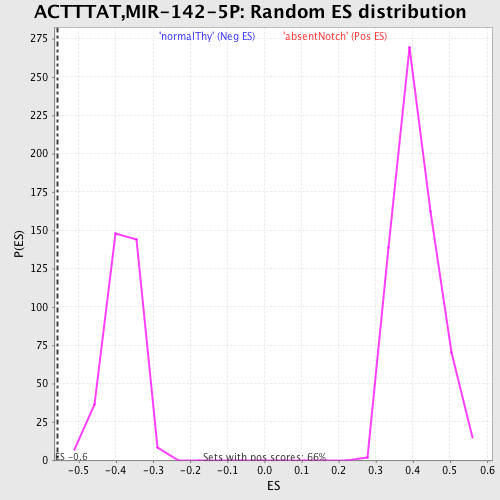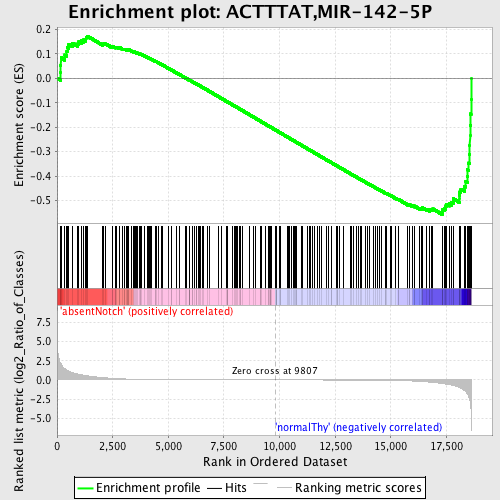
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_normalThy.phenotype_absentNotch_versus_normalThy.cls #absentNotch_versus_normalThy.phenotype_absentNotch_versus_normalThy.cls #absentNotch_versus_normalThy_repos |
| Phenotype | phenotype_absentNotch_versus_normalThy.cls#absentNotch_versus_normalThy_repos |
| Upregulated in class | normalThy |
| GeneSet | ACTTTAT,MIR-142-5P |
| Enrichment Score (ES) | -0.55650365 |
| Normalized Enrichment Score (NES) | -1.4627081 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.2558495 |
| FWER p-Value | 0.926 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | EGLN3 | 1660484 | 157 | 2.151 | 0.0230 | No | ||
| 2 | RGMA | 2450711 | 164 | 2.113 | 0.0537 | No | ||
| 3 | CBX3 | 1780592 3940008 | 174 | 2.077 | 0.0836 | No | ||
| 4 | CELSR1 | 6510348 | 350 | 1.434 | 0.0952 | No | ||
| 5 | BVES | 5340072 | 437 | 1.289 | 0.1094 | No | ||
| 6 | DNAJC7 | 4920048 | 465 | 1.225 | 0.1259 | No | ||
| 7 | GCH1 | 670364 6550358 | 525 | 1.126 | 0.1392 | No | ||
| 8 | APRIN | 2810050 | 692 | 0.931 | 0.1438 | No | ||
| 9 | RNH1 | 840438 | 935 | 0.759 | 0.1418 | No | ||
| 10 | CAPN7 | 5550050 | 978 | 0.732 | 0.1503 | No | ||
| 11 | RAB11FIP5 | 4570600 | 1104 | 0.653 | 0.1530 | No | ||
| 12 | HERPUD1 | 7100440 | 1202 | 0.607 | 0.1567 | No | ||
| 13 | ALS2 | 3130546 6400070 | 1293 | 0.561 | 0.1600 | No | ||
| 14 | PPM1G | 610725 | 1298 | 0.558 | 0.1680 | No | ||
| 15 | HOXB4 | 540131 | 1363 | 0.526 | 0.1722 | No | ||
| 16 | GRSF1 | 2100184 | 2059 | 0.303 | 0.1389 | No | ||
| 17 | BTBD1 | 5720619 | 2100 | 0.290 | 0.1410 | No | ||
| 18 | PAIP1 | 3870576 | 2185 | 0.268 | 0.1403 | No | ||
| 19 | H3F3A | 2900086 | 2485 | 0.208 | 0.1271 | No | ||
| 20 | ATXN7L2 | 4010161 | 2486 | 0.207 | 0.1302 | No | ||
| 21 | TGFBR2 | 1780711 1980537 6550398 | 2610 | 0.183 | 0.1262 | No | ||
| 22 | IL18BP | 3450128 | 2632 | 0.179 | 0.1277 | No | ||
| 23 | TRIM36 | 1400600 1980082 5130037 | 2685 | 0.172 | 0.1274 | No | ||
| 24 | MBIP | 2640524 5570441 | 2792 | 0.155 | 0.1239 | No | ||
| 25 | IGF1 | 1990193 3130377 3290280 | 2798 | 0.153 | 0.1259 | No | ||
| 26 | TPBG | 630161 | 2956 | 0.133 | 0.1193 | No | ||
| 27 | PBX3 | 1300424 3710577 6180575 | 3026 | 0.126 | 0.1174 | No | ||
