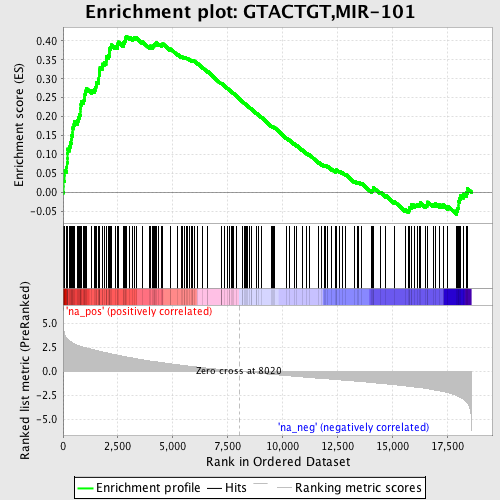
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_wtNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | GTACTGT,MIR-101 |
| Enrichment Score (ES) | 0.41260028 |
| Normalized Enrichment Score (NES) | 2.0980241 |
| Nominal p-value | 0.0 |
| FDR q-value | 7.570554E-4 |
| FWER p-Value | 0.0080 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ERBB2IP | 29 | 4.297 | 0.0145 | Yes | ||
| 2 | PPFIA1 | 31 | 4.282 | 0.0305 | Yes | ||
| 3 | UBE2Q1 | 61 | 3.971 | 0.0438 | Yes | ||
| 4 | CDYL | 64 | 3.934 | 0.0584 | Yes | ||
| 5 | MAGI1 | 158 | 3.456 | 0.0663 | Yes | ||
| 6 | EED | 177 | 3.421 | 0.0781 | Yes | ||
| 7 | PAPOLG | 187 | 3.392 | 0.0903 | Yes | ||
| 8 | ANKRD17 | 193 | 3.382 | 0.1027 | Yes | ||
| 9 | LRRFIP2 | 203 | 3.333 | 0.1146 | Yes | ||
| 10 | REV3L | 304 | 3.147 | 0.1210 | Yes | ||
| 11 | SRPK2 | 330 | 3.115 | 0.1313 | Yes | ||
| 12 | CTCF | 372 | 3.040 | 0.1404 | Yes | ||
| 13 | TERF2 | 396 | 3.011 | 0.1505 | Yes | ||
| 14 | WSB1 | 435 | 2.947 | 0.1594 | Yes | ||
| 15 | RAB1A | 439 | 2.944 | 0.1703 | Yes | ||
| 16 | KPNB1 | 493 | 2.884 | 0.1782 | Yes | ||
| 17 | AP3S1 | 517 | 2.860 | 0.1876 | Yes | ||
| 18 | OGT | 665 | 2.718 | 0.1898 | Yes | ||
| 19 | RBBP7 | 696 | 2.697 | 0.1983 | Yes | ||
| 20 | NLK | 741 | 2.665 | 0.2059 | Yes | ||
| 21 | MBNL1 | 798 | 2.617 | 0.2126 | Yes | ||
| 22 | TLK2 | 800 | 2.616 | 0.2224 | Yes | ||
| 23 | PREI3 | 811 | 2.607 | 0.2316 | Yes | ||
| 24 | RANBP9 | 833 | 2.593 | 0.2401 | Yes | ||
| 25 | SLC39A10 | 949 | 2.516 | 0.2433 | Yes | ||
| 26 | TULP4 | 986 | 2.492 | 0.2507 | Yes | ||
| 27 | UBE2A | 993 | 2.488 | 0.2597 | Yes | ||
| 28 | PURG | 1015 | 2.478 | 0.2678 | Yes | ||
| 29 | STK4 | 1059 | 2.443 | 0.2746 | Yes | ||
| 30 | TXNDC4 | 1301 | 2.297 | 0.2701 | Yes | ||
| 31 | RAP2C | 1416 | 2.232 | 0.2723 | Yes | ||
| 32 | ARID1A | 1460 | 2.208 | 0.2782 | Yes | ||
| 33 | PUM2 | 1500 | 2.184 | 0.2843 | Yes | ||
| 34 | UBE2D3 | 1526 | 2.167 | 0.2910 | Yes | ||
| 35 | SPOP | 1608 | 2.123 | 0.2946 | Yes | ||
| 36 | CLCN3 | 1631 | 2.113 | 0.3013 | Yes | ||
| 37 | TGFBR1 | 1635 | 2.109 | 0.3090 | Yes | ||
| 38 | PANK3 | 1649 | 2.099 | 0.3162 | Yes | ||
| 39 | FEM1C | 1661 | 2.094 | 0.3234 | Yes | ||
| 40 | DDX3Y | 1680 | 2.085 | 0.3302 | Yes | ||
| 41 | GLRA2 | 1779 | 2.035 | 0.3325 | Yes | ||
