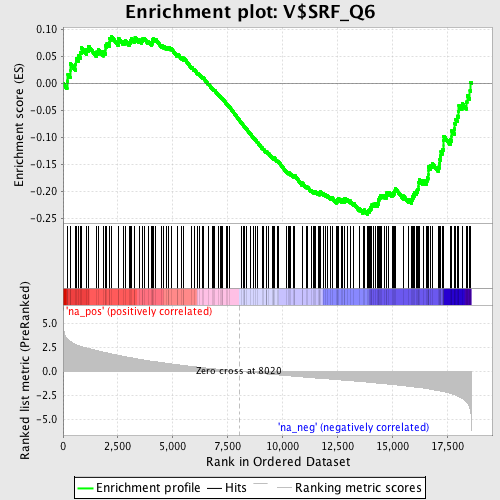
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_wtNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$SRF_Q6 |
| Enrichment Score (ES) | -0.24257919 |
| Normalized Enrichment Score (NES) | -1.2549441 |
| Nominal p-value | 0.075789474 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HOXB4 | 180 | 3.412 | 0.0046 | No | ||
| 2 | ROCK2 | 217 | 3.300 | 0.0165 | No | ||
| 3 | CNTF | 323 | 3.120 | 0.0239 | No | ||
| 4 | TPM3 | 329 | 3.115 | 0.0367 | No | ||
| 5 | SIN3A | 566 | 2.813 | 0.0357 | No | ||
| 6 | SVIL | 587 | 2.790 | 0.0463 | No | ||
| 7 | RBBP7 | 696 | 2.697 | 0.0517 | No | ||
| 8 | MBNL1 | 798 | 2.617 | 0.0573 | No | ||
| 9 | SCOC | 830 | 2.594 | 0.0665 | No | ||
| 10 | PADI2 | 1071 | 2.437 | 0.0637 | No | ||
| 11 | SYNCRIP | 1162 | 2.383 | 0.0688 | No | ||
| 12 | UBE2D3 | 1526 | 2.167 | 0.0582 | No | ||
| 13 | PPP1R12A | 1602 | 2.128 | 0.0630 | No | ||
| 14 | CFL2 | 1833 | 2.009 | 0.0590 | No | ||
| 15 | MATR3 | 1929 | 1.952 | 0.0620 | No | ||
| 16 | CACNA1B | 1931 | 1.951 | 0.0702 | No | ||
| 17 | MYL7 | 1997 | 1.921 | 0.0747 | No | ||
| 18 | RRM1 | 2114 | 1.857 | 0.0762 | No | ||
| 19 | H3F3A | 2122 | 1.854 | 0.0836 | No | ||
| 20 | ABR | 2192 | 1.822 | 0.0875 | No | ||
| 21 | RERE | 2517 | 1.672 | 0.0770 | No | ||
| 22 | NNAT | 2528 | 1.666 | 0.0834 | No | ||
| 23 | ATF4 | 2753 | 1.564 | 0.0778 | No | ||
| 24 | PICALM | 2847 | 1.519 | 0.0791 | No | ||
| 25 | HIVEP3 | 3026 | 1.443 | 0.0755 | No | ||
| 26 | SDPR | 3073 | 1.419 | 0.0790 | No | ||
| 27 | DHH | 3111 | 1.407 | 0.0829 | No | ||
| 28 | GPR20 | 3239 | 1.352 | 0.0817 | No | ||
| 29 | EML4 | 3275 | 1.339 | 0.0854 | No | ||
| 30 | PHF12 | 3462 | 1.268 | 0.0806 | No | ||
| 31 | PPAP2B | 3595 | 1.214 | 0.0786 | No | ||
| 32 | FOXP3 | 3618 | 1.207 | 0.0824 | No | ||
| 33 | KRTCAP2 | 3687 | 1.178 | 0.0837 | No | ||
| 34 | HOXA3 | 3878 | 1.112 | 0.0780 | No | ||
| 35 | SLC25A4 | 4028 | 1.055 | 0.0744 | No | ||
| 36 | RASSF2 | 4058 | 1.042 | 0.0772 | No | ||
| 37 | CFL1 | 4060 | 1.041 | 0.0815 | No | ||
| 38 | FOXP2 | 4119 | 1.024 | 0.0827 | No | ||
| 39 | ACTR3 | 4212 | 0.999 | 0.0819 | No | ||
| 40 | DGKG | 4492 | 0.910 | 0.0705 | No | ||
| 41 | BDNF | 4590 | 0.876 | 0.0689 | No | ||
| 42 | IGF1 | 4695 | 0.840 | 0.0668 | No | ||
| 43 | PRKAB2 | 4782 | 0.818 | 0.0656 | No | ||
| 44 | STARD13 | 4815 | 0.811 | 0.0673 | No | ||
| 45 | RAB27A | 4918 | 0.780 | 0.0650 | No | ||
