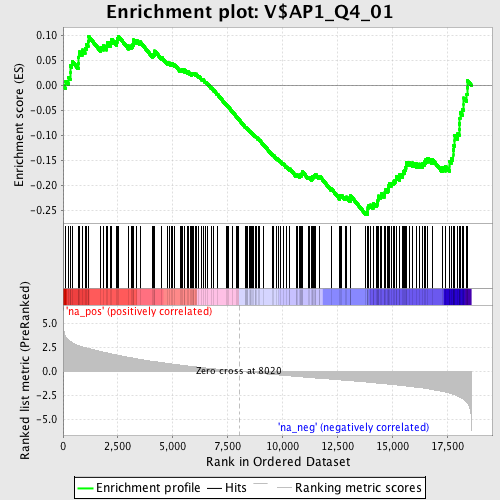
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_wtNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$AP1_Q4_01 |
| Enrichment Score (ES) | -0.25752857 |
| Normalized Enrichment Score (NES) | -1.326049 |
| Nominal p-value | 0.016393442 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GTF2B | 121 | 3.569 | 0.0092 | No | ||
| 2 | UBE2E3 | 257 | 3.230 | 0.0162 | No | ||
| 3 | CNTF | 323 | 3.120 | 0.0264 | No | ||
| 4 | SRPK2 | 330 | 3.115 | 0.0399 | No | ||
| 5 | PPP2CA | 405 | 2.984 | 0.0491 | No | ||
| 6 | RBBP7 | 696 | 2.697 | 0.0453 | No | ||
| 7 | VAPA | 706 | 2.692 | 0.0567 | No | ||
| 8 | TTC1 | 724 | 2.678 | 0.0676 | No | ||
| 9 | H3F3B | 864 | 2.572 | 0.0714 | No | ||
| 10 | LAMC2 | 1000 | 2.484 | 0.0751 | No | ||
| 11 | SFRS2 | 1068 | 2.439 | 0.0823 | No | ||
| 12 | PSMD4 | 1139 | 2.395 | 0.0890 | No | ||
| 13 | XLKD1 | 1170 | 2.379 | 0.0979 | No | ||
| 14 | RAB3D | 1720 | 2.065 | 0.0773 | No | ||
| 15 | RNF34 | 1834 | 2.008 | 0.0800 | No | ||
| 16 | BLR1 | 1998 | 1.921 | 0.0797 | No | ||
| 17 | TIAL1 | 2032 | 1.898 | 0.0863 | No | ||
| 18 | PSME4 | 2176 | 1.828 | 0.0866 | No | ||
| 19 | ATP1B1 | 2217 | 1.812 | 0.0924 | No | ||
| 20 | TAF15 | 2440 | 1.702 | 0.0879 | No | ||
| 21 | TOB1 | 2460 | 1.695 | 0.0944 | No | ||
| 22 | EMP1 | 2512 | 1.676 | 0.0990 | No | ||
| 23 | MYB | 2992 | 1.455 | 0.0795 | No | ||
| 24 | XPOT | 3106 | 1.408 | 0.0796 | No | ||
| 25 | TMEM24 | 3179 | 1.378 | 0.0818 | No | ||
| 26 | FGF12 | 3212 | 1.365 | 0.0861 | No | ||
| 27 | LAMA3 | 3222 | 1.361 | 0.0916 | No | ||
| 28 | NEFH | 3350 | 1.313 | 0.0905 | No | ||
| 29 | CRYGS | 3506 | 1.253 | 0.0876 | No | ||
| 30 | PSMD11 | 4075 | 1.037 | 0.0614 | No | ||
| 31 | GRIA1 | 4138 | 1.020 | 0.0625 | No | ||
| 32 | HCLS1 | 4173 | 1.009 | 0.0652 | No | ||
| 33 | DTNA | 4179 | 1.008 | 0.0693 | No | ||
| 34 | RIN1 | 4488 | 0.912 | 0.0567 | No | ||
| 35 | OMG | 4749 | 0.827 | 0.0462 | No | ||
| 36 | UCN2 | 4830 | 0.807 | 0.0454 | No | ||
| 37 | SNCB | 4921 | 0.778 | 0.0440 | No | ||
| 38 | KCNA2 | 4986 | 0.762 | 0.0439 | No | ||
| 39 | GTPBP2 | 5088 | 0.726 | 0.0416 | No | ||
| 40 | BCL9L | 5348 | 0.649 | 0.0304 | No | ||
| 41 | DYRK1A | 5380 | 0.637 | 0.0316 | No | ||
| 42 | SSH2 | 5422 | 0.627 | 0.0321 | No | ||
| 43 | SNX10 | 5518 | 0.603 | 0.0296 | No | ||
| 44 | WNT6 | 5531 | 0.599 | 0.0316 | No | ||
| 45 | SLC16A6 | 5662 | 0.561 | 0.0271 | No | ||
