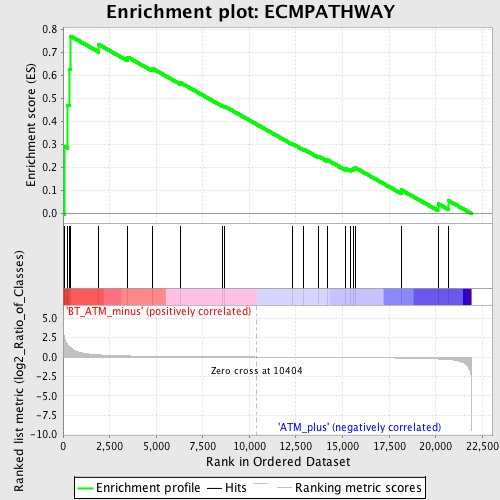
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | BT_ATM_minus |
| GeneSet | ECMPATHWAY |
| Enrichment Score (ES) | 0.7710982 |
| Normalized Enrichment Score (NES) | 1.6572752 |
| Nominal p-value | 0.0088495575 |
| FDR q-value | 0.3799558 |
| FWER p-Value | 0.995 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GSN | 1415812_at 1436991_x_at 1437171_x_at 1441225_at 1456312_x_at 1456568_at 1456569_x_at | 80 | 2.482 | 0.2922 | Yes | ||
| 2 | ITGB1 | 1426918_at 1426919_at 1426920_x_at 1427771_x_at 1438119_at 1452545_a_at | 251 | 1.572 | 0.4718 | Yes | ||
| 3 | PIK3R1 | 1425514_at 1425515_at 1438682_at 1444591_at 1451737_at | 343 | 1.346 | 0.6281 | Yes | ||
| 4 | FYN | 1417558_at 1441647_at 1448765_at | 398 | 1.220 | 0.7711 | Yes | ||
| 5 | PIK3CA | 1423144_at 1429434_at 1429435_x_at 1440054_at 1453134_at 1460326_at | 1919 | 0.273 | 0.7342 | No | ||
| 6 | ROCK1 | 1423444_at 1423445_at 1441162_at 1446518_at 1450994_at 1460729_at | 3461 | 0.137 | 0.6803 | No | ||
| 7 | SHC1 | 1422853_at 1422854_at | 4792 | 0.092 | 0.6306 | No | ||
| 8 | ITGA1 | 1439713_at 1446569_at 1455251_at | 6318 | 0.060 | 0.5681 | No | ||
| 9 | MYLK | 1425504_at 1425505_at 1425506_at | 8585 | 0.024 | 0.4676 | No | ||
| 10 | HRAS | 1422407_s_at 1424132_at | 8692 | 0.023 | 0.4655 | No | ||
| 11 | SRC | 1423240_at 1450918_s_at | 12329 | -0.025 | 0.3026 | No | ||
| 12 | RAF1 | 1416078_s_at 1425419_a_at | 12932 | -0.033 | 0.2791 | No | ||
| 13 | PFN1 | 1449018_at | 13688 | -0.044 | 0.2499 | No | ||
| 14 | MAPK1 | 1419568_at 1426585_s_at 1442876_at 1453104_at | 14178 | -0.052 | 0.2337 | No | ||
| 15 | MYL2 | 1448394_at | 15148 | -0.067 | 0.1975 | No | ||
| 16 | PXN | 1424027_at 1426085_a_at 1456135_s_at | 15420 | -0.072 | 0.1937 | No | ||
| 17 | TLN1 | 1436042_at 1448402_at 1457782_at | 15572 | -0.075 | 0.1957 | No | ||
| 18 | ARHGAP5 | 1423194_at 1450896_at 1450897_at 1457410_at | 15698 | -0.077 | 0.1992 | No | ||
| 19 | MAPK3 | 1427060_at | 18157 | -0.138 | 0.1034 | No | ||
| 20 | PTK2 | 1423059_at 1430827_a_at 1439198_at 1440082_at 1441475_at 1443384_at 1445137_at | 20132 | -0.251 | 0.0433 | No | ||
| 21 | MAP2K1 | 1416351_at | 20684 | -0.326 | 0.0570 | No |