| 28 | CPEB2 | 4760338 | 3042 | 0.124 | 0.1184 | No | ||
| 29 | VPS54 | 1660168 | 3112 | 0.116 | 0.1163 | No | ||
| 30 | FBXL3 | 5910142 | 3154 | 0.112 | 0.1157 | No | ||
| 31 | STC1 | 360161 | 3185 | 0.109 | 0.1157 | No | ||
| 32 | CAMK2A | 1740333 1940112 4150292 | 3217 | 0.106 | 0.1156 | No | ||
| 33 | TRIP10 | 460438 | 3224 | 0.105 | 0.1168 | No | ||
| 34 | ABCG4 | 3440458 | 3350 | 0.094 | 0.1114 | No | ||
| 35 | MGAT4B | 130053 6840100 | 3430 | 0.087 | 0.1084 | No | ||
| 36 | CUL4A | 1170088 2320008 2470278 | 3457 | 0.085 | 0.1082 | No | ||
| 37 | TCF12 | 3610324 7000156 | 3512 | 0.081 | 0.1065 | No | ||
| 38 | RHOT1 | 6760692 | 3586 | 0.077 | 0.1036 | No | ||
| 39 | KIT | 7040095 | 3619 | 0.075 | 0.1030 | No | ||
| 40 | SLC36A1 | 5080242 | 3688 | 0.070 | 0.1003 | No | ||
| 41 | SCOC | 610048 2230053 | 3742 | 0.067 | 0.0984 | No | ||
| 42 | EGR3 | 6940128 | 3745 | 0.067 | 0.0993 | No | ||
| 43 | ARID4B | 730754 6660026 | 3777 | 0.065 | 0.0986 | No | ||
| 44 | UBE4A | 6100520 | 3910 | 0.058 | 0.0923 | No | ||
| 45 | REV3L | 1090717 | 4078 | 0.051 | 0.0839 | No | ||
| 46 | CD69 | 380167 4730088 | 4094 | 0.050 | 0.0839 | No | ||
| 47 | RHOC | 2060242 | 4130 | 0.048 | 0.0827 | No | ||
| 48 | ZFX | 5900400 | 4176 | 0.047 | 0.0809 | No | ||
| 49 | STAG1 | 1190300 1400722 4590100 | 4260 | 0.044 | 0.0770 | No | ||
| 50 | SPOCK2 | 6650369 | 4415 | 0.039 | 0.0692 | No | ||
| 51 | SNX16 | 870446 | 4427 | 0.038 | 0.0692 | No | ||
| 52 | FZD7 | 7050706 | 4453 | 0.038 | 0.0684 | No | ||
| 53 | ST8SIA2 | 2680082 | 4545 | 0.035 | 0.0640 | No | ||
| 54 | NRG1 | 1050332 | 4694 | 0.032 | 0.0564 | No | ||
| 55 | NFAT5 | 2510411 5890195 6550152 | 4733 | 0.031 | 0.0548 | No | ||
| 56 | ETV1 | 70735 2940603 5080463 | 5007 | 0.026 | 0.0403 | No | ||
| 57 | CUL2 | 4200278 | 5136 | 0.024 | 0.0337 | No | ||
| 58 | ARID2 | 110647 5390435 | 5163 | 0.023 | 0.0326 | No | ||
| 59 | RTN1 | 4150452 4850176 | 5345 | 0.021 | 0.0231 | No | ||
| 60 | HIAT1 | 1580176 | 5483 | 0.019 | 0.0160 | No | ||
| 61 | LUZP2 | 60673 | 5486 | 0.019 | 0.0161 | No | ||
| 62 | HMGCLL1 | 2570091 | 5750 | 0.017 | 0.0021 | No | ||
| 63 | ZCCHC14 | 5900288 | 5811 | 0.016 | -0.0009 | No | ||
| 64 | LRP2 | 1400204 3290706 | 5929 | 0.015 | -0.0071 | No | ||
| 65 | FNDC3A | 2690097 3290044 | 5935 | 0.015 | -0.0071 | No | ||
