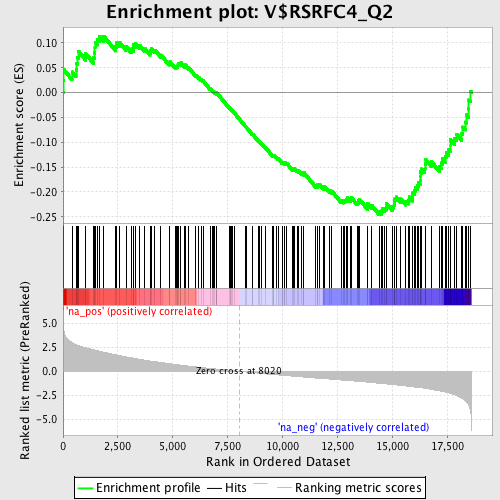
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_wtNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$RSRFC4_Q2 |
| Enrichment Score (ES) | -0.24547124 |
| Normalized Enrichment Score (NES) | -1.2253028 |
| Nominal p-value | 0.08713693 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PIK3R3 | 12 | 4.890 | 0.0249 | No | ||
| 2 | RG9MTD2 | 32 | 4.258 | 0.0462 | No | ||
| 3 | NFAT5 | 408 | 2.982 | 0.0415 | No | ||
| 4 | ERG | 589 | 2.789 | 0.0463 | No | ||
| 5 | ANKMY2 | 623 | 2.757 | 0.0590 | No | ||
| 6 | WBP4 | 668 | 2.716 | 0.0708 | No | ||
| 7 | CUL3 | 707 | 2.691 | 0.0828 | No | ||
| 8 | TRIM33 | 1034 | 2.459 | 0.0780 | No | ||
| 9 | TPP2 | 1392 | 2.244 | 0.0705 | No | ||
| 10 | RAP2C | 1416 | 2.232 | 0.0809 | No | ||
| 11 | MBNL2 | 1452 | 2.211 | 0.0906 | No | ||
| 12 | SLAMF1 | 1459 | 2.209 | 0.1018 | No | ||
| 13 | JMJD1C | 1572 | 2.143 | 0.1069 | No | ||
| 14 | CUTL1 | 1662 | 2.094 | 0.1131 | No | ||
| 15 | MYH4 | 1846 | 2.002 | 0.1137 | No | ||
| 16 | CDC42EP3 | 2401 | 1.721 | 0.0927 | No | ||
| 17 | ATP1B2 | 2430 | 1.708 | 0.1001 | No | ||
| 18 | SLC26A6 | 2560 | 1.653 | 0.1017 | No | ||
| 19 | CASQ2 | 2869 | 1.511 | 0.0930 | No | ||
| 20 | GRIN2B | 3102 | 1.411 | 0.0878 | No | ||
| 21 | LUC7L | 3195 | 1.371 | 0.0900 | No | ||
| 22 | FGF12 | 3212 | 1.365 | 0.0963 | No | ||
| 23 | PLAGL2 | 3302 | 1.330 | 0.0984 | No | ||
| 24 | HAPLN1 | 3487 | 1.260 | 0.0950 | No | ||
| 25 | AMPD1 | 3731 | 1.163 | 0.0880 | No | ||
| 26 | MEOX2 | 3961 | 1.079 | 0.0812 | No | ||
| 27 | LYN | 3996 | 1.067 | 0.0849 | No | ||
| 28 | PCBP2 | 4040 | 1.051 | 0.0881 | No | ||
| 29 | RASGRP3 | 4184 | 1.005 | 0.0856 | No | ||
| 30 | CNTN1 | 4458 | 0.919 | 0.0756 | No | ||
| 31 | EPHA7 | 4832 | 0.807 | 0.0597 | No | ||
| 32 | WFDC1 | 4870 | 0.798 | 0.0618 | No | ||
| 33 | HIBADH | 5105 | 0.721 | 0.0529 | No | ||
| 34 | TGFB3 | 5173 | 0.702 | 0.0530 | No | ||
| 35 | EYA1 | 5231 | 0.687 | 0.0535 | No | ||
| 36 | MITF | 5235 | 0.686 | 0.0569 | No | ||
| 37 | LZTS2 | 5270 | 0.677 | 0.0586 | No | ||
| 38 | BCL9L | 5348 | 0.649 | 0.0578 | No | ||
| 39 | GPC4 | 5372 | 0.640 | 0.0599 | No | ||
| 40 | ART5 | 5552 | 0.593 | 0.0534 | No | ||
| 41 | SLC8A3 | 5566 | 0.589 | 0.0557 | No | ||
| 42 | ZHX2 | 5732 | 0.541 | 0.0496 | No | ||
| 43 | MYF6 | 6049 | 0.464 | 0.0349 | No | ||
| 44 | STAC | 6176 | 0.429 | 0.0304 | No | ||
| 45 | KPNA3 | 6286 | 0.403 | 0.0266 | No | ||
| 46 | MYL1 | 6380 | 0.379 | 0.0235 | No | ||
| 47 | MGST3 | 6703 | 0.302 | 0.0077 | No | ||
| 48 | MYL3 | 6813 | 0.276 | 0.0032 | No | ||
| 49 | ASB16 | 6870 | 0.263 | 0.0015 | No | ||
| 50 | IRS1 | 6920 | 0.249 | 0.0002 | No | ||
| 51 | PAX3 | 6985 | 0.235 | -0.0020 | No | ||
| 52 | MUSK | 6990 | 0.234 | -0.0010 | No | ||
| 53 | SLC2A4 | 6993 | 0.234 | 0.0001 | No | ||
| 54 | HOXD4 | 7586 | 0.100 | -0.0315 | No | ||
| 55 | NME5 | 7613 | 0.094 | -0.0324 | No | ||
| 56 | COL8A1 | 7640 | 0.088 | -0.0333 | No | ||
| 57 | BZW2 | 7691 | 0.078 | -0.0356 | No | ||
| 58 | FBXW11 | 7737 | 0.066 | -0.0377 | No | ||
| 59 | LECT1 | 7824 | 0.049 | -0.0421 | No | ||
| 60 | EDG1 | 8326 | -0.068 | -0.0689 | No | ||
| 61 | PDGFRA | 8362 | -0.075 | -0.0704 | No | ||
| 62 | ZCCHC5 | 8629 | -0.130 | -0.0841 | No | ||
| 63 | THBS2 | 8649 | -0.134 | -0.0844 | No | ||
| 64 | TNNI3K | 8898 | -0.185 | -0.0969 | No | ||
| 65 | HOXC4 | 8957 | -0.197 | -0.0990 | No | ||
| 66 | ESR1 | 9054 | -0.220 | -0.1031 | No | ||
| 67 | SSH3 | 9240 | -0.262 | -0.1117 | No | ||
| 68 | TRDN | 9551 | -0.329 | -0.1268 | No | ||
| 69 | TMEPAI | 9583 | -0.337 | -0.1267 | No | ||
| 70 | ADAM11 | 9605 | -0.341 | -0.1260 | No | ||
| 71 | TAF5L | 9711 | -0.362 | -0.1298 | No | ||
| 72 | TNNC1 | 9808 | -0.381 | -0.1330 | No | ||
| 73 | PCDH9 | 10005 | -0.414 | -0.1415 | No | ||
| 74 | MYL2 | 10016 | -0.416 | -0.1399 | No | ||
| 75 | STOML2 | 10111 | -0.434 | -0.1427 | No | ||
| 76 | SLC30A1 | 10112 | -0.434 | -0.1404 | No | ||
| 77 | GABRB2 | 10183 | -0.448 | -0.1418 | No | ||
| 78 | SV2A | 10462 | -0.500 | -0.1543 | No | ||
| 79 | ELAVL4 | 10495 | -0.507 | -0.1534 | No | ||
| 80 | DNAJA4 | 10528 | -0.514 | -0.1524 | No | ||
| 81 | KCNJ9 | 10662 | -0.538 | -0.1568 | No | ||
| 82 | MRPS23 | 10735 | -0.552 | -0.1578 | No | ||
| 83 | DKK2 | 10858 | -0.574 | -0.1614 | No | ||
| 84 | TRPS1 | 10949 | -0.593 | -0.1632 | No | ||
| 85 | HAO1 | 10956 | -0.595 | -0.1604 | No | ||
| 86 | BAI3 | 11517 | -0.699 | -0.1871 | No | ||
| 87 | SSPN | 11582 | -0.708 | -0.1868 | No | ||
| 88 | ATF3 | 11613 | -0.714 | -0.1847 | No | ||
| 89 | PRDM1 | 11699 | -0.726 | -0.1855 | No | ||
| 90 | KCNN1 | 11880 | -0.757 | -0.1913 | No | ||
| 91 | HDAC9 | 11899 | -0.760 | -0.1883 | No | ||
| 92 | SLC29A2 | 12137 | -0.804 | -0.1969 | No | ||
| 93 | GPR153 | 12240 | -0.824 | -0.1981 | No | ||
| 94 | BNC2 | 12666 | -0.900 | -0.2164 | No | ||
| 95 | POFUT1 | 12776 | -0.921 | -0.2175 | No | ||
| 96 | ZFPM2 | 12845 | -0.934 | -0.2163 | No | ||
| 97 | KBTBD5 | 12917 | -0.947 | -0.2152 | No | ||
| 98 | FBXO40 | 12950 | -0.955 | -0.2120 | No | ||
| 99 | DIRAS1 | 13102 | -0.980 | -0.2150 | No | ||
| 100 | RALY | 13132 | -0.985 | -0.2114 | No | ||
| 101 | NEDD4 | 13400 | -1.039 | -0.2204 | No | ||
| 102 | CASQ1 | 13449 | -1.047 | -0.2176 | No | ||
| 103 | TNNC2 | 13496 | -1.056 | -0.2145 | No | ||
| 104 | ITGA7 | 13878 | -1.127 | -0.2293 | No | ||
| 105 | BNIP3 | 13886 | -1.129 | -0.2237 | No | ||
| 106 | TPM2 | 14053 | -1.164 | -0.2266 | No | ||
| 107 | CKM | 14402 | -1.244 | -0.2390 | Yes | ||
| 108 | INPP4A | 14515 | -1.269 | -0.2384 | Yes | ||
| 109 | TCEA3 | 14556 | -1.276 | -0.2339 | Yes | ||
| 110 | CRTAP | 14659 | -1.296 | -0.2326 | Yes | ||
| 111 | DMD | 14733 | -1.309 | -0.2297 | Yes | ||
| 112 | HDAC7A | 14737 | -1.310 | -0.2230 | Yes | ||
| 113 | NR4A1 | 14996 | -1.372 | -0.2298 | Yes | ||
| 114 | TYRO3 | 15081 | -1.390 | -0.2271 | Yes | ||
| 115 | SMPX | 15098 | -1.394 | -0.2207 | Yes | ||
| 116 | MYOZ2 | 15116 | -1.398 | -0.2143 | Yes | ||
| 117 | SDHC | 15178 | -1.410 | -0.2102 | Yes | ||
| 118 | SOX5 | 15391 | -1.461 | -0.2140 | Yes | ||
| 119 | LIF | 15622 | -1.523 | -0.2185 | Yes | ||
| 120 | ARR3 | 15731 | -1.550 | -0.2163 | Yes | ||
| 121 | SLCO3A1 | 15770 | -1.564 | -0.2101 | Yes | ||
| 122 | SMYD1 | 15925 | -1.607 | -0.2101 | Yes | ||
| 123 | PTPRO | 15935 | -1.610 | -0.2021 | Yes | ||
| 124 | CYP2E1 | 16009 | -1.633 | -0.1975 | Yes | ||
| 125 | HFE2 | 16057 | -1.645 | -0.1915 | Yes | ||
| 126 | CAPN3 | 16137 | -1.666 | -0.1870 | Yes | ||
| 127 | NR2F1 | 16196 | -1.681 | -0.1814 | Yes | ||
| 128 | POU4F1 | 16280 | -1.704 | -0.1770 | Yes | ||
| 129 | CLCN1 | 16286 | -1.705 | -0.1683 | Yes | ||
| 130 | BRUNOL4 | 16292 | -1.707 | -0.1596 | Yes | ||
| 131 | MAB21L2 | 16351 | -1.726 | -0.1537 | Yes | ||
| 132 | TNFRSF17 | 16494 | -1.778 | -0.1521 | Yes | ||
| 133 | CACNA2D3 | 16500 | -1.782 | -0.1431 | Yes | ||
| 134 | USP2 | 16532 | -1.793 | -0.1354 | Yes | ||
| 135 | EEF1A2 | 16774 | -1.888 | -0.1385 | Yes | ||
| 136 | TTYH2 | 17157 | -2.038 | -0.1486 | Yes | ||
| 137 | KTN1 | 17230 | -2.074 | -0.1416 | Yes | ||
| 138 | SLC26A9 | 17280 | -2.095 | -0.1333 | Yes | ||
| 139 | ALS2CR2 | 17415 | -2.166 | -0.1292 | Yes | ||
| 140 | TNNI1 | 17458 | -2.188 | -0.1201 | Yes | ||
| 141 | PRKAG1 | 17581 | -2.261 | -0.1148 | Yes | ||
| 142 | TWIST1 | 17639 | -2.293 | -0.1059 | Yes | ||
| 143 | PTPN1 | 17641 | -2.295 | -0.0940 | Yes | ||
| 144 | ASPH | 17852 | -2.461 | -0.0924 | Yes | ||
| 145 | HAS2 | 17943 | -2.542 | -0.0840 | Yes | ||
| 146 | PDLIM1 | 18174 | -2.796 | -0.0818 | Yes | ||
| 147 | MAP2K5 | 18221 | -2.871 | -0.0693 | Yes | ||
| 148 | RGS3 | 18334 | -3.081 | -0.0593 | Yes | ||
| 149 | NRXN3 | 18376 | -3.171 | -0.0449 | Yes | ||
| 150 | DTNB | 18478 | -3.435 | -0.0324 | Yes | ||
| 151 | RASGEF1B | 18493 | -3.482 | -0.0149 | Yes | ||
| 152 | ATP2A3 | 18568 | -4.113 | 0.0026 | Yes |
