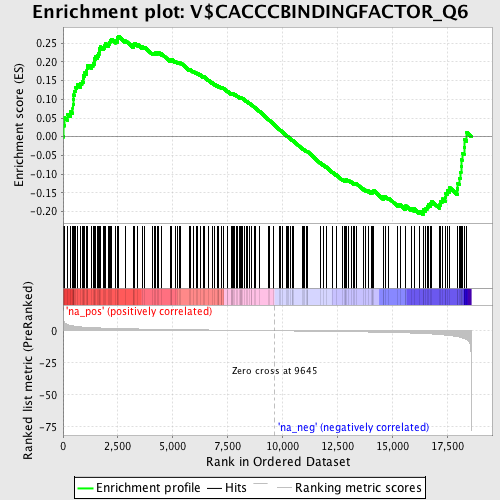
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_wtNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$CACCCBINDINGFACTOR_Q6 |
| Enrichment Score (ES) | 0.26917398 |
| Normalized Enrichment Score (NES) | 1.2085172 |
| Nominal p-value | 0.06918239 |
| FDR q-value | 0.5706085 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EIF5A | 17 | 8.994 | 0.0304 | Yes | ||
| 2 | PCBP4 | 52 | 6.597 | 0.0515 | Yes | ||
| 3 | ATP5G1 | 202 | 4.726 | 0.0599 | Yes | ||
| 4 | OTUB1 | 335 | 4.183 | 0.0673 | Yes | ||
| 5 | SFRS5 | 446 | 3.781 | 0.0745 | Yes | ||
| 6 | CDK4 | 450 | 3.776 | 0.0875 | Yes | ||
| 7 | DLD | 461 | 3.746 | 0.1000 | Yes | ||
| 8 | SLC6A9 | 484 | 3.705 | 0.1117 | Yes | ||
| 9 | EFEMP2 | 523 | 3.588 | 0.1222 | Yes | ||
| 10 | EIF4E | 566 | 3.497 | 0.1321 | Yes | ||
| 11 | SMYD5 | 632 | 3.322 | 0.1401 | Yes | ||
| 12 | EPHB2 | 786 | 3.039 | 0.1424 | Yes | ||
| 13 | EEF1A1 | 861 | 2.912 | 0.1485 | Yes | ||
| 14 | MYO1C | 919 | 2.836 | 0.1553 | Yes | ||
| 15 | NLN | 931 | 2.821 | 0.1646 | Yes | ||
| 16 | KCNAB3 | 991 | 2.755 | 0.1709 | Yes | ||
| 17 | YTHDF2 | 1071 | 2.684 | 0.1760 | Yes | ||
| 18 | CDKN1A | 1088 | 2.662 | 0.1844 | Yes | ||
| 19 | OTX1 | 1122 | 2.622 | 0.1918 | Yes | ||
| 20 | YARS | 1288 | 2.461 | 0.1914 | Yes | ||
| 21 | RGMA | 1364 | 2.390 | 0.1956 | Yes | ||
| 22 | BRUNOL4 | 1416 | 2.346 | 0.2010 | Yes | ||
| 23 | EPHA7 | 1427 | 2.337 | 0.2086 | Yes | ||
| 24 | FKBP2 | 1480 | 2.298 | 0.2138 | Yes | ||
| 25 | SDC1 | 1559 | 2.248 | 0.2174 | Yes | ||
| 26 | SLC25A13 | 1606 | 2.217 | 0.2227 | Yes | ||
| 27 | PPP1R1B | 1640 | 2.200 | 0.2285 | Yes | ||
| 28 | CRK | 1653 | 2.190 | 0.2355 | Yes | ||
| 29 | LRP1 | 1700 | 2.158 | 0.2405 | Yes | ||
| 30 | TIAL1 | 1850 | 2.070 | 0.2397 | Yes | ||
| 31 | BMF | 1901 | 2.039 | 0.2440 | Yes | ||
| 32 | HOXC6 | 1944 | 2.013 | 0.2488 | Yes | ||
| 33 | HMGN2 | 2089 | 1.941 | 0.2477 | Yes | ||
| 34 | GFAP | 2114 | 1.930 | 0.2531 | Yes | ||
| 35 | DLX1 | 2142 | 1.913 | 0.2583 | Yes | ||
| 36 | PMP22 | 2212 | 1.878 | 0.2611 | Yes | ||
| 37 | BLMH | 2408 | 1.793 | 0.2568 | Yes | ||
| 38 | OTX2 | 2480 | 1.750 | 0.2591 | Yes | ||
| 39 | MAPT | 2484 | 1.749 | 0.2650 | Yes | ||
| 40 | SEMA4C | 2519 | 1.731 | 0.2692 | Yes | ||
| 41 | HR | 2858 | 1.590 | 0.2564 | No | ||
| 42 | STAG2 | 3198 | 1.469 | 0.2431 | No | ||
| 43 | UBE2R2 | 3206 | 1.466 | 0.2478 | No | ||
| 44 | PLTP | 3266 | 1.445 | 0.2497 | No | ||
| 45 | CASKIN2 | 3399 | 1.397 | 0.2474 | No | ||
| 46 | SPRY2 | 3611 | 1.332 | 0.2406 | No | ||
| 47 | FLNC | 3717 | 1.300 | 0.2394 | No | ||
| 48 | ENTPD2 | 4085 | 1.189 | 0.2236 | No | ||
