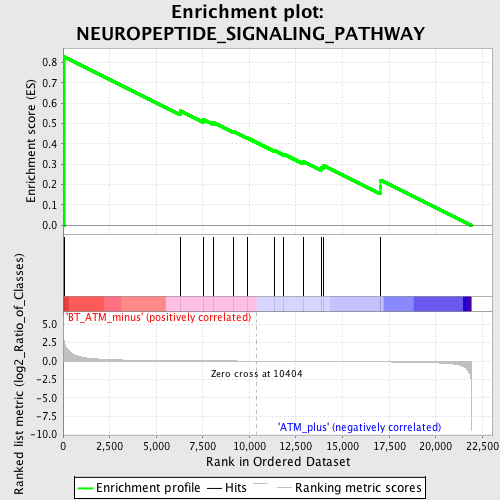
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | BT_ATM_minus |
| GeneSet | NEUROPEPTIDE_SIGNALING_PATHWAY |
| Enrichment Score (ES) | 0.8273029 |
| Normalized Enrichment Score (NES) | 1.6197841 |
| Nominal p-value | 0.011257036 |
| FDR q-value | 0.5049072 |
| FWER p-Value | 0.999 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | AGRP | 1421690_s_at 1457362_at | 66 | 2.575 | 0.8273 | Yes | ||
| 2 | GLRA2 | 1434098_at 1443351_at | 6313 | 0.060 | 0.5617 | No | ||
| 3 | GALR3 | 1450581_at | 7539 | 0.040 | 0.5186 | No | ||
| 4 | ECEL1 | 1422586_at | 8079 | 0.032 | 0.5042 | No | ||
| 5 | NPFF | 1420606_at | 9133 | 0.017 | 0.4617 | No | ||
| 6 | NMUR1 | 1421667_at | 9894 | 0.007 | 0.4292 | No | ||
| 7 | GRP | 1424525_at | 11334 | -0.012 | 0.3675 | No | ||
| 8 | NMUR2 | 1425943_at | 11852 | -0.019 | 0.3499 | No | ||
| 9 | GLRB | 1422504_at 1459850_x_at | 12896 | -0.033 | 0.3129 | No | ||
| 10 | GLRA1 | 1422277_at 1437139_at | 13849 | -0.047 | 0.2845 | No | ||
| 11 | GALR1 | 1427704_a_at 1441329_at 1450568_at | 13977 | -0.048 | 0.2943 | No | ||
| 12 | HCRTR2 | 1444223_at | 17031 | -0.106 | 0.1892 | No | ||
| 13 | HCRTR1 | 1436295_at | 17062 | -0.107 | 0.2222 | No |

