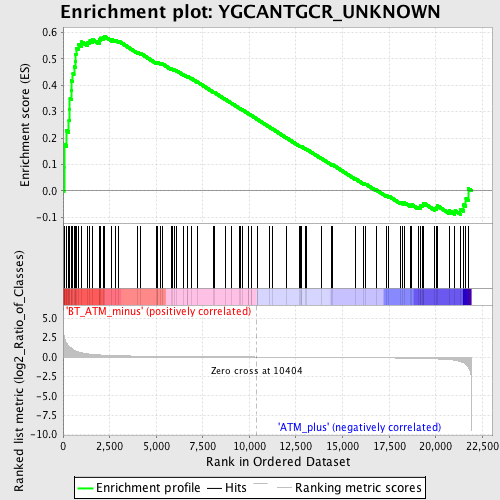
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | BT_ATM_minus |
| GeneSet | YGCANTGCR_UNKNOWN |
| Enrichment Score (ES) | 0.58650655 |
| Normalized Enrichment Score (NES) | 1.5888746 |
| Nominal p-value | 0.00729927 |
| FDR q-value | 0.25445706 |
| FWER p-Value | 0.796 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | EWSR1 | 1417238_at 1425919_at 1436884_x_at 1441074_at 1456461_at | 48 | 2.776 | 0.0874 | Yes | ||
| 2 | RTN3 | 1418101_a_at 1425539_a_at 1442365_at 1443220_at | 50 | 2.724 | 0.1752 | Yes | ||
| 3 | CLTC | 1419407_at 1439094_at 1440457_at 1454626_at | 179 | 1.818 | 0.2280 | Yes | ||
| 4 | DDX6 | 1424598_at 1457618_at | 314 | 1.424 | 0.2678 | Yes | ||
| 5 | VSNL1 | 1420955_at 1450055_at | 362 | 1.289 | 0.3073 | Yes | ||
| 6 | MAT2A | 1423667_at 1433576_at 1438386_x_at 1438630_x_at 1438976_x_at 1439386_x_at 1456702_x_at | 368 | 1.284 | 0.3485 | Yes | ||
| 7 | SYNJ1 | 1436333_a_at 1436334_at 1439672_at 1439997_at 1454961_at | 439 | 1.139 | 0.3820 | Yes | ||
| 8 | USP8 | 1422883_at 1443420_at 1446807_at 1447386_at | 457 | 1.110 | 0.4170 | Yes | ||
| 9 | GANAB | 1415787_at 1437812_x_at | 528 | 0.955 | 0.4446 | Yes | ||
| 10 | TOPORS | 1417754_at 1417755_at 1448838_at | 586 | 0.880 | 0.4704 | Yes | ||
| 11 | KPNA1 | 1419548_at 1449503_at 1449504_at 1460260_s_at | 664 | 0.783 | 0.4922 | Yes | ||
| 12 | PHEX | 1421979_at 1450445_at | 681 | 0.768 | 0.5162 | Yes | ||
| 13 | EIF4G2 | 1415863_at 1452758_s_at | 696 | 0.753 | 0.5399 | Yes | ||
| 14 | SRGAP2 | 1427967_at 1429884_at 1434406_at 1434407_at | 812 | 0.651 | 0.5556 | Yes | ||
| 15 | CALM3 | 1426710_at 1438825_at 1438826_x_at 1450864_at | 998 | 0.538 | 0.5645 | Yes | ||
| 16 | FBXO11 | 1434108_at | 1325 | 0.401 | 0.5625 | Yes | ||
| 17 | CENTG2 | 1435432_at 1435433_at 1437394_at 1442632_at 1457978_at 1459053_at 1459389_at | 1441 | 0.367 | 0.5691 | Yes | ||
| 18 | SEMA7A | 1422040_at 1459903_at | 1583 | 0.335 | 0.5735 | Yes | ||
| 19 | MAP2K7 | 1421416_at 1425393_a_at 1425512_at 1425513_at 1440442_at 1451736_a_at 1457182_at | 1939 | 0.269 | 0.5659 | Yes | ||
| 20 | IGSF4D | 1435145_at 1435146_s_at 1435147_x_at 1436743_at 1440412_at 1444664_at 1459761_x_at | 1964 | 0.266 | 0.5734 | Yes | ||
| 21 | NONO | 1415820_x_at 1431239_at 1448103_s_at | 2016 | 0.259 | 0.5794 | Yes | ||
| 22 | KCNS3 | 1445519_at 1457799_at | 2151 | 0.240 | 0.5810 | Yes | ||
| 23 | BMPR2 | 1419616_at 1434310_at 1441652_at | 2199 | 0.236 | 0.5865 | Yes | ||
| 24 | MLL5 | 1427236_a_at 1432601_at 1434704_at 1439107_a_at 1439108_at | 2613 | 0.190 | 0.5737 | No | ||
| 25 | FGF13 | 1418497_at 1418498_at | 2812 | 0.174 | 0.5703 | No | ||
| 26 | MBTPS2 | 1436883_at 1437495_at | 2995 | 0.162 | 0.5672 | No | ||
| 27 | APP | 1420621_a_at 1427442_a_at 1438373_at 1438374_x_at 1444705_at 1447347_at 1458525_at | 3980 | 0.116 | 0.5259 | No | ||
| 28 | STX16 | 1429237_at | 4158 | 0.110 | 0.5213 | No | ||
| 29 | MAGED1 | 1450062_a_at | 5026 | 0.086 | 0.4844 | No | ||
| 30 | CAST | 1426098_a_at 1435972_at 1451413_at | 5064 | 0.085 | 0.4855 | No | ||
| 31 | GSK3A | 1435638_at | 5076 | 0.085 | 0.4877 | No | ||
| 32 | PCM1 | 1418524_at 1418525_at 1431287_at 1436908_at 1438528_at 1446140_at 1449120_a_at | 5247 | 0.082 | 0.4825 | No | ||
| 33 | CALM2 | 1422414_a_at 1423807_a_at | 5336 | 0.080 | 0.4811 | No | ||
| 34 | SLC39A6 | 1424674_at 1424675_at 1441949_x_at 1447893_x_at | 5795 | 0.070 | 0.4624 | No | ||
| 35 | CTCF | 1418330_at 1449042_at | 5882 | 0.068 | 0.4606 | No | ||
| 36 | CAMK1D | 1426389_at 1438643_at 1452050_at | 5981 | 0.066 | 0.4583 | No | ||
| 37 | TRIM9 | 1434595_at 1443989_at | 6088 | 0.065 | 0.4555 | No | ||
| 38 | SLC9A3R2 | 1428954_at 1428955_x_at 1431208_a_at 1439368_a_at 1439369_x_at 1452976_a_at | 6460 | 0.057 | 0.4404 | No | ||
| 39 | CHD2 | 1444246_at 1445843_at | 6687 | 0.054 | 0.4318 | No | ||
| 40 | PAQR9 | 1455025_at | 6696 | 0.054 | 0.4331 | No | ||
| 41 | AHI1 | 1439083_at 1455177_at | 6900 | 0.050 | 0.4254 | No | ||
| 42 | IL1RAPL1 | 1445821_at 1458964_at | 7191 | 0.045 | 0.4136 | No | ||
| 43 | ACCN1 | 1417994_a_at | 8066 | 0.032 | 0.3746 | No | ||
| 44 | ADNP | 1423069_at | 8145 | 0.031 | 0.3721 | No | ||
| 45 | OLFML3 | 1448475_at | 8703 | 0.023 | 0.3473 | No | ||
| 46 | NOL5A | 1426533_at 1455035_s_at | 9040 | 0.018 | 0.3325 | No | ||
| 47 | HGF | 1425379_at 1442884_at 1451866_a_at | 9457 | 0.013 | 0.3139 | No | ||
| 48 | EIF5A | 1437859_x_at 1451470_s_at | 9520 | 0.012 | 0.3114 | No | ||
| 49 | ZDHHC15 | 1442274_at | 9621 | 0.011 | 0.3072 | No | ||
| 50 | PURA | 1420628_at 1438219_at 1449934_at 1453783_at 1456898_at | 9944 | 0.006 | 0.2926 | No | ||
| 51 | TMEM14A | 1428447_at | 9963 | 0.006 | 0.2920 | No | ||
| 52 | PFTK1 | 1419249_at 1419250_a_at 1441692_at 1445984_at 1453956_a_at | 10104 | 0.004 | 0.2857 | No | ||
| 53 | SFRS9 | 1417727_at | 10127 | 0.004 | 0.2848 | No | ||
| 54 | SYT12 | 1422878_at | 10413 | -0.000 | 0.2718 | No | ||
| 55 | GAB2 | 1419829_a_at 1420785_at 1439786_at 1446860_at | 10440 | -0.000 | 0.2706 | No | ||
| 56 | PSAP | 1415687_a_at 1421813_a_at | 11097 | -0.009 | 0.2408 | No | ||
| 57 | FGFR1 | 1424050_s_at 1425911_a_at 1436551_at | 11255 | -0.011 | 0.2340 | No | ||
| 58 | VAMP1 | 1421862_a_at 1421863_at | 11978 | -0.020 | 0.2016 | No | ||
| 59 | NLGN3 | 1456384_at | 12703 | -0.030 | 0.1694 | No | ||
| 60 | LRFN5 | 1441502_at 1454687_at | 12732 | -0.030 | 0.1691 | No | ||
| 61 | SFXN5 | 1436618_at 1440428_at | 12798 | -0.031 | 0.1672 | No | ||
| 62 | ASCL1 | 1437086_at 1450164_at | 12802 | -0.031 | 0.1680 | No | ||
| 63 | EPHB1 | 1455188_at | 13030 | -0.034 | 0.1588 | No | ||
| 64 | ELOVL4 | 1424306_at 1451308_at | 13069 | -0.035 | 0.1581 | No | ||
| 65 | HMGCLL1 | 1434913_at | 13878 | -0.047 | 0.1227 | No | ||
| 66 | DGKD | 1433564_at 1439873_at 1442254_at 1454655_at | 14432 | -0.056 | 0.0992 | No | ||
| 67 | MIR16 | 1418444_a_at | 14461 | -0.056 | 0.0997 | No | ||
| 68 | ARHGAP5 | 1423194_at 1450896_at 1450897_at 1457410_at | 15698 | -0.077 | 0.0456 | No | ||
| 69 | LSM11 | 1453755_at | 16154 | -0.085 | 0.0275 | No | ||
| 70 | MGEA5 | 1422900_at 1422901_at 1422902_s_at 1433654_at 1439081_at | 16257 | -0.087 | 0.0257 | No | ||
| 71 | SARS | 1426257_a_at 1452000_s_at | 16811 | -0.100 | 0.0036 | No | ||
| 72 | BAI2 | 1435558_at | 17356 | -0.114 | -0.0177 | No | ||
| 73 | GSPT1 | 1426736_at 1426737_at 1435000_at 1446550_at 1452168_x_at 1455173_at | 17499 | -0.118 | -0.0203 | No | ||
| 74 | CPNE1 | 1433439_at | 18111 | -0.136 | -0.0439 | No | ||
| 75 | PNMA3 | 1456829_at | 18243 | -0.140 | -0.0454 | No | ||
| 76 | UVRAG | 1454706_at | 18349 | -0.143 | -0.0456 | No | ||
| 77 | PDCL3 | 1424002_at | 18677 | -0.156 | -0.0555 | No | ||
| 78 | HS3ST3B1 | 1421331_at 1421332_at 1433977_at 1442816_at 1444055_at | 18730 | -0.158 | -0.0528 | No | ||
| 79 | CLSTN1 | 1421860_at 1421861_at 1445085_at | 19073 | -0.173 | -0.0629 | No | ||
| 80 | MNT | 1418192_at | 19190 | -0.179 | -0.0624 | No | ||
| 81 | NRXN1 | 1428240_at 1433413_at 1439358_a_at 1439359_x_at 1442501_at 1447216_at 1454691_at 1460540_at | 19201 | -0.179 | -0.0571 | No | ||
| 82 | STATIP1 | 1415774_at 1438179_s_at | 19291 | -0.184 | -0.0552 | No | ||
| 83 | NRXN3 | 1419825_at 1431153_at 1432931_at 1433788_at 1438193_at 1439629_at 1442423_at 1444700_at 1456137_at 1460101_at | 19334 | -0.187 | -0.0511 | No | ||
| 84 | KLC2 | 1418214_at 1431487_at | 19357 | -0.188 | -0.0460 | No | ||
| 85 | GBF1 | 1415709_s_at 1415757_at 1432677_at 1438207_at | 19945 | -0.231 | -0.0655 | No | ||
| 86 | KCNA1 | 1417416_at 1437230_at 1442413_at 1455785_at | 20050 | -0.241 | -0.0625 | No | ||
| 87 | NNAT | 1423506_a_at | 20106 | -0.248 | -0.0570 | No | ||
| 88 | KCNN2 | 1445676_at 1448927_at | 20739 | -0.334 | -0.0751 | No | ||
| 89 | ANXA6 | 1415818_at 1429246_a_at 1429247_at 1442288_at | 21043 | -0.422 | -0.0754 | No | ||
| 90 | TACC2 | 1425745_a_at 1447891_at | 21354 | -0.592 | -0.0705 | No | ||
| 91 | FREQ | 1434887_at 1450146_at 1460293_at | 21496 | -0.731 | -0.0533 | No | ||
| 92 | ASCL2 | 1422396_s_at 1432018_at 1460514_s_at | 21633 | -0.920 | -0.0298 | No | ||
| 93 | PLEKHA1 | 1417965_at 1443111_at | 21759 | -1.350 | 0.0080 | No |

