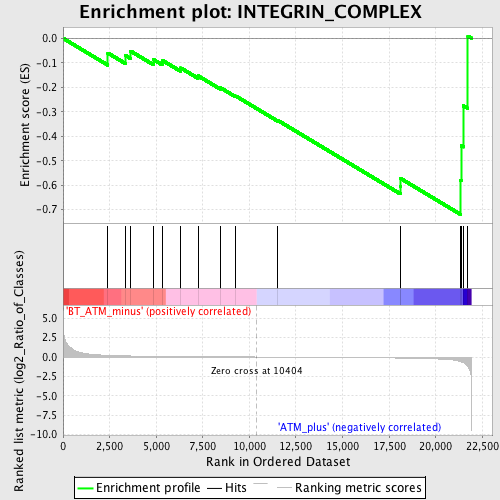
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | INTEGRIN_COMPLEX |
| Enrichment Score (ES) | -0.7195633 |
| Normalized Enrichment Score (NES) | -1.6188139 |
| Nominal p-value | 0.018604651 |
| FDR q-value | 0.7378261 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ITGB4 | 1427387_a_at | 2403 | 0.212 | -0.0601 | No | ||
| 2 | ITGA5 | 1423267_s_at 1423268_at 1457561_at 1458996_at | 3362 | 0.142 | -0.0707 | No | ||
| 3 | ITGA9 | 1420860_at 1450029_s_at 1460285_at | 3617 | 0.130 | -0.0518 | No | ||
| 4 | ITGAM | 1422046_at | 4849 | 0.090 | -0.0869 | No | ||
| 5 | ITGA10 | 1440235_at | 5319 | 0.080 | -0.0896 | No | ||
| 6 | ITGA1 | 1439713_at 1446569_at 1455251_at | 6318 | 0.060 | -0.1211 | No | ||
| 7 | ITGB6 | 1422983_at 1432281_a_at | 7245 | 0.044 | -0.1531 | No | ||
| 8 | ITGA3 | 1421997_s_at 1455158_at 1460305_at | 8439 | 0.026 | -0.2014 | No | ||
| 9 | ITGA8 | 1427489_at 1454966_at | 9269 | 0.015 | -0.2356 | No | ||
| 10 | ITGA2 | 1450501_at | 11529 | -0.015 | -0.3353 | No | ||
| 11 | ITGB8 | 1436223_at | 18103 | -0.136 | -0.6033 | Yes | ||
| 12 | ITGAX | 1419128_at | 18125 | -0.137 | -0.5724 | Yes | ||
| 13 | ITGAE | 1447541_s_at 1449216_at | 21352 | -0.591 | -0.5814 | Yes | ||
| 14 | MYH9 | 1417472_at 1420170_at 1420171_s_at 1420172_at 1440708_at | 21376 | -0.614 | -0.4389 | Yes | ||
| 15 | ITGB7 | 1418741_at | 21488 | -0.721 | -0.2754 | Yes | ||
| 16 | ITGB3 | 1421511_at 1455257_at | 21738 | -1.266 | 0.0089 | Yes |