| 66 | BAI3 | 940692 | 5943 | 0.015 | -0.0073 | No | ||
| 67 | ZFPM2 | 460072 | 5951 | 0.015 | -0.0074 | No | ||
| 68 | ALCAM | 1050019 | 6077 | 0.014 | -0.0140 | No | ||
| 69 | EYA4 | 5360286 | 6098 | 0.014 | -0.0149 | No | ||
| 70 | MSH6 | 4480064 6520093 | 6156 | 0.014 | -0.0178 | No | ||
| 71 | UBR1 | 3800132 | 6250 | 0.013 | -0.0226 | No | ||
| 72 | QKI | 5220093 6130044 | 6276 | 0.013 | -0.0238 | No | ||
| 73 | ADAMTS5 | 5890592 | 6349 | 0.012 | -0.0275 | No | ||
| 74 | FXYD3 | 2320450 | 6384 | 0.012 | -0.0292 | No | ||
| 75 | UCHL3 | 1240735 | 6456 | 0.012 | -0.0329 | No | ||
| 76 | PAPOLB | 3840450 | 6529 | 0.011 | -0.0366 | No | ||
| 77 | ZCCHC11 | 3390731 5360039 | 6595 | 0.011 | -0.0400 | No | ||
| 78 | RPS25 | 520139 | 6740 | 0.010 | -0.0477 | No | ||
| 79 | MASTL | 380433 4480215 5390301 | 6838 | 0.010 | -0.0528 | No | ||
| 80 | FIGN | 7000601 | 7259 | 0.008 | -0.0755 | No | ||
| 81 | HDLBP | 2350692 6590520 | 7378 | 0.007 | -0.0818 | No | ||
| 82 | UNC84A | 1450632 3520408 | 7629 | 0.007 | -0.0953 | No | ||
| 83 | HSPA8 | 2120315 1050132 6550746 | 7647 | 0.007 | -0.0961 | No | ||
| 84 | RCN2 | 840324 | 7657 | 0.006 | -0.0965 | No | ||
| 85 | SLC17A7 | 7000037 | 7672 | 0.006 | -0.0972 | No | ||
| 86 | NFE2L2 | 2810619 3390162 | 7884 | 0.006 | -0.1086 | No | ||
| 87 | KLF11 | 6840739 | 7975 | 0.005 | -0.1134 | No | ||
| 88 | SLC4A4 | 1780075 1850040 4810068 | 8010 | 0.005 | -0.1151 | No | ||
| 89 | NPAS4 | 2320204 4150154 | 8057 | 0.005 | -0.1176 | No | ||
| 90 | LHX8 | 5700347 | 8124 | 0.005 | -0.1211 | No | ||
| 91 | PTPN4 | 1410047 | 8193 | 0.005 | -0.1247 | No | ||
| 92 | BTG3 | 7050079 | 8249 | 0.004 | -0.1276 | No | ||
| 93 | TAOK2 | 110575 6380692 | 8258 | 0.004 | -0.1280 | No | ||
| 94 | STK35 | 2360121 | 8338 | 0.004 | -0.1322 | No | ||
| 95 | KLF10 | 4850056 | 8632 | 0.003 | -0.1481 | No | ||
| 96 | HOXA13 | 3120647 | 8843 | 0.003 | -0.1595 | No | ||
| 97 | GPR88 | 2450435 | 8918 | 0.002 | -0.1634 | No | ||
| 98 | BTBD7 | 730288 | 9131 | 0.002 | -0.1749 | No | ||
| 99 | MBD2 | 5900546 | 9145 | 0.002 | -0.1756 | No | ||
| 100 | MMP24 | 5340154 | 9204 | 0.002 | -0.1787 | No | ||
| 101 | SLC18A2 | 6420541 | 9359 | 0.001 | -0.1871 | No | ||
| 102 | UBE2D1 | 1850400 | 9493 | 0.001 | -0.1943 | No | ||