| 46 | IL11 | 5694 | 0.551 | 0.0278 | No | ||
| 47 | CDKN1A | 5813 | 0.522 | 0.0237 | No | ||
| 48 | BAZ2A | 5872 | 0.508 | 0.0228 | No | ||
| 49 | GNAI1 | 5896 | 0.500 | 0.0238 | No | ||
| 50 | HCN3 | 5928 | 0.493 | 0.0243 | No | ||
| 51 | CALB2 | 5962 | 0.484 | 0.0246 | No | ||
| 52 | LAMC1 | 6041 | 0.465 | 0.0224 | No | ||
| 53 | ADAM15 | 6063 | 0.462 | 0.0233 | No | ||
| 54 | ABHD4 | 6186 | 0.427 | 0.0186 | No | ||
| 55 | DHRS3 | 6324 | 0.394 | 0.0129 | No | ||
| 56 | MYH14 | 6375 | 0.380 | 0.0119 | No | ||
| 57 | TRAF3 | 6495 | 0.351 | 0.0070 | No | ||
| 58 | NR0B2 | 6589 | 0.331 | 0.0034 | No | ||
| 59 | RELA | 6756 | 0.290 | -0.0043 | No | ||
| 60 | SYTL1 | 6875 | 0.262 | -0.0096 | No | ||
| 61 | FGF11 | 7043 | 0.222 | -0.0177 | No | ||
| 62 | FBS1 | 7427 | 0.137 | -0.0378 | No | ||
| 63 | PTPRN | 7497 | 0.119 | -0.0411 | No | ||
| 64 | FBXO2 | 7554 | 0.107 | -0.0436 | No | ||
| 65 | FBXW11 | 7737 | 0.066 | -0.0532 | No | ||
| 66 | WNT7B | 7900 | 0.031 | -0.0619 | No | ||
| 67 | RGS2 | 7944 | 0.020 | -0.0641 | No | ||
| 68 | ABCF3 | 7982 | 0.009 | -0.0661 | No | ||
| 69 | PHLDA2 | 8303 | -0.061 | -0.0832 | No | ||
| 70 | EDG1 | 8326 | -0.068 | -0.0841 | No | ||
| 71 | SLC24A4 | 8337 | -0.070 | -0.0843 | No | ||
| 72 | CHST1 | 8369 | -0.076 | -0.0856 | No | ||
| 73 | GJA1 | 8400 | -0.082 | -0.0869 | No | ||
| 74 | LOR | 8401 | -0.082 | -0.0865 | No | ||
| 75 | BLMH | 8487 | -0.102 | -0.0907 | No | ||
| 76 | PLCD1 | 8531 | -0.112 | -0.0925 | No | ||
| 77 | AP2M1 | 8542 | -0.114 | -0.0926 | No | ||
| 78 | IGSF9 | 8599 | -0.125 | -0.0951 | No | ||
| 79 | SYT2 | 8611 | -0.127 | -0.0951 | No | ||
| 80 | CMAS | 8696 | -0.144 | -0.0990 | No | ||
| 81 | LMOD3 | 8751 | -0.155 | -0.1013 | No | ||
| 82 | FABP4 | 8771 | -0.158 | -0.1016 | No | ||
| 83 | ESRRG | 8806 | -0.163 | -0.1027 | No | ||
| 84 | BTK | 8888 | -0.183 | -0.1063 | No | ||
| 85 | RBP4 | 8956 | -0.197 | -0.1091 | No | ||
| 86 | PPP1R9B | 9138 | -0.238 | -0.1179 | No | ||
| 87 | COL7A1 | 9531 | -0.326 | -0.1377 | No | ||
| 88 | MAMDC2 | 9587 | -0.338 | -0.1392 | No | ||
| 89 | PVALB | 9745 | -0.369 | -0.1461 | No | ||
| 90 | MAPRE3 | 9817 | -0.382 | -0.1482 | No | ||
| 91 | SH3RF2 | 9903 | -0.397 | -0.1511 | No | ||
| 92 | MMP9 | 10035 | -0.419 | -0.1563 | No | ||
| 93 | FOSL1 | 10184 | -0.449 | -0.1624 | No | ||
| 94 | DDIT3 | 10322 | -0.473 | -0.1677 | No | ||
| 95 | PKP3 | 10338 | -0.478 | -0.1664 | No | ||
| 96 | AK5 | 10618 | -0.531 | -0.1792 | No | ||