| 42 | PDE4D | 1800 | 2.027 | 0.3390 | Yes | ||
| 43 | RAP1B | 1871 | 1.987 | 0.3427 | Yes | ||
| 44 | UBE2D2 | 1956 | 1.936 | 0.3454 | Yes | ||
| 45 | ICK | 1960 | 1.935 | 0.3524 | Yes | ||
| 46 | MPPE1 | 1984 | 1.925 | 0.3584 | Yes | ||
| 47 | ING3 | 2068 | 1.878 | 0.3609 | Yes | ||
| 48 | DDEF1 | 2109 | 1.859 | 0.3657 | Yes | ||
| 49 | RRM1 | 2114 | 1.857 | 0.3724 | Yes | ||
| 50 | MTMR2 | 2121 | 1.855 | 0.3791 | Yes | ||
| 51 | EZH2 | 2166 | 1.831 | 0.3835 | Yes | ||
| 52 | STAG2 | 2183 | 1.825 | 0.3895 | Yes | ||
| 53 | FBXW7 | 2368 | 1.732 | 0.3860 | Yes | ||
| 54 | LRRC4 | 2485 | 1.684 | 0.3860 | Yes | ||
| 55 | DDX3X | 2492 | 1.681 | 0.3919 | Yes | ||
| 56 | EMP1 | 2512 | 1.676 | 0.3972 | Yes | ||
| 57 | SFRS5 | 2739 | 1.569 | 0.3908 | Yes | ||
| 58 | ZNF24 | 2762 | 1.561 | 0.3954 | Yes | ||
| 59 | QKI | 2799 | 1.540 | 0.3993 | Yes | ||
| 60 | AEBP2 | 2833 | 1.525 | 0.4032 | Yes | ||
| 61 | POGZ | 2850 | 1.518 | 0.4080 | Yes | ||
| 62 | EXOC5 | 2870 | 1.510 | 0.4126 | Yes | ||
| 63 | NACA | 3027 | 1.443 | 0.4095 | No | ||
| 64 | CDH11 | 3180 | 1.378 | 0.4064 | No | ||
| 65 | SLC19A2 | 3232 | 1.357 | 0.4088 | No | ||
| 66 | SOCS5 | 3330 | 1.320 | 0.4084 | No | ||
| 67 | USP47 | 3601 | 1.213 | 0.3983 | No | ||
| 68 | DAG1 | 3935 | 1.089 | 0.3843 | No | ||
| 69 | LMNB1 | 3967 | 1.076 | 0.3867 | No | ||
| 70 | NEGR1 | 4084 | 1.035 | 0.3843 | No | ||
| 71 | PALM2 | 4099 | 1.031 | 0.3874 | No | ||
| 72 | PURB | 4143 | 1.019 | 0.3888 | No | ||
| 73 | PTGS2 | 4168 | 1.011 | 0.3913 | No | ||
| 74 | CDK5R1 | 4222 | 0.995 | 0.3922 | No | ||
| 75 | BICD2 | 4237 | 0.990 | 0.3951 | No | ||
| 76 | ABCC5 | 4369 | 0.948 | 0.3915 | No | ||
| 77 | ADAMTS10 | 4506 | 0.906 | 0.3876 | No | ||
| 78 | THRB | 4514 | 0.903 | 0.3906 | No | ||
| 79 | STC1 | 4549 | 0.889 | 0.3920 | No | ||
| 80 | CDH5 | 4877 | 0.795 | 0.3773 | No | ||
| 81 | SOCS2 | 4904 | 0.784 | 0.3788 | No | ||
| 82 | EYA1 | 5231 | 0.687 | 0.3637 | No | ||
| 83 | DYRK1A | 5380 | 0.637 | 0.3580 | No | ||
| 84 | KIF1A | 5431 | 0.624 | 0.3577 | No | ||
| 85 | TAL1 | 5513 | 0.604 | 0.3555 | No | ||
| 86 | GNB1 | 5549 | 0.594 | 0.3559 | No | ||
| 87 | SIX4 | 5611 | 0.576 | 0.3547 | No | ||
| 88 | MYRIP | 5661 | 0.561 | 0.3541 | No | ||
| 89 | MYST2 | 5755 | 0.536 | 0.3511 | No | ||
| 90 | BAZ2A | 5872 | 0.508 | 0.3467 | No | ||