| 103 | EXOC5 | 7000711 | 9495 | 0.001 | -0.1943 | No | ||
| 104 | EPAS1 | 5290156 | 9554 | 0.001 | -0.1975 | No | ||
| 105 | MYF5 | 1050739 | 9573 | 0.001 | -0.1984 | No | ||
| 106 | PRPF4B | 940181 2570300 | 9595 | 0.001 | -0.1996 | No | ||
| 107 | CCNT2 | 2100390 4150563 | 9605 | 0.001 | -0.2001 | No | ||
| 108 | RERG | 5910324 | 9656 | 0.000 | -0.2028 | No | ||
| 109 | DCHS1 | 4850487 | 9798 | 0.000 | -0.2104 | No | ||
| 110 | WDR26 | 1090138 | 9865 | -0.000 | -0.2140 | No | ||
| 111 | PURA | 3870156 | 9881 | -0.000 | -0.2148 | No | ||
| 112 | ABCA1 | 6290156 | 9882 | -0.000 | -0.2148 | No | ||
| 113 | BHLHB3 | 3450438 | 10001 | -0.001 | -0.2212 | No | ||
| 114 | MEA1 | 2230347 6350338 | 10035 | -0.001 | -0.2230 | No | ||
| 115 | SHANK2 | 1500452 | 10059 | -0.001 | -0.2242 | No | ||
| 116 | SREBF1 | 4780333 | 10333 | -0.002 | -0.2390 | No | ||
| 117 | NPAT | 3800594 4670026 | 10414 | -0.002 | -0.2434 | No | ||
| 118 | GARNL1 | 5670398 | 10439 | -0.002 | -0.2446 | No | ||
| 119 | MKLN1 | 70504 1850129 | 10536 | -0.002 | -0.2498 | No | ||
| 120 | COL4A3 | 5910075 | 10617 | -0.002 | -0.2541 | No | ||
| 121 | DIO2 | 380504 540403 | 10682 | -0.002 | -0.2576 | No | ||
| 122 | ROBO1 | 3710398 | 10713 | -0.003 | -0.2592 | No | ||
| 123 | LRP1B | 1450193 | 10754 | -0.003 | -0.2613 | No | ||
| 124 | BMP2 | 6620687 | 10779 | -0.003 | -0.2626 | No | ||
| 125 | RHOQ | 520161 | 11000 | -0.003 | -0.2745 | No | ||
| 126 | SIX4 | 6760022 | 11017 | -0.004 | -0.2753 | No | ||
| 127 | OTX2 | 1190471 | 11235 | -0.004 | -0.2870 | No | ||
| 128 | CRSP2 | 1940059 | 11355 | -0.005 | -0.2934 | No | ||
| 129 | MAT2B | 1690139 2510706 | 11381 | -0.005 | -0.2947 | No | ||
| 130 | CNNM4 | 1740347 | 11456 | -0.005 | -0.2986 | No | ||
| 131 | ADAMTS1 | 3140286 5860463 | 11558 | -0.005 | -0.3040 | No | ||
| 132 | SON | 3120148 5860239 7040403 | 11712 | -0.006 | -0.3123 | No | ||
| 133 | GRM7 | 4230110 | 11793 | -0.006 | -0.3165 | No | ||
| 134 | KCNJ3 | 50184 | 11871 | -0.007 | -0.3206 | No | ||
| 135 | SYT7 | 2450528 3390500 3610687 | 12128 | -0.008 | -0.3344 | No | ||
| 136 | RNF128 | 1230300 5670273 | 12182 | -0.008 | -0.3372 | No | ||
| 137 | ELAVL4 | 50735 3360086 5220167 | 12316 | -0.009 | -0.3443 | No | ||
| 138 | PAPPA | 4230463 | 12352 | -0.009 | -0.3460 | No | ||
