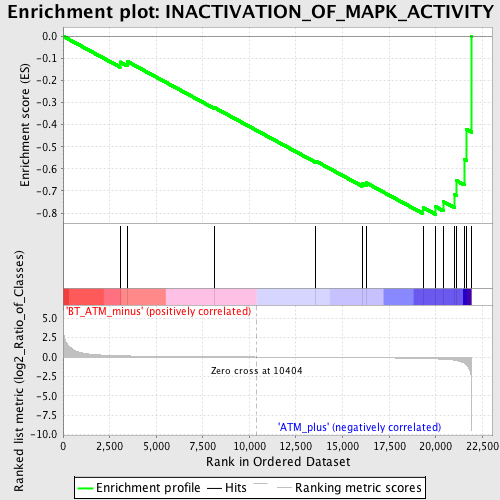
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | INACTIVATION_OF_MAPK_ACTIVITY |
| Enrichment Score (ES) | -0.80640185 |
| Normalized Enrichment Score (NES) | -1.7425275 |
| Nominal p-value | 0.014084507 |
| FDR q-value | 0.60492796 |
| FWER p-Value | 0.95 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MBIP | 1424405_at | 3056 | 0.158 | -0.1163 | No | ||
| 2 | SPRED2 | 1423001_at 1434403_at 1436892_at 1441415_at 1441802_at | 3444 | 0.138 | -0.1138 | No | ||
| 3 | DUSP8 | 1418714_at | 8136 | 0.031 | -0.3233 | No | ||
| 4 | GPS1 | 1415699_a_at | 13574 | -0.043 | -0.5651 | No | ||
| 5 | DUSP22 | 1441723_at 1448985_at | 16051 | -0.083 | -0.6658 | No | ||
| 6 | GPS2 | 1417796_at | 16282 | -0.088 | -0.6634 | No | ||
| 7 | SPRED1 | 1423160_at 1423161_s_at 1423162_s_at 1428777_at 1441517_at 1442859_at 1443652_x_at 1445228_at 1452911_at 1460116_s_at | 19326 | -0.187 | -0.7750 | Yes | ||
| 8 | RGS4 | 1416286_at 1416287_at 1448285_at | 20016 | -0.238 | -0.7717 | Yes | ||
| 9 | LAX1 | 1438687_at | 20411 | -0.284 | -0.7480 | Yes | ||
| 10 | DUSP2 | 1450698_at | 21013 | -0.410 | -0.7155 | Yes | ||
| 11 | DUSP9 | 1433844_a_at 1433845_x_at 1454737_at | 21142 | -0.460 | -0.6541 | Yes | ||
| 12 | DUSP6 | 1415834_at | 21545 | -0.791 | -0.5568 | Yes | ||
| 13 | DUSP16 | 1418401_a_at 1440615_at 1440651_at | 21647 | -0.958 | -0.4212 | Yes | ||
| 14 | RGS3 | 1425701_a_at 1427648_at 1449516_a_at 1454026_a_at | 21919 | -2.970 | 0.0007 | Yes |