| 49 | SH3GLB2 | 4160 | 1.170 | 0.2237 | No | ||
| 50 | SULF1 | 4222 | 1.152 | 0.2244 | No | ||
| 51 | TRIM33 | 4287 | 1.137 | 0.2249 | No | ||
| 52 | MRPL14 | 4353 | 1.125 | 0.2253 | No | ||
| 53 | LBX1 | 4485 | 1.089 | 0.2220 | No | ||
| 54 | RTKN | 4891 | 0.994 | 0.2035 | No | ||
| 55 | KCNH8 | 4936 | 0.983 | 0.2045 | No | ||
| 56 | ZADH2 | 4948 | 0.981 | 0.2073 | No | ||
| 57 | INHA | 5121 | 0.939 | 0.2013 | No | ||
| 58 | GRK5 | 5209 | 0.915 | 0.1998 | No | ||
| 59 | SLC12A5 | 5287 | 0.895 | 0.1987 | No | ||
| 60 | SIX4 | 5366 | 0.872 | 0.1975 | No | ||
| 61 | RARG | 5744 | 0.786 | 0.1798 | No | ||
| 62 | MORF4L2 | 5791 | 0.776 | 0.1800 | No | ||
| 63 | PLS3 | 5922 | 0.750 | 0.1756 | No | ||
| 64 | DUSP14 | 6059 | 0.718 | 0.1707 | No | ||
| 65 | HOXB8 | 6120 | 0.707 | 0.1699 | No | ||
| 66 | ABCG4 | 6241 | 0.681 | 0.1658 | No | ||
| 67 | AMBN | 6378 | 0.650 | 0.1606 | No | ||
| 68 | PRKAG1 | 6428 | 0.642 | 0.1602 | No | ||
| 69 | XPO1 | 6628 | 0.603 | 0.1515 | No | ||
| 70 | GEMIN7 | 6800 | 0.569 | 0.1442 | No | ||
| 71 | MIP | 6915 | 0.545 | 0.1400 | No | ||
| 72 | HES1 | 7013 | 0.526 | 0.1365 | No | ||
| 73 | KCNN4 | 7087 | 0.512 | 0.1343 | No | ||
| 74 | ZFX | 7101 | 0.510 | 0.1354 | No | ||
| 75 | HOXC4 | 7214 | 0.484 | 0.1310 | No | ||
| 76 | CPLX1 | 7234 | 0.478 | 0.1317 | No | ||
| 77 | KCNH7 | 7297 | 0.465 | 0.1299 | No | ||
| 78 | EFNA3 | 7488 | 0.427 | 0.1211 | No | ||
| 79 | JARID2 | 7512 | 0.423 | 0.1213 | No | ||
| 80 | BMP2 | 7679 | 0.391 | 0.1137 | No | ||
| 81 | SERPINE1 | 7689 | 0.390 | 0.1146 | No | ||
| 82 | SOX15 | 7708 | 0.386 | 0.1149 | No | ||
| 83 | PTPRJ | 7716 | 0.384 | 0.1159 | No | ||
| 84 | CNKSR2 | 7767 | 0.375 | 0.1145 | No | ||
| 85 | LGI1 | 7830 | 0.360 | 0.1124 | No | ||
| 86 | RGS6 | 7887 | 0.347 | 0.1105 | No | ||
| 87 | SGTB | 7960 | 0.334 | 0.1078 | No | ||
| 88 | PCF11 | 8060 | 0.313 | 0.1035 | No | ||
| 89 | JPH4 | 8064 | 0.312 | 0.1044 | No | ||
| 90 | LGI4 | 8105 | 0.302 | 0.1033 | No | ||
| 91 | PLA2G6 | 8114 | 0.301 | 0.1039 | No | ||
| 92 | FGF11 | 8151 | 0.293 | 0.1030 | No | ||
| 93 | MID2 | 8187 | 0.286 | 0.1021 | No | ||
| 94 | LTBR | 8282 | 0.264 | 0.0979 | No | ||
| 95 | LTB4R | 8347 | 0.254 | 0.0954 | No | ||
| 96 | F11R | 8388 | 0.248 | 0.0940 | No | ||
| 97 | GSH1 | 8500 | 0.224 | 0.0888 | No | ||
| 98 | WNT3 | 8575 | 0.210 | 0.0855 | No | ||
| 99 | PVRL4 | 8592 | 0.206 | 0.0854 | No | ||
| 100 | RAD23B | 8738 | 0.176 | 0.0781 | No | ||
| 101 | CALM1 | 8771 | 0.170 | 0.0770 | No | ||
| 102 | CHRM1 | 8940 | 0.136 | 0.0683 | No | ||
| 103 | NINJ2 | 9365 | 0.057 | 0.0456 | No | ||
| 104 | LAD1 | 9410 | 0.047 | 0.0433 | No | ||
| 105 | TAGLN | