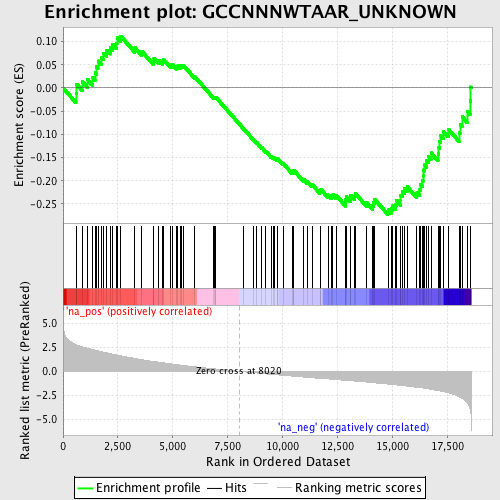
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_wtNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | GCCNNNWTAAR_UNKNOWN |
| Enrichment Score (ES) | -0.27269652 |
| Normalized Enrichment Score (NES) | -1.2812556 |
| Nominal p-value | 0.088794924 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ERG | 589 | 2.789 | -0.0110 | No | ||
| 2 | HPS3 | 613 | 2.770 | 0.0083 | No | ||
| 3 | PTMA | 867 | 2.571 | 0.0138 | No | ||
| 4 | PHEX | 1114 | 2.412 | 0.0185 | No | ||
| 5 | ITPR1 | 1346 | 2.271 | 0.0229 | No | ||
| 6 | ARID1A | 1460 | 2.208 | 0.0333 | No | ||
| 7 | UBE2D3 | 1526 | 2.167 | 0.0459 | No | ||
| 8 | RBMX | 1593 | 2.135 | 0.0582 | No | ||
| 9 | ZADH2 | 1742 | 2.055 | 0.0656 | No | ||
| 10 | MPP2 | 1837 | 2.007 | 0.0754 | No | ||
| 11 | OTX1 | 1994 | 1.922 | 0.0813 | No | ||
| 12 | EN2 | 2144 | 1.841 | 0.0870 | No | ||
| 13 | ITGA3 | 2252 | 1.792 | 0.0945 | No | ||
| 14 | MID1 | 2443 | 1.701 | 0.0969 | No | ||
| 15 | XPO7 | 2458 | 1.696 | 0.1088 | No | ||
| 16 | SOX4 | 2626 | 1.618 | 0.1119 | No | ||
| 17 | EML4 | 3275 | 1.339 | 0.0868 | No | ||
| 18 | HMGA2 | 3580 | 1.219 | 0.0795 | No | ||
| 19 | FOXP2 | 4119 | 1.024 | 0.0581 | No | ||
| 20 | KLC2 | 4139 | 1.020 | 0.0646 | No | ||
| 21 | ABCC5 | 4369 | 0.948 | 0.0593 | No | ||
| 22 | PPARGC1A | 4538 | 0.893 | 0.0569 | No | ||
| 23 | OTP | 4589 | 0.876 | 0.0607 | No | ||
| 24 | SOCS2 | 4904 | 0.784 | 0.0496 | No | ||
| 25 | APBA2BP | 4996 | 0.758 | 0.0503 | No | ||
| 26 | TGFB3 | 5173 | 0.702 | 0.0460 | No | ||
| 27 | MITF | 5235 | 0.686 | 0.0479 | No | ||
| 28 | ADAM12 | 5333 | 0.655 | 0.0475 | No | ||
| 29 | ASGR1 | 5414 | 0.629 | 0.0479 | No | ||
| 30 | VCL | 5501 | 0.607 | 0.0477 | No | ||
| 31 | SFXN5 | 6010 | 0.475 | 0.0238 | No | ||
| 32 | NPR3 | 6872 | 0.262 | -0.0207 | No | ||
| 33 | SLC5A7 | 6893 | 0.257 | -0.0199 | No | ||
| 34 | KCNH3 | 6962 | 0.240 | -0.0218 | No | ||
| 35 | SHH | 6966 | 0.239 | -0.0201 | No | ||
| 36 | ADAMTS5 | 8243 | -0.048 | -0.0887 | No | ||
| 37 | MSX1 | 8669 | -0.138 | -0.1106 | No | ||
| 38 | ISL1 | 8824 | -0.168 | -0.1177 | No | ||
| 39 | BHLHB3 | 9060 | -0.221 | -0.1287 | No | ||
| 40 | TBCC | 9241 | -0.263 | -0.1365 | No | ||
| 41 | NFIL3 | 9483 | -0.315 | -0.1472 | No | ||
| 42 | NFATC4 | 9593 | -0.339 | -0.1505 | No | ||
| 43 | IMPDH2 | 9646 | -0.350 | -0.1507 | No | ||
| 44 | STK35 | 9766 | -0.373 | -0.1544 | No | ||
| 45 | SFTPC | 9782 | -0.376 | -0.1524 | No | ||
| 46 | MMP9 | 10035 | -0.419 | -0.1629 | No | ||
| 47 | NPTX1 | 10441 | -0.496 | -0.1811 | No | ||
| 48 | ELAVL4 | 10495 | -0.507 | -0.1801 | No | ||
| 49 | RFX4 | 10515 | -0.511 | -0.1774 | No | ||
