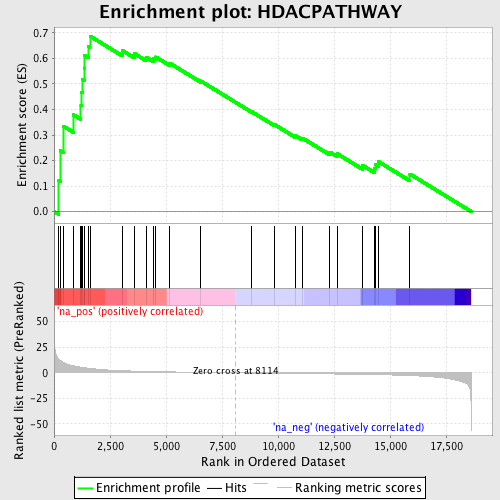
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | HDACPATHWAY |
| Enrichment Score (ES) | 0.68748343 |
| Normalized Enrichment Score (NES) | 1.837445 |
| Nominal p-value | 0.0019607844 |
| FDR q-value | 0.08451717 |
| FWER p-Value | 0.253 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NFATC1 | 213 | 13.199 | 0.1211 | Yes | ||
| 2 | MEF2C | 271 | 12.059 | 0.2391 | Yes | ||
| 3 | CAMK1G | 406 | 10.279 | 0.3351 | Yes | ||
| 4 | MEF2A | 873 | 6.858 | 0.3788 | Yes | ||
| 5 | CAMK1 | 1198 | 5.428 | 0.4159 | Yes | ||
| 6 | IGF1 | 1226 | 5.346 | 0.4681 | Yes | ||
| 7 | PPP3CC | 1281 | 5.185 | 0.5173 | Yes | ||
| 8 | CALM3 | 1361 | 4.940 | 0.5626 | Yes | ||
| 9 | MAPK14 | 1366 | 4.922 | 0.6118 | Yes | ||
| 10 | PPP3CB | 1536 | 4.496 | 0.6479 | Yes | ||
| 11 | IGF1R | 1603 | 4.301 | 0.6875 | Yes | ||
| 12 | PPP3CA | 3049 | 2.097 | 0.6308 | No | ||
| 13 | CALM2 | 3582 | 1.696 | 0.6192 | No | ||
| 14 | HDAC5 | 4107 | 1.376 | 0.6048 | No | ||
| 15 | PIK3CA | 4439 | 1.215 | 0.5992 | No | ||
| 16 | PIK3R1 | 4537 | 1.166 | 0.6057 | No | ||
| 17 | MEF2B | 5162 | 0.922 | 0.5814 | No | ||
| 18 | SYT1 | 6540 | 0.456 | 0.5119 | No | ||
| 19 | CABIN1 | 8824 | -0.187 | 0.3909 | No | ||
| 20 | NFATC2 | 9829 | -0.432 | 0.3413 | No | ||
| 21 | INSR | 10783 | -0.669 | 0.2967 | No | ||
| 22 | AVP | 11083 | -0.735 | 0.2880 | No | ||
| 23 | CALM1 | 12307 | -1.052 | 0.2328 | No | ||
| 24 | YWHAH | 12632 | -1.149 | 0.2269 | No | ||
| 25 | AKT1 | 13777 | -1.515 | 0.1805 | No | ||
| 26 | MAP2K6 | 14286 | -1.709 | 0.1704 | No | ||
| 27 | MYOD1 | 14356 | -1.745 | 0.1842 | No | ||
| 28 | MEF2D | 14489 | -1.807 | 0.1952 | No | ||
| 29 | MAPK7 | 15883 | -2.664 | 0.1470 | No |