| 91 | DR1 | 5882 | 0.503 | 0.3481 | No | ||
| 92 | NRK | 5913 | 0.496 | 0.3483 | No | ||
| 93 | FZD6 | 5972 | 0.484 | 0.3470 | No | ||
| 94 | CCNJ | 6129 | 0.444 | 0.3402 | No | ||
| 95 | PPFIA4 | 6365 | 0.383 | 0.3289 | No | ||
| 96 | NDFIP1 | 6580 | 0.333 | 0.3185 | No | ||
| 97 | CBL | 6592 | 0.331 | 0.3192 | No | ||
| 98 | PHLDA1 | 7196 | 0.191 | 0.2872 | No | ||
| 99 | UBE2D1 | 7208 | 0.187 | 0.2873 | No | ||
| 100 | SOX11 | 7214 | 0.185 | 0.2877 | No | ||
| 101 | TEAD3 | 7219 | 0.184 | 0.2882 | No | ||
| 102 | CTTNBP2 | 7372 | 0.147 | 0.2805 | No | ||
| 103 | HNRPF | 7505 | 0.115 | 0.2737 | No | ||
| 104 | HOXD4 | 7586 | 0.100 | 0.2698 | No | ||
| 105 | LRRN3 | 7599 | 0.097 | 0.2695 | No | ||
| 106 | BZW2 | 7691 | 0.078 | 0.2648 | No | ||
| 107 | FBXW11 | 7737 | 0.066 | 0.2626 | No | ||
| 108 | JDP2 | 7784 | 0.058 | 0.2604 | No | ||
| 109 | SPRED1 | 7924 | 0.025 | 0.2529 | No | ||
| 110 | RIN2 | 8165 | -0.031 | 0.2400 | No | ||
| 111 | CAV3 | 8168 | -0.032 | 0.2400 | No | ||
| 112 | SMARCA1 | 8272 | -0.053 | 0.2346 | No | ||
| 113 | ZCCHC2 | 8334 | -0.069 | 0.2316 | No | ||
| 114 | RASD2 | 8336 | -0.070 | 0.2318 | No | ||
| 115 | GJA1 | 8400 | -0.082 | 0.2287 | No | ||
| 116 | ASPN | 8489 | -0.102 | 0.2243 | No | ||
| 117 | SLC30A7 | 8568 | -0.119 | 0.2205 | No | ||
| 118 | DCBLD2 | 8572 | -0.120 | 0.2208 | No | ||
| 119 | LRRN1 | 8800 | -0.162 | 0.2091 | No | ||
| 120 | ZFHX4 | 8805 | -0.163 | 0.2095 | No | ||
| 121 | MAK | 8914 | -0.188 | 0.2043 | No | ||
| 122 | SGPL1 | 9042 | -0.217 | 0.1982 | No | ||
| 123 | DDIT4 | 9491 | -0.317 | 0.1751 | No | ||
| 124 | APP | 9545 | -0.328 | 0.1735 | No | ||
| 125 | LASS2 | 9581 | -0.337 | 0.1728 | No | ||
| 126 | ZFAND3 | 9645 | -0.350 | 0.1707 | No | ||
| 127 | PACRG | 10190 | -0.450 | 0.1429 | No | ||
| 128 | JUNB | 10304 | -0.471 | 0.1385 | No | ||
| 129 | FOS | 10529 | -0.514 | 0.1283 | No | ||
| 130 | CEBPA | 10653 | -0.537 | 0.1237 | No | ||
| 131 | KHDRBS3 | 10905 | -0.585 | 0.1122 | No | ||
| 132 | TMEFF1 | 11108 | -0.624 | 0.1036 | No | ||
| 133 | RIPK5 | 11224 | -0.646 | 0.0998 | No | ||
| 134 | NEUROD1 | 11657 | -0.720 | 0.0790 | No | ||
| 135 | RAB5A | 11791 | -0.738 | 0.0746 | No | ||
| 136 | FZD4 | 11909 | -0.762 | 0.0711 | No | ||
| 137 | RGS1 | 11939 | -0.769 | 0.0724 | No | ||