| 50 | PHOX2B | 10967 | -0.597 | -0.1973 | No | ||
| 51 | JPH4 | 11146 | -0.631 | -0.2022 | No | ||
| 52 | DPF3 | 11344 | -0.667 | -0.2079 | No | ||
| 53 | MEF2D | 11709 | -0.727 | -0.2221 | No | ||
| 54 | EGFL6 | 11748 | -0.732 | -0.2187 | No | ||
| 55 | HOXA4 | 12079 | -0.795 | -0.2306 | No | ||
| 56 | GDNF | 12227 | -0.822 | -0.2324 | No | ||
| 57 | ABCA1 | 12299 | -0.834 | -0.2300 | No | ||
| 58 | H2AFX | 12446 | -0.860 | -0.2315 | No | ||
| 59 | BCL9 | 12875 | -0.940 | -0.2476 | No | ||
| 60 | FGF9 | 12884 | -0.942 | -0.2410 | No | ||
| 61 | ATOH1 | 12896 | -0.944 | -0.2346 | No | ||
| 62 | NDRG2 | 13101 | -0.980 | -0.2383 | No | ||
| 63 | VAX1 | 13111 | -0.981 | -0.2315 | No | ||
| 64 | TRIB2 | 13295 | -1.019 | -0.2338 | No | ||
| 65 | COCH | 13309 | -1.021 | -0.2269 | No | ||
| 66 | PDGFB | 13835 | -1.119 | -0.2469 | No | ||
| 67 | RAB33A | 14105 | -1.175 | -0.2527 | No | ||
| 68 | CDKN1B | 14151 | -1.185 | -0.2463 | No | ||
| 69 | PTPRG | 14208 | -1.198 | -0.2404 | No | ||
| 70 | KCTD15 | 14807 | -1.328 | -0.2628 | Yes | ||
| 71 | MMP24 | 14948 | -1.359 | -0.2602 | Yes | ||
| 72 | FHL3 | 15027 | -1.378 | -0.2542 | Yes | ||
| 73 | NKX2-5 | 15168 | -1.408 | -0.2513 | Yes | ||
| 74 | SLC39A5 | 15202 | -1.416 | -0.2425 | Yes | ||
| 75 | SOX5 | 15391 | -1.461 | -0.2418 | Yes | ||
| 76 | HOXA10 | 15394 | -1.462 | -0.2310 | Yes | ||
| 77 | FES | 15456 | -1.479 | -0.2233 | Yes | ||
| 78 | CCBL1 | 15551 | -1.504 | -0.2172 | Yes | ||
| 79 | INPPL1 | 15679 | -1.538 | -0.2126 | Yes | ||
| 80 | DAB1 | 16127 | -1.663 | -0.2243 | Yes | ||
| 81 | ATP5G2 | 16259 | -1.698 | -0.2187 | Yes | ||
| 82 | BRUNOL4 | 16292 | -1.707 | -0.2078 | Yes | ||
| 83 | HOXB8 | 16370 | -1.734 | -0.1990 | Yes | ||
| 84 | EGR1 | 16405 | -1.746 | -0.1878 | Yes | ||
| 85 | GRASP | 16442 | -1.760 | -0.1767 | Yes | ||
| 86 | SLC25A10 | 16486 | -1.775 | -0.1658 | Yes | ||
| 87 | CDKN2C | 16578 | -1.811 | -0.1572 | Yes | ||
| 88 | LRP5 | 16644 | -1.834 | -0.1471 | Yes | ||
| 89 | PEX5 | 16771 | -1.887 | -0.1398 | Yes | ||
| 90 | FYN | 17088 | -2.012 | -0.1419 | Yes | ||
| 91 | RPL10 | 17126 | -2.030 | -0.1288 | Yes | ||
| 92 | RPP21 | 17153 | -2.036 | -0.1150 | Yes | ||
| 93 | LDHD | 17195 | -2.055 | -0.1019 | Yes | ||
| 94 | CTDSP1 | 17339 | -2.121 | -0.0939 | Yes | ||
| 95 | HTR2C | 17552 | -2.244 | -0.0886 | Yes | ||
| 96 | GGA1 | 18049 | -2.648 | -0.0957 | Yes | ||
| 97 | VASP | 18094 | -2.698 | -0.0779 | Yes | ||
| 98 | CAMK2D | 18184 | -2.810 | -0.0618 | Yes | ||
| 99 | ETV5 | 18434 | -3.305 | -0.0507 | Yes | ||
| 100 | GNAQ | 18552 | -3.955 | -0.0275 | Yes | ||
| 101 | ETV6 | 18576 | -4.153 | 0.0022 | Yes |