| 97 | ANXA7 | 10625 | -0.532 | -0.1772 | No | ||
| 98 | CAMKK1 | 10693 | -0.543 | -0.1784 | No | ||
| 99 | COBL | 10792 | -0.563 | -0.1813 | No | ||
| 100 | SLC26A1 | 10803 | -0.565 | -0.1793 | No | ||
| 101 | MYBPH | 10819 | -0.567 | -0.1776 | No | ||
| 102 | NTN4 | 10881 | -0.579 | -0.1784 | No | ||
| 103 | IBRDC2 | 10885 | -0.580 | -0.1760 | No | ||
| 104 | SYNGR1 | 10893 | -0.582 | -0.1738 | No | ||
| 105 | TLL1 | 10904 | -0.584 | -0.1717 | No | ||
| 106 | AKT3 | 11164 | -0.635 | -0.1830 | No | ||
| 107 | EN1 | 11217 | -0.645 | -0.1829 | No | ||
| 108 | BACH1 | 11341 | -0.667 | -0.1867 | No | ||
| 109 | CMYA1 | 11352 | -0.669 | -0.1842 | No | ||
| 110 | USP3 | 11353 | -0.669 | -0.1813 | No | ||
| 111 | LRRTM3 | 11418 | -0.680 | -0.1817 | No | ||
| 112 | FSTL1 | 11441 | -0.684 | -0.1799 | No | ||
| 113 | CRYBA2 | 11482 | -0.692 | -0.1790 | No | ||
| 114 | NRAS | 11504 | -0.696 | -0.1771 | No | ||
| 115 | HOXA11 | 11663 | -0.721 | -0.1825 | No | ||
| 116 | PRDM1 | 11699 | -0.726 | -0.1812 | No | ||
| 117 | GDNF | 12227 | -0.822 | -0.2061 | No | ||
| 118 | SLC38A3 | 12615 | -0.893 | -0.2232 | No | ||
| 119 | GADD45A | 12625 | -0.894 | -0.2197 | No | ||
| 120 | VDR | 12686 | -0.903 | -0.2190 | No | ||
| 121 | ABCD1 | 12847 | -0.935 | -0.2235 | No | ||
| 122 | PRX | 12906 | -0.946 | -0.2225 | No | ||
| 123 | GKN1 | 13085 | -0.977 | -0.2278 | No | ||
| 124 | NDRG2 | 13101 | -0.980 | -0.2243 | No | ||
| 125 | DIRAS1 | 13102 | -0.980 | -0.2200 | No | ||
| 126 | TNXB | 13795 | -1.112 | -0.2526 | Yes | ||
| 127 | BNIP3 | 13886 | -1.129 | -0.2525 | Yes | ||
| 128 | CLIC1 | 13891 | -1.131 | -0.2477 | Yes | ||
| 129 | SHC3 | 13894 | -1.132 | -0.2428 | Yes | ||
| 130 | LTBP3 | 13934 | -1.140 | -0.2399 | Yes | ||
| 131 | OLR1 | 14003 | -1.153 | -0.2385 | Yes | ||
| 132 | MARK1 | 14128 | -1.180 | -0.2400 | Yes | ||
| 133 | ELAVL2 | 14146 | -1.184 | -0.2357 | Yes | ||
| 134 | SFN | 14273 | -1.213 | -0.2371 | Yes | ||
| 135 | CSF1R | 14309 | -1.222 | -0.2336 | Yes | ||
| 136 | PSTPIP1 | 14346 | -1.231 | -0.2301 | Yes | ||
| 137 | PADI4 | 14354 | -1.233 | -0.2251 | Yes | ||
| 138 | ITM2B | 14371 | -1.238 | -0.2205 | Yes | ||
| 139 | COL16A1 | 14484 | -1.260 | -0.2210 | Yes | ||
| 140 | MPV17 | 14497 | -1.264 | -0.2160 | Yes | ||
| 141 | RIMS1 | 14629 | -1.291 | -0.2174 | Yes | ||
| 142 | ADCY8 | 14668 | -1.297 | -0.2138 | Yes | ||
| 143 | SPATS2 | 14673 | -1.299 | -0.2082 | Yes | ||
