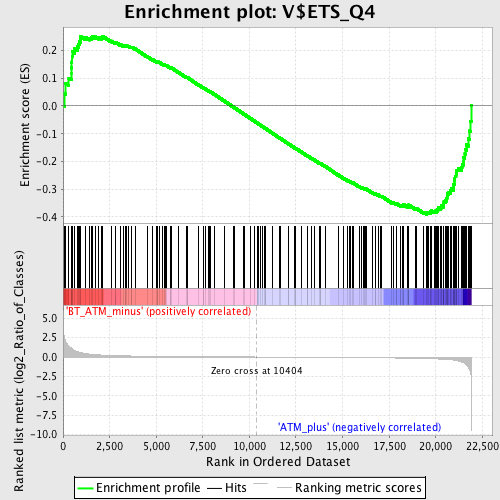
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | V$ETS_Q4 |
| Enrichment Score (ES) | -0.39116922 |
| Normalized Enrichment Score (NES) | -1.3096462 |
| Nominal p-value | 0.034632035 |
| FDR q-value | 0.68496335 |
| FWER p-Value | 0.999 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SRPR | 1423670_a_at 1427570_at | 78 | 2.503 | 0.0444 | No | ||
| 2 | DPP3 | 1431594_at 1448215_a_at | 124 | 2.065 | 0.0820 | No | ||
| 3 | SOST | 1421245_at 1436240_at 1450179_at | 312 | 1.424 | 0.1007 | No | ||
| 4 | SCN3A | 1421705_at 1439204_at | 446 | 1.131 | 0.1163 | No | ||
| 5 | SEC61A1 | 1416189_a_at 1416190_a_at 1416191_at 1434986_a_at 1448242_at 1455987_at | 450 | 1.121 | 0.1377 | No | ||
| 6 | ITPR2 | 1421678_at 1424833_at 1424834_s_at 1427287_s_at 1427693_at 1444418_at 1444551_at 1447328_at | 468 | 1.072 | 0.1575 | No | ||
| 7 | PPP1R16B | 1425414_at 1455080_at | 477 | 1.045 | 0.1772 | No | ||
| 8 | ARHGAP4 | 1419296_at 1443360_x_at 1458190_at | 485 | 1.037 | 0.1968 | No | ||
| 9 | ERG | 1425370_a_at 1440244_at | 631 | 0.826 | 0.2060 | No | ||
| 10 | UTX | 1427235_at 1427672_a_at 1440524_at 1441201_at 1445198_at 1446234_at 1446278_at | 761 | 0.691 | 0.2133 | No | ||
| 11 | FOXP1 | 1421140_a_at 1421141_a_at 1421142_s_at 1435222_at 1438802_at 1443258_at 1446280_at 1447209_at 1455242_at 1456851_at | 815 | 0.649 | 0.2233 | No | ||
| 12 | FOXO3A | 1434831_a_at 1434832_at 1439598_at 1444226_at 1450236_at 1458279_at | 884 | 0.600 | 0.2317 | No | ||
| 13 | ETF1 | 1420023_at 1420024_s_at 1424013_at 1440629_at 1451208_at | 914 | 0.584 | 0.2416 | No | ||
| 14 | CNOT10 | 1433882_at 1453654_at 1459756_at 1459757_x_at | 940 | 0.569 | 0.2513 | No | ||
| 15 | ARHGAP15 | 1429881_at 1435959_at 1441650_at | 1190 | 0.457 | 0.2487 | No | ||
| 16 | ZFP91 | 1426326_at 1429615_at 1429616_at 1459444_at | 1425 | 0.371 | 0.2450 | No | ||
| 17 | RAP2C | 1428119_a_at 1437016_x_at 1460430_at | 1511 | 0.351 | 0.2479 | No | ||
| 18 | CHIC2 | 1419659_s_at 1429050_at 1445911_at | 1585 | 0.335 | 0.2509 | No | ||
| 19 | EXTL2 | 1422538_at 1422539_at | 1738 | 0.307 | 0.2498 | No | ||
| 20 | SEC24C | 1430250_at 1452147_at | 1921 | 0.272 | 0.2467 | No | ||
| 21 | NASP | 1416042_s_at 1416043_at 1440328_at 1442728_at | 2058 | 0.252 | 0.2453 | No | ||
| 22 | ERCC1 | 1417328_at 1437447_s_at 1441838_at 1445266_at | 2104 | 0.247 | 0.2479 | No | ||