| 139 | ELAVL2 | 360181 | 12551 | -0.010 | -0.3566 | No | ||
| 140 | AHR | 4560671 | 12618 | -0.010 | -0.3601 | No | ||
| 141 | TRIM3 | 5570156 | 12623 | -0.010 | -0.3602 | No | ||
| 142 | KCNRG | 3460315 3940072 | 12677 | -0.010 | -0.3629 | No | ||
| 143 | CPEB3 | 3940164 | 12861 | -0.011 | -0.3727 | No | ||
| 144 | GOPC | 6620400 | 13181 | -0.014 | -0.3898 | No | ||
| 145 | SOX5 | 2370576 2900167 3190128 5050528 | 13250 | -0.014 | -0.3933 | No | ||
| 146 | MYO1D | 6760292 | 13303 | -0.015 | -0.3959 | No | ||
| 147 | PLEKHA3 | 5080068 | 13436 | -0.016 | -0.4028 | No | ||
| 148 | GAS7 | 2120358 6450494 | 13545 | -0.017 | -0.4084 | No | ||
| 149 | DDHD1 | 3450241 3940523 6380156 | 13620 | -0.018 | -0.4122 | No | ||
| 150 | LMX1A | 2810239 5690324 | 13678 | -0.018 | -0.4150 | No | ||
| 151 | KITLG | 2120047 6220300 | 13878 | -0.021 | -0.4255 | No | ||
| 152 | VANGL1 | 380162 | 13952 | -0.022 | -0.4292 | No | ||
| 153 | FOS | 1850315 | 14022 | -0.023 | -0.4326 | No | ||
| 154 | CBLN4 | 4010451 | 14206 | -0.026 | -0.4421 | No | ||
| 155 | RIPK5 | 1230647 1340114 | 14303 | -0.028 | -0.4469 | No | ||
| 156 | LRP12 | 2370112 | 14416 | -0.030 | -0.4526 | No | ||
| 157 | RAB6B | 5860121 | 14477 | -0.032 | -0.4554 | No | ||
| 158 | PCBP2 | 5690048 | 14601 | -0.035 | -0.4615 | No | ||
| 159 | SLC25A27 | 6200707 6940019 | 14757 | -0.040 | -0.4694 | No | ||
| 160 | JMJD1C | 940575 2120025 | 14762 | -0.040 | -0.4690 | No | ||
| 161 | SRI | 3390446 4850064 | 14820 | -0.043 | -0.4715 | No | ||
| 162 | NTRK3 | 4050687 4070167 4610026 6100484 | 14992 | -0.052 | -0.4800 | No | ||
| 163 | CHD9 | 430037 2570129 5420398 | 15012 | -0.053 | -0.4802 | No | ||
| 164 | SP2 | 6400026 | 15213 | -0.066 | -0.4901 | No | ||
| 165 | PCTK2 | 5700673 | 15323 | -0.073 | -0.4950 | No | ||
| 166 | PLXNA2 | 6450132 | 15343 | -0.075 | -0.4949 | No | ||
| 167 | RPS6KA5 | 2120563 7040546 | 15771 | -0.117 | -0.5164 | No | ||
| 168 | HN1 | 3360156 | 15821 | -0.123 | -0.5173 | No | ||
| 169 | HIPK1 | 110193 | 15832 | -0.125 | -0.5160 | No | ||
| 170 | OSR1 | 1500025 | 15952 | -0.137 | -0.5204 | No | ||
| 171 | RPS6KA4 | 2190524 | 16057 | -0.151 | -0.5239 | No | ||
| 172 | AP1G1 | 2360671 | 16058 | -0.151 | -0.5216 | No | ||
| 173 | MGLL | 2030446 4210520 | 16297 | -0.185 | -0.5318 | No | ||
