | -0.929 | -0.3388 | Yes | ||
| 148 | SESN1 | 2120021 | 18075 | -0.963 | -0.3229 | Yes | ||
| 149 | TOB2 | 1240465 | 18083 | -0.970 | -0.3057 | Yes | ||
| 150 | PSAP | 2810338 | 18107 | -0.996 | -0.2889 | Yes | ||
| 151 | RAB6IP1 | 4150044 | 18143 | -1.040 | -0.2720 | Yes | ||
| 152 | PTPRF | 1770528 3190044 | 18211 | -1.146 | -0.2548 | Yes | ||
| 153 | NEDD9 | 1740373 4230053 | 18276 | -1.287 | -0.2349 | Yes | ||
| 154 | RASSF2 | 2480079 5720546 | 18520 | -2.347 | -0.2056 | Yes | ||
| 155 | SEMA4A | 5890022 | 18535 | -2.487 | -0.1612 | Yes | ||
| 156 | TRPC4AP | 840520 1660402 | 18552 | -2.618 | -0.1146 | Yes | ||
| 157 | MIDN | 2260132 5220377 | 18616 | -6.512 | -0.0000 | Yes |