| 23 | RBM4 | 1416920_at 1437322_at 1455517_at | 2120 | 0.245 | 0.2520 | No | ||
| 24 | TCERG1 | 1421033_a_at 1434434_s_at 1450100_a_at | 2584 | 0.192 | 0.2344 | No | ||
| 25 | DPPA4 | 1429597_at | 2793 | 0.175 | 0.2282 | No | ||
| 26 | POU3F4 | 1422164_at 1422165_at | 2811 | 0.174 | 0.2307 | No | ||
| 27 | HMGA1 | 1416184_s_at | 3099 | 0.155 | 0.2205 | No | ||
| 28 | BAZ2A | 1427209_at 1427210_at 1438192_s_at | 3215 | 0.149 | 0.2181 | No | ||
| 29 | CHES1 | 1434002_at 1436925_at 1438255_at 1440607_at 1444555_at 1447325_at 1453094_at | 3324 | 0.144 | 0.2159 | No | ||
| 30 | MGAT4A | 1444347_at | 3345 | 0.142 | 0.2177 | No | ||
| 31 | ITPR1 | 1417279_at 1443492_at 1457189_at 1460203_at | 3416 | 0.139 | 0.2171 | No | ||
| 32 | DOCK11 | 1429028_at 1443467_at | 3512 | 0.135 | 0.2154 | No | ||
| 33 | CRADD | 1427505_a_at 1443430_at | 3645 | 0.129 | 0.2118 | No | ||
| 34 | CSAD | 1427981_a_at | 3690 | 0.127 | 0.2122 | No | ||
| 35 | SPIB | 1460407_at | 3869 | 0.120 | 0.2063 | No | ||
| 36 | USP3 | 1425022_at 1425023_at 1435531_at 1441056_at 1446886_at | 4519 | 0.099 | 0.1784 | No | ||
| 37 | SOX14 | 1427609_at | 4808 | 0.092 | 0.1669 | No | ||
| 38 | GMFG | 1419193_a_at 1419194_s_at | 4994 | 0.087 | 0.1601 | No | ||
| 39 | CAST | 1426098_a_at 1435972_at 1451413_at | 5064 | 0.085 | 0.1586 | No | ||
| 40 | CANX | 1415692_s_at 1422845_at 1428935_at | 5074 | 0.085 | 0.1598 | No | ||
| 41 | NEDD4 | 1421955_a_at 1444044_at 1450431_a_at 1451109_a_at | 5155 | 0.084 | 0.1577 | No | ||
| 42 | CALM2 | 1422414_a_at 1423807_a_at | 5336 | 0.080 | 0.1510 | No | ||
| 43 | AMD1 | 1448484_at | 5430 | 0.077 | 0.1482 | No | ||
| 44 | GJA5 | 1422192_at 1429101_at | 5486 | 0.076 | 0.1471 | No | ||
| 45 | CPNE8 | 1430520_at 1430521_s_at 1431146_a_at | 5550 | 0.075 | 0.1457 | No | ||
| 46 | RIN1 | 1424507_at 1437460_x_at | 5778 | 0.070 | 0.1366 | No | ||
| 47 | MARK1 | 1419834_x_at 1425509_at 1425510_at 1425511_at 1449630_s_at | 5788 | 0.070 | 0.1375 | No | ||
| 48 | HOXC4 | 1422870_at | 5840 | 0.069 | 0.1365 | No | ||
| 49 | FGFR2 | 1420847_a_at 1433489_s_at 1443996_at | 6213 | 0.062 | 0.1206 | No | ||
| 50 | ARSB | 1429190_at 1458281_at | 6612 | 0.055 | 0.1033 | No | ||
| 51 | PPP1R9B | 1447247_at 1447486_at | 6657 | 0.054 | 0.1024 | No | ||
| 52 | CHD2 | 1444246_at 1445843_at | 6687 | 0.054 | 0.1021 | No | ||
| 53 | PAX6 | 1419271_at 1425960_s_at 1437816_at 1452526_a_at 1456342_at | 7256 | 0.044 | 0.0768 | No | ||
| 54 | LIF | 1421206_at 1421207_at 1450160_at | 7545 | 0.039 | 0.0643 | No | ||
| 55 | NME2 | 1448808_a_at | 7547 | 0.039 | 0.0650 | No | ||
| 56 | TEAD3 | 1420593_a_at | 7657 | 0.038 | 0.0607 | No | ||
| 57 | KCNAB2 | 1416956_at 1425885_a_at | 7803 | 0.036 | 0.0548 | No | ||
| 58 | AGPAT1 | 1421024_at 1421025_at | 7844 | 0.035 | 0.0536 | No | ||
| 59 | DDX23 | 1430050_at | 7913 | 0.034 | 0.0511 | No | ||
| 60 | CD2BP2 | 1417223_at 1417224_a_at 1448624_at | 8128 | 0.031 | 0.0419 | No | ||
| 61 | CORO1A | 1416246_a_at 1435288_at 1455269_a_at | 8643 | 0.024 | 0.0187 | No | ||
| 62 | HOXA1 | 1420565_at | 9131 | 0.017 | -0.0033 | No | ||
| 63 | NR1D1 | 1421802_at 1426464_at | 9215 | 0.016 | -0.0068 | No | ||
| 64 | TIMP4 | 1423405_at 1450974_at | 9660 | 0.010 | -0.0271 | No | ||
| 65 | STARD13 | 1452604_at | 9757 | 0.009 | -0.0313 | No | ||
| 66 | EPN3 | 1424960_at | 10061 | 0.004 | -0.0452 | No | ||
| 67 | NDUFA4 | 1424085_at | 10281 | 0.002 | -0.0552 | No | ||
| 68 | SEMA4C | 1433920_at | 10425 | -0.000 | -0.0617 | No | ||
| 69 | GAB2 | 1419829_a_at 1420785_at 1439786_at 1446860_at | 10440 | -0.000 | -0.0624 | No | ||
| 70 | TRIM41 | 1424275_s_at 1440268_at | 10487 | -0.001 | -0.0645 | No | ||
| 71 | RPL41 | 1422623_x_at 1455578_x_at | 10596 | -0.002 | -0.0694 | No | ||
| 72 | RHOV | 1424976_at | 10710 | -0.004 | -0.0745 | No | ||
| 73 | POF1B | 1427492_at | 10802 | -0.005 | -0.0786 | No | ||
| 74 | PPP1R14C | 1417701_at 1443799_at 1446671_at | 10856 | -0.006 | -0.0809 | No | ||
| 75 | MMP3 | 1418945_at | 10883 | -0.006 | -0.0820 | No | ||
| 76 | SLC9A3R1 | 1438115_a_at 1438116_x_at 1450982_at | 11226 | -0.011 | -0.0975 | No | ||
| 77 | ZNF35 | 1460252_s_at | 11599 | -0.016 | -0.1143 | No | ||
| 78 | TWIST1 | 1418733_at | 11661 | -0.016 | -0.1168 | No | ||
| 79 | CENTB1 | 1434873_a_at 1434874_x_at 1437117_at 1455125_at | 12108 | -0.022 | -0.1369 | No | ||
| 80 | UXT | 1418986_a_at | 12125 | -0.022 | -0.1372 | No | ||
| 81 | ELK3 | 1417662_at 1442986_at 1445886_at 1448797_at | 12442 | -0.026 | -0.1512 | No | ||
| 82 | IRAK4 | 1421670_a_at 1451749_at | 12465 | -0.027 | -0.1517 | No | ||
| 83 | TPP2 | 1421893_a_at 1421894_a_at 1430575_a_at 1430576_at 1438264_a_at | 12501 | -0.027 | -0.1528 | No | ||
| 84 | E2F5 | 1417444_at 1447625_at 1460207_s_at | 12774 | -0.031 | -0.1647 | No | ||
| 85 | GFI1 | 1417679_at | 13112 | -0.035 | -0.1795 | No | ||
| 86 | TENC1 | 1452264_at | 13128 | -0.036 | -0.1795 | No | ||
| 87 | SOX2 | 1416967_at | 13341 | -0.039 | -0.1885 | No | ||
| 88 | NDUFS2 | 1451096_at | 13485 | -0.041 | -0.1943 | No | ||
| 89 | HOXA10 | 1431475_a_at 1446408_at | 13767 | -0.045 | -0.2063 | No | ||
| 90 | PLCB2 | 1452481_at | 13797 | -0.046 | -0.2068 | No | ||
| 91 | DRP2 | 1440452_at 1441224_at 1441542_at | 13831 | -0.046 | -0.2074 | No | ||
| 92 | CRSP7 | 1452282_at 1458377_at | 14099 | -0.050 | -0.2187 | No | ||
| 93 | KLF13 | 1432543_a_at 1456030_at | 14771 | -0.061 | -0.2484 | No | ||
| 94 | ACTR3 | 1426392_a_at 1434968_a_at 1452051_at | 15065 | -0.066 | -0.2606 | No | ||
| 95 | CENTD2 | 1428064_at | 15282 | -0.069 | -0.2692 | No | ||
| 96 | TBN | 1416450_at 1416451_s_at | 15385 | -0.071 | -0.2725 | No | ||
| 97 | RPS3 | 1435151_a_at 1447563_at 1455600_at | 15455 | -0.073 | -0.2743 | No | ||
| 98 | TGFB3 | 1417455_at | 15532 | -0.074 | -0.2764 | No | ||
| 99 | VCAM1 | 1415989_at 1436003_at 1448162_at 1451314_a_at | 15580 | -0.075 | -0.2771 | No | ||
| 100 | KIF3B | 1420987_at 1450073_at 1450074_at | 15892 | -0.081 | -0.2898 | No | ||
| 101 | LTBR | 1416435_at | 16045 | -0.083 | -0.2952 | No | ||
| 102 | DDIT3 | 1417516_at 1443897_at | 16124 | -0.085 | -0.2972 | No | ||
| 103 | FURIN | 1418518_at | 16201 | -0.086 | -0.2990 | No | ||
| 104 | ADAMTS4 | 1452595_at | 16226 | -0.087 | -0.2984 | No | ||
| 105 | RGS14 | 1419221_a_at | 16278 | -0.088 | -0.2991 | No | ||
| 106 | BLNK | 1451780_at | 16631 | -0.096 | -0.3134 | No | ||
| 107 | LRFN4 | 1454700_at | 16755 | -0.099 | -0.3172 | No | ||
| 108 | PDGFB | 1450413_at 1450414_at | 16775 | -0.099 | -0.3162 | No | ||
| 109 | GGTL3 | 1451094_at | 16912 | -0.103 | -0.3204 | No | ||
| 110 | ADCY4 | 1418098_at | 17055 | -0.107 | -0.3249 | No | ||
| 111 | LSM1 | 1423873_at 1441452_at | 17104 | -0.108 | -0.3251 | No | ||
| 112 | PDAP1 | 1434019_at 1434020_at | 17650 | -0.123 | -0.3478 | No | ||
| 113 | SEPT1 | 1449898_at | 17727 | -0.124 | -0.3489 | No | ||
| 114 | CENTG1 | 1419789_at | 17899 | -0.130 | -0.3542 | No | ||
| 115 | IL13 | 1420802_at | 17902 | -0.130 | -0.3518 | No | ||
| 116 | AP1G2 | 1419113_at | 18133 | -0.137 | -0.3598 | No | ||
| 117 | DMTF1 | 1422636_at 1440349_at 1456697_x_at | 18135 | -0.137 | -0.3572 | No | ||
| 118 | STRBP | 1421125_at 1442465_s_at 1442622_at 1444001_at 1452061_s_at 1457478_at | 18207 | -0.139 | -0.3578 | No | ||
| 119 | HYAL2 | 1448679_at | 18291 | -0.142 | -0.3589 | No | ||
| 120 | MAP4K2 | 1428297_at 1434833_at 1450244_a_at | 18294 | -0.142 | -0.3563 | No | ||
| 121 | ACSL5 | 1428082_at | 18484 | -0.148 | -0.3621 | No | ||
| 122 | NEDD8 | 1423715_a_at 1440996_at 1445811_at | 18540 | -0.151 | -0.3618 | No | ||
| 123 | BIN3 | 1417691_at | 18550 | -0.151 | -0.3593 | No | ||
| 124 | BAG4 | 1418707_at 1439024_at 1449186_at | 18561 | -0.152 | -0.3568 | No | ||
| 125 | RBMS2 | 1423540_at 1433979_at | 18926 | -0.166 | -0.3704 | No | ||
| 126 | CCL2 | 1420380_at | 18955 | -0.168 | -0.3684 | No | ||
| 127 | VAV1 | 1422932_a_at | 19344 | -0.187 | -0.3827 | No | ||
| 128 | DAB2IP | 1433558_at 1441170_a_at | 19530 | -0.199 | -0.3874 | Yes | ||
| 129 | TNFSF11 | 1419083_at 1451944_a_at | 19561 | -0.201 | -0.3849 | Yes | ||
| 130 | PTPN11 | 1421196_at 1427699_a_at 1451225_at | 19624 | -0.205 | -0.3838 | Yes | ||
| 131 | POLD4 | 1427885_at | 19721 | -0.213 | -0.3841 | Yes | ||
| 132 | HCLS1 | 1418842_at | 19728 | -0.213 | -0.3803 | Yes | ||
| 133 | ACSL4 | 1418911_s_at 1433531_at 1451828_a_at | 19757 | -0.214 | -0.3775 | Yes | ||
| 134 | GATA3 | 1448886_at | 19920 | -0.230 | -0.3805 | Yes | ||
| 135 | PRKAG1 | 1417690_at 1433533_x_at 1457803_at | 19974 | -0.233 | -0.3785 | Yes | ||
| 136 | STAT5B | 1422102_a_at 1422103_a_at | 20068 | -0.243 | -0.3781 | Yes | ||
| 137 | SIGIRR | 1449163_at | 20086 | -0.245 | -0.3742 | Yes | ||
| 138 | PTK2 | 1423059_at 1430827_a_at 1439198_at 1440082_at 1441475_at 1443384_at 1445137_at | 20132 | -0.251 | -0.3714 | Yes | ||
| 139 | CAP1 | 1417461_at 1417462_at | 20136 | -0.252 | -0.3667 | Yes | ||
| 140 | LCP1 | 1415983_at 1448160_at | 20278 | -0.267 | -0.3681 | Yes | ||
| 141 | FCHO1 | 1428575_at 1436077_a_at 1436078_at | 20296 | -0.268 | -0.3637 | Yes | ||
| 142 | ARPC1B | 1416226_at | 20309 | -0.271 | -0.3591 | Yes | ||
| 143 | FXYD5 | 1418296_at | 20425 | -0.287 | -0.3589 | Yes | ||
| 144 | MADD | 1425735_at 1455502_at | 20444 | -0.289 | -0.3542 | Yes | ||
| 145 | PIK3R4 | 1460352_s_at | 20450 | -0.290 | -0.3488 | Yes | ||
| 146 | ETV5 | 1420998_at 1428142_at 1450082_s_at | 20451 | -0.290 | -0.3433 | Yes | ||
| 147 | PRKCBP1 | 1426614_at 1429415_at 1445827_at 1456143_at 1458617_at | 20521 | -0.299 | -0.3407 | Yes | ||
| 148 | RIN3 | 1434684_at 1442530_at | 20579 | -0.309 | -0.3374 | Yes | ||
| 149 | DGKA | 1418578_at | 20600 | -0.313 | -0.3323 | Yes | ||
| 150 | ST7L | 1427916_at 1456363_at | 20617 | -0.316 | -0.3270 | Yes | ||