| 174 | ACIN1 | 3390278 6350377 6620142 | 16374 | -0.199 | -0.5330 | No | ||
| 175 | PCBP1 | 1050088 | 16401 | -0.205 | -0.5315 | No | ||
| 176 | CRK | 1230162 4780128 | 16406 | -0.206 | -0.5287 | No | ||
| 177 | ZFYVE21 | 1940056 | 16614 | -0.251 | -0.5362 | No | ||
| 178 | TAF10 | 2760020 | 16743 | -0.279 | -0.5391 | No | ||
| 179 | MED28 | 3420025 | 16748 | -0.280 | -0.5352 | No | ||
| 180 | APBB3 | 6020168 | 16834 | -0.303 | -0.5354 | No | ||
| 181 | SSH2 | 360056 | 16893 | -0.318 | -0.5339 | No | ||
| 182 | CDK5 | 940348 | 17311 | -0.458 | -0.5498 | Yes | ||
| 183 | CCNG2 | 3190095 | 17333 | -0.468 | -0.5441 | Yes | ||
| 184 | BECN1 | 3840433 | 17336 | -0.470 | -0.5373 | Yes | ||
| 185 | ACTN4 | 3840301 4590390 7050132 | 17404 | -0.498 | -0.5336 | Yes | ||
| 186 | UBE2A | 4010563 | 17457 | -0.522 | -0.5288 | Yes | ||
| 187 | ZFAND3 | 2760504 | 17471 | -0.529 | -0.5217 | Yes | ||
| 188 | ATG16L1 | 5390292 | 17489 | -0.535 | -0.5148 | Yes | ||
| 189 | BAZ2A | 730184 | 17621 | -0.608 | -0.5130 | Yes | ||
| 190 | ULK1 | 6100315 | 17720 | -0.669 | -0.5085 | Yes | ||
| 191 | FOXJ2 | 2450441 | 17804 | -0.717 | -0.5025 | Yes | ||
| 192 | DUSP2 | 2370184 | 17814 | -0.722 | -0.4924 | Yes | ||
| 193 | ATP1B1 | 3130594 | 18074 | -0.962 | -0.4924 | Yes | ||
| 194 | SPEN | 2060041 | 18077 | -0.965 | -0.4783 | Yes | ||
| 195 | UBE3C | 3140673 | 18089 | -0.976 | -0.4646 | Yes | ||
| 196 | CIC | 6370161 | 18149 | -1.044 | -0.4525 | Yes | ||
| 197 | MCL1 | 1660672 | 18315 | -1.362 | -0.4415 | Yes | ||
| 198 | LRRIQ2 | 3990064 6370673 6380372 | 18370 | -1.543 | -0.4218 | Yes | ||
| 199 | SDCBP | 460487 2190039 5270441 | 18439 | -1.771 | -0.3995 | Yes | ||
| 200 | CLCN3 | 510195 780176 1660093 3830372 | 18450 | -1.787 | -0.3739 | Yes | ||
| 201 | BNIP2 | 1410475 6770088 | 18487 | -2.026 | -0.3461 | Yes | ||
| 202 | FGF13 | 630575 1570440 5360121 | 18542 | -2.538 | -0.3118 | Yes | ||
| 203 | TRPC4AP | 840520 1660402 | 18552 | -2.618 | -0.2739 | Yes | ||
| 204 | TLE4 | 1450064 1500519 | 18558 | -2.750 | -0.2339 | Yes | ||
| 205 | ZFP36 | 2030605 | 18563 | -2.788 | -0.1932 | Yes | ||
| 206 | SRF | 70139 2320039 | 18588 | -3.326 | -0.1458 | Yes | ||
| 207 | BTG1 | 4200735 6040131 6200133 | 18607 | -4.199 | -0.0852 | Yes | ||
| 208 | RHOA | 580142 5900131 5340450 | 18613 | -5.837 | 0.0002 | Yes |