| 138 | PRKCE | 12056 | -0.791 | 0.0691 | No | ||
| 139 | PRKAA1 | 12247 | -0.826 | 0.0618 | No | ||
| 140 | PRPF4B | 12408 | -0.852 | 0.0564 | No | ||
| 141 | ROBO2 | 12437 | -0.859 | 0.0580 | No | ||
| 142 | SLC1A1 | 12469 | -0.865 | 0.0596 | No | ||
| 143 | CDK6 | 12593 | -0.888 | 0.0563 | No | ||
| 144 | NUDT11 | 12733 | -0.914 | 0.0521 | No | ||
| 145 | BCL9 | 12875 | -0.940 | 0.0480 | No | ||
| 146 | IGFBP5 | 13298 | -1.020 | 0.0289 | No | ||
| 147 | RNF38 | 13398 | -1.039 | 0.0274 | No | ||
| 148 | BTBD3 | 13471 | -1.053 | 0.0275 | No | ||
| 149 | PLXNB1 | 13601 | -1.079 | 0.0245 | No | ||
| 150 | CPEB2 | 14049 | -1.163 | 0.0046 | No | ||
| 151 | PDE4A | 14111 | -1.176 | 0.0057 | No | ||
| 152 | MARK1 | 14128 | -1.180 | 0.0092 | No | ||
| 153 | FLRT2 | 14141 | -1.183 | 0.0130 | No | ||
| 154 | ADCY6 | 14458 | -1.254 | 0.0006 | No | ||
| 155 | SOX9 | 14712 | -1.304 | -0.0083 | No | ||
| 156 | CPEB3 | 15108 | -1.397 | -0.0245 | No | ||
| 157 | PCDH8 | 15599 | -1.517 | -0.0454 | No | ||
| 158 | ATXN1 | 15736 | -1.552 | -0.0470 | No | ||
| 159 | COL10A1 | 15776 | -1.567 | -0.0432 | No | ||
| 160 | TSC22D1 | 15802 | -1.574 | -0.0387 | No | ||
| 161 | TRIB1 | 15881 | -1.594 | -0.0369 | No | ||
| 162 | PLCG1 | 15887 | -1.596 | -0.0312 | No | ||
| 163 | FLRT3 | 16037 | -1.642 | -0.0332 | No | ||
| 164 | KCNH7 | 16131 | -1.664 | -0.0320 | No | ||
| 165 | DNMT3A | 16236 | -1.692 | -0.0313 | No | ||
| 166 | ACCN2 | 16277 | -1.703 | -0.0271 | No | ||
| 167 | RXRB | 16536 | -1.795 | -0.0344 | No | ||
| 168 | SMARCD1 | 16587 | -1.813 | -0.0303 | No | ||
| 169 | DOT1L | 16621 | -1.825 | -0.0253 | No | ||
| 170 | ATRX | 16863 | -1.921 | -0.0312 | No | ||
| 171 | MMP15 | 16962 | -1.961 | -0.0291 | No | ||
| 172 | FBN2 | 17164 | -2.042 | -0.0324 | No | ||
| 173 | RAPH1 | 17314 | -2.109 | -0.0326 | No | ||
| 174 | USP38 | 17536 | -2.238 | -0.0362 | No | ||
| 175 | HAS2 | 17943 | -2.542 | -0.0487 | No | ||
| 176 | FGA | 17990 | -2.588 | -0.0415 | No | ||
| 177 | PIP5K1C | 18028 | -2.624 | -0.0337 | No | ||
| 178 | DLG5 | 18030 | -2.628 | -0.0239 | No | ||
| 179 | GPR85 | 18081 | -2.683 | -0.0166 | No | ||
| 180 | BCL2L11 | 18124 | -2.738 | -0.0086 | No | ||
| 181 | CACNB2 | 18241 | -2.911 | -0.0040 | No | ||
| 182 | DUSP1 | 18382 | -3.185 | 0.0003 | No | ||
| 183 | ETV5 | 18434 | -3.305 | 0.0099 | No |