| 151 | SLC41A1 | 1444511_at 1460565_at | 20620 | -0.316 | -0.3210 | Yes | ||
| 152 | HCST | 1419119_at | 20631 | -0.317 | -0.3154 | Yes | ||
| 153 | SLC30A7 | 1450697_at | 20700 | -0.329 | -0.3122 | Yes | ||
| 154 | ARRB2 | 1426239_s_at 1451987_at | 20802 | -0.348 | -0.3102 | Yes | ||
| 155 | E2F3 | 1427462_at 1434564_at | 20809 | -0.350 | -0.3037 | Yes | ||
| 156 | UBE2L3 | 1417907_at 1417908_s_at 1448879_at 1448880_at | 20832 | -0.357 | -0.2979 | Yes | ||
| 157 | RAE1 | 1429528_at 1444256_at 1459822_at | 20952 | -0.389 | -0.2959 | Yes | ||
| 158 | DGKZ | 1426738_at 1452169_a_at | 20974 | -0.396 | -0.2893 | Yes | ||
| 159 | GMPR2 | 1416356_at 1429180_at | 20976 | -0.396 | -0.2817 | Yes | ||
| 160 | JUNB | 1415899_at | 21007 | -0.408 | -0.2753 | Yes | ||
| 161 | PDLIM2 | 1423946_at | 21008 | -0.409 | -0.2674 | Yes | ||
| 162 | CD79A | 1418830_at | 21039 | -0.421 | -0.2607 | Yes | ||
| 163 | PML | 1448757_at 1456103_at 1459137_at | 21044 | -0.423 | -0.2528 | Yes | ||
| 164 | LCP2 | 1418641_at 1418642_at | 21099 | -0.441 | -0.2468 | Yes | ||
| 165 | HM13 | 1417287_at 1428855_at 1428856_at 1438456_at 1438865_at 1455631_at | 21124 | -0.453 | -0.2392 | Yes | ||
| 166 | LPXN | 1424965_at | 21133 | -0.457 | -0.2308 | Yes | ||
| 167 | IKBKB | 1426207_at 1426333_a_at 1432275_at 1445141_at 1454184_a_at | 21209 | -0.491 | -0.2249 | Yes | ||
| 168 | TREML2 | 1419837_at 1440298_at 1440938_at 1444718_at | 21393 | -0.627 | -0.2212 | Yes | ||
| 169 | GPR150 | 1442261_at 1447485_at | 21423 | -0.656 | -0.2100 | Yes | ||
| 170 | ITGB7 | 1418741_at | 21488 | -0.721 | -0.1991 | Yes | ||
| 171 | CAPZA1 | 1439455_x_at 1440380_at | 21495 | -0.731 | -0.1853 | Yes | ||
| 172 | SLC9A9 | 1433719_at 1437488_at 1447149_at | 21579 | -0.842 | -0.1730 | Yes | ||
| 173 | RNF12 | 1417250_at 1418730_at 1418731_at 1439403_x_at 1440850_at 1443167_at | 21625 | -0.911 | -0.1576 | Yes | ||
| 174 | ELMO1 | 1424523_at 1446610_at 1450208_a_at | 21665 | -1.016 | -0.1399 | Yes | ||
| 175 | PCDH7 | 1427592_at 1437442_at 1449249_at 1456214_at | 21768 | -1.369 | -0.1183 | Yes | ||
| 176 | TCF12 | 1421908_a_at 1427670_a_at 1438762_at 1439209_at 1439619_at 1443012_at 1444620_at 1445093_at 1445133_at 1457524_at 1458337_at | 21810 | -1.573 | -0.0900 | Yes | ||
| 177 | KLF12 | 1439846_at 1439847_s_at 1441040_at 1455521_at | 21889 | -2.012 | -0.0550 | Yes | ||
| 178 | RGS3 | 1425701_a_at 1427648_at 1449516_a_at 1454026_a_at | 21919 | -2.970 | 0.0007 | Yes |