| 144 | ADCK4 | 14776 | -1.321 | -0.2079 | Yes | ||
| 145 | SV2B | 14834 | -1.332 | -0.2051 | Yes | ||
| 146 | HDAC3 | 14846 | -1.334 | -0.1998 | Yes | ||
| 147 | RIT1 | 14872 | -1.341 | -0.1953 | Yes | ||
| 148 | WDFY3 | 14976 | -1.369 | -0.1948 | Yes | ||
| 149 | EPHB2 | 15046 | -1.382 | -0.1924 | Yes | ||
| 150 | MYOZ2 | 15116 | -1.398 | -0.1900 | Yes | ||
| 151 | PAPPA | 15172 | -1.409 | -0.1867 | Yes | ||
| 152 | FBXO44 | 15189 | -1.412 | -0.1814 | Yes | ||
| 153 | RPS6KA4 | 15342 | -1.450 | -0.1832 | Yes | ||
| 154 | HSPB8 | 15345 | -1.450 | -0.1769 | Yes | ||
| 155 | NR1D1 | 15465 | -1.482 | -0.1768 | Yes | ||
| 156 | DUSP14 | 15492 | -1.491 | -0.1716 | Yes | ||
| 157 | ECM1 | 15572 | -1.510 | -0.1692 | Yes | ||
| 158 | TREX1 | 15598 | -1.517 | -0.1639 | Yes | ||
| 159 | SEC24D | 15629 | -1.525 | -0.1588 | Yes | ||
| 160 | ELK3 | 15642 | -1.529 | -0.1526 | Yes | ||
| 161 | VIT | 15772 | -1.565 | -0.1527 | Yes | ||
| 162 | BAG2 | 15934 | -1.610 | -0.1543 | Yes | ||
| 163 | CSMD3 | 16099 | -1.658 | -0.1559 | Yes | ||
| 164 | ITPKC | 16251 | -1.697 | -0.1566 | Yes | ||
| 165 | PEA15 | 16381 | -1.738 | -0.1559 | Yes | ||
| 166 | CHST4 | 16458 | -1.766 | -0.1522 | Yes | ||
| 167 | USP2 | 16532 | -1.793 | -0.1483 | Yes | ||
| 168 | AQP5 | 16619 | -1.824 | -0.1449 | Yes | ||
| 169 | CD68 | 16828 | -1.910 | -0.1477 | Yes | ||
| 170 | SLC26A9 | 17280 | -2.095 | -0.1629 | Yes | ||
| 171 | ITGB4 | 17425 | -2.168 | -0.1611 | Yes | ||
| 172 | CSF3 | 17608 | -2.276 | -0.1609 | Yes | ||
| 173 | TUBB4 | 17609 | -2.279 | -0.1509 | Yes | ||
| 174 | SLC35B1 | 17686 | -2.330 | -0.1447 | Yes | ||
| 175 | P2RXL1 | 17770 | -2.396 | -0.1386 | Yes | ||
| 176 | RHBDF1 | 17772 | -2.396 | -0.1281 | Yes | ||
| 177 | ALDOA | 17804 | -2.426 | -0.1190 | Yes | ||
| 178 | NCDN | 17843 | -2.457 | -0.1102 | Yes | ||
| 179 | CDH23 | 17846 | -2.460 | -0.0994 | Yes | ||
| 180 | MNT | 17992 | -2.589 | -0.0959 | Yes | ||
| 181 | SQSTM1 | 18052 | -2.652 | -0.0873 | Yes | ||
| 182 | MAP4K5 | 18060 | -2.658 | -0.0759 | Yes | ||
| 183 | ROM1 | 18082 | -2.685 | -0.0652 | Yes | ||
| 184 | VASP | 18094 | -2.698 | -0.0539 | Yes | ||
| 185 | RB1CC1 | 18195 | -2.822 | -0.0468 | Yes | ||
| 186 | MARK4 | 18264 | -2.944 | -0.0375 | Yes | ||
| 187 | ENO1 | 18267 | -2.947 | -0.0246 | Yes | ||
| 188 | TYSND1 | 18401 | -3.223 | -0.0175 | Yes | ||
| 189 | IL10 | 18421 | -3.274 | -0.0041 | Yes | ||
| 190 | ETV5 | 18434 | -3.305 | 0.0099 | Yes |