| 46 | THBS1 | 5190 | 0.697 | 0.0532 | No | ||
| 47 | FOSB | 5227 | 0.687 | 0.0542 | No | ||
| 48 | ADAMTS12 | 5398 | 0.632 | 0.0476 | No | ||
| 49 | KIF1B | 5494 | 0.610 | 0.0450 | No | ||
| 50 | VCL | 5501 | 0.607 | 0.0472 | No | ||
| 51 | EFHD1 | 5868 | 0.508 | 0.0295 | No | ||
| 52 | FOXP1 | 5973 | 0.483 | 0.0259 | No | ||
| 53 | CABP1 | 6136 | 0.442 | 0.0189 | No | ||
| 54 | DUSP2 | 6198 | 0.425 | 0.0174 | No | ||
| 55 | TNMD | 6342 | 0.388 | 0.0113 | No | ||
| 56 | MYL1 | 6380 | 0.379 | 0.0109 | No | ||
| 57 | INSM1 | 6628 | 0.322 | -0.0012 | No | ||
| 58 | CORO1C | 6828 | 0.273 | -0.0109 | No | ||
| 59 | NPR3 | 6872 | 0.262 | -0.0121 | No | ||
| 60 | TRIM46 | 6901 | 0.254 | -0.0125 | No | ||
| 61 | EGR2 | 7090 | 0.214 | -0.0218 | No | ||
| 62 | MAP2K6 | 7150 | 0.203 | -0.0242 | No | ||
| 63 | MYL9 | 7222 | 0.183 | -0.0273 | No | ||
| 64 | SP7 | 7249 | 0.176 | -0.0280 | No | ||
| 65 | KCNK3 | 7267 | 0.171 | -0.0282 | No | ||
| 66 | IL17B | 7453 | 0.132 | -0.0376 | No | ||
| 67 | FST | 7512 | 0.114 | -0.0403 | No | ||
| 68 | GADD45G | 7569 | 0.104 | -0.0429 | No | ||
| 69 | MYH11 | 8138 | -0.024 | -0.0736 | No | ||
| 70 | KCNA1 | 8231 | -0.045 | -0.0784 | No | ||
| 71 | LSP1 | 8254 | -0.050 | -0.0794 | No | ||
| 72 | LDHA | 8255 | -0.050 | -0.0792 | No | ||
| 73 | RASD2 | 8336 | -0.070 | -0.0833 | No | ||
| 74 | KCNMB1 | 8561 | -0.118 | -0.0949 | No | ||
| 75 | MSX1 | 8669 | -0.138 | -0.1001 | No | ||
| 76 | HOXA5 | 8764 | -0.157 | -0.1046 | No | ||
| 77 | HOXD10 | 8856 | -0.177 | -0.1088 | No | ||
| 78 | NPAS2 | 9066 | -0.222 | -0.1192 | No | ||
| 79 | RSU1 | 9133 | -0.237 | -0.1218 | No | ||
| 80 | PDLIM3 | 9247 | -0.265 | -0.1268 | No | ||
| 81 | DIXDC1 | 9251 | -0.266 | -0.1259 | No | ||
| 82 | PLCB3 | 9259 | -0.267 | -0.1251 | No | ||
| 83 | TAGLN | 9365 | -0.289 | -0.1296 | No | ||
| 84 | TRDN | 9551 | -0.329 | -0.1383 | No | ||
| 85 | NFATC4 | 9593 | -0.339 | -0.1391 | No | ||
| 86 | NKX6-1 | 9595 | -0.339 | -0.1377 | No | ||
| 87 | ACTA1 | 9616 | -0.343 | -0.1373 | No | ||
| 88 | AQP2 | 9760 | -0.372 | -0.1435 | No | ||
| 89 | TNNC1 | 9808 | -0.381 | -0.1445 | No | ||
| 90 | FOSL1 | 10184 | -0.449 | -0.1630 | No | ||
| 91 | MYO1E | 10274 | -0.466 | -0.1658 | No | ||
| 92 | JUNB | 10304 | -0.471 | -0.1654 | No | ||
| 93 | HOXA1 | 10385 | -0.487 | -0.1677 | No | ||
| 94 | ELAVL4 | 10495 | -0.507 | -0.1715 | No | ||
| 95 | FOS | 10529 | -0.514 | -0.1711 | No | ||
| 96 | GNG8 | 10549 | -0.518 | -0.1700 | No | ||
| 97 | JPH2 | 10891 | -0.582 | -0.1861 | No | ||
| 98 | RAP1A | 10900 | -0.583 | -0.1840 | No | ||
| 99 | CNN1 | 11078 | -0.619 | -0.1911 | No | ||
| 100 | RAI2 | 11138 | -0.629 | -0.1916 | No | ||
| 101 | NKX2-2 | 11302 | -0.661 | -0.1977 | No | ||
| 102 | WNT5A | 11403 | -0.677 | -0.2003 | No | ||
| 103 | ACTG2 | 11456 | -0.687 | -0.2002 | No | ||
| 104 | SLC7A1 | 11502 | -0.696 | -0.1997 | No | ||
| 105 | MYADM | 11626 | -0.716 | -0.2034 | No | ||
| 106 | LPP | 11687 | -0.723 | -0.2036 | No | ||
| 107 | PRDM1 | 11699 | -0.726 | -0.2012 | No | ||
| 108 | EMILIN2 | 11722 | -0.728 | -0.1993 | No | ||
| 109 | PLN | 11869 | -0.755 | -0.2041 | No | ||
| 110 | SRF | 11958 | -0.772 | -0.2056 | No | ||
| 111 | RUNDC1 | 12053 | -0.791 | -0.2074 | No | ||
| 112 | FGFRL1 | 12186 | -0.813 | -0.2111 | No | ||
| 113 | CITED2 | 12257 | -0.828 | -0.2114 | No | ||
| 114 | H2AFX | 12446 | -0.860 | -0.2180 | No | ||
| 115 | TES | 12487 | -0.867 | -0.2166 | No | ||
| 116 | SIPA1 | 12507 | -0.872 | -0.2139 | No | ||
| 117 | RARB | 12546 | -0.879 | -0.2123 | No | ||
| 118 | SEPX1 | 12689 | -0.904 | -0.2162 | No | ||
| 119 | RHOJ | 12735 | -0.914 | -0.2148 | No | ||
| 120 | SIX2 | 12836 | -0.932 | -0.2163 | No | ||
| 121 | COL1A2 | 12837 | -0.932 | -0.2124 | No | ||
| 122 | NOL4 | 12966 | -0.957 | -0.2154 | No | ||
| 123 | HOXD13 | 13080 | -0.976 | -0.2174 | No | ||
| 124 | MYO18B | 13241 | -1.007 | -0.2219 | No | ||
| 125 | FGF8 | 13507 | -1.059 | -0.2318 | No | ||
| 126 | STAT5B | 13671 | -1.090 | -0.2361 | No | ||
| 127 | HOXB5 | 13718 | -1.098 | -0.2340 | No | ||
| 128 | ITGA7 | 13878 | -1.127 | -0.2378 | Yes | ||
| 129 | PDLIM4 | 13906 | -1.134 | -0.2346 | Yes | ||
| 130 | EPHB1 | 13980 | -1.148 | -0.2337 | Yes | ||
| 131 | SCN3B | 14002 | -1.153 | -0.2300 | Yes | ||
| 132 | TPM2 | 14053 | -1.164 | -0.2278 | Yes | ||
| 133 | MYLK | 14075 | -1.166 | -0.2241 | Yes | ||
| 134 | CDKN1B | 14151 | -1.185 | -0.2232 | Yes | ||
| 135 | PFN1 | 14232 | -1.203 | -0.2224 | Yes | ||
| 136 | DVL3 | 14330 | -1.228 | -0.2226 | Yes | ||
| 137 | PODN | 14367 | -1.236 | -0.2193 | Yes | ||
| 138 | FOXE3 | 14378 | -1.238 | -0.2147 | Yes | ||
| 139 | CKM | 14402 | -1.244 | -0.2107 | Yes | ||
| 140 | ATP1A2 | 14448 | -1.252 | -0.2079 | Yes | ||
| 141 | TAZ | 14527 | -1.272 | -0.2068 | Yes | ||
| 142 | RIMS1 | 14629 | -1.291 | -0.2068 | Yes | ||
| 143 | DMD | 14733 | -1.309 | -0.2069 | Yes | ||
| 144 | HOXC5 | 14755 | -1.315 | -0.2025 | Yes | ||
| 145 | COL1A1 | 14840 | -1.333 | -0.2015 | Yes | ||
| 146 | NR4A1 | 14996 | -1.372 | -0.2042 | Yes | ||
| 147 | MAP1A | 15060 | -1.386 | -0.2018 | Yes | ||
| 148 | PCDH7 | 15119 | -1.399 | -0.1990 | Yes | ||
| 149 | NR2F2 | 15134 | -1.401 | -0.1939 | Yes | ||
| 150 | ANXA6 | 15511 | -1.496 | -0.2080 | Yes | ||
| 151 | LRRTM4 | 15741 | -1.554 | -0.2140 | Yes | ||
| 152 | TLE3 | 15895 | -1.599 | -0.2155 | Yes | ||
| 153 | PRELP | 15910 | -1.602 | -0.2096 | Yes | ||
| 154 | NAV1 | 15963 | -1.619 | -0.2056 | Yes | ||
| 155 | ITGB1BP2 | 16025 | -1.638 | -0.2020 | Yes | ||
| 156 | CSMD3 | 16099 | -1.658 | -0.1990 | Yes | ||
| 157 | EGR3 | 16167 | -1.673 | -0.1957 | Yes | ||
| 158 | CALD1 | 16193 | -1.681 | -0.1900 | Yes | ||
| 159 | NR2F1 | 16196 | -1.681 | -0.1830 | Yes | ||
| 160 | LRRFIP1 | 16227 | -1.690 | -0.1775 | Yes | ||
| 161 | EGR1 | 16405 | -1.746 | -0.1798 | Yes | ||
| 162 | MRGPRF | 16549 | -1.800 | -0.1800 | Yes | ||
| 163 | CX3CL1 | 16590 | -1.816 | -0.1746 | Yes | ||
| 164 | HIST1H4I | 16635 | -1.830 | -0.1693 | Yes | ||
| 165 | NPM3 | 16640 | -1.831 | -0.1618 | Yes | ||
| 166 | LRP5 | 16644 | -1.834 | -0.1543 | Yes | ||
| 167 | ACTB | 16743 | -1.871 | -0.1517 | Yes | ||
| 168 | GPR158 | 16820 | -1.907 | -0.1478 | Yes | ||
| 169 | CNN2 | 17103 | -2.020 | -0.1547 | Yes | ||
| 170 | SRD5A2 | 17143 | -2.034 | -0.1482 | Yes | ||
| 171 | POU3F4 | 17169 | -2.043 | -0.1410 | Yes | ||
| 172 | DUSP10 | 17201 | -2.060 | -0.1341 | Yes | ||
| 173 | MUS81 | 17219 | -2.071 | -0.1263 | Yes | ||
| 174 | ACAA2 | 17306 | -2.106 | -0.1221 | Yes | ||
| 175 | TITF1 | 17338 | -2.121 | -0.1149 | Yes | ||
| 176 | CTDSP1 | 17339 | -2.121 | -0.1060 | Yes | ||
| 177 | BLOC1S1 | 17349 | -2.128 | -0.0975 | Yes | ||
| 178 | NPPA | 17648 | -2.303 | -0.1040 | Yes | ||
| 179 | PDLIM7 | 17707 | -2.343 | -0.0974 | Yes | ||
| 180 | SLC4A3 | 17711 | -2.345 | -0.0877 | Yes | ||
| 181 | MYL6 | 17836 | -2.450 | -0.0841 | Yes | ||
| 182 | ASPH | 17852 | -2.461 | -0.0746 | Yes | ||
| 183 | ARPC4 | 17893 | -2.491 | -0.0663 | Yes | ||
| 184 | TGFB1I1 | 17983 | -2.580 | -0.0603 | Yes | ||
| 185 | TADA3L | 18027 | -2.624 | -0.0516 | Yes | ||
| 186 | GRK6 | 18038 | -2.636 | -0.0411 | Yes | ||
| 187 | PPARG | 18183 | -2.809 | -0.0371 | Yes | ||
| 188 | CSNK1E | 18377 | -3.173 | -0.0343 | Yes | ||
| 189 | ABL1 | 18411 | -3.245 | -0.0224 | Yes | ||
| 190 | FLNA | 18543 | -3.845 | -0.0134 | Yes | ||
| 191 | MAPKAPK2 | 18571 | -4.126 | 0.0024 | Yes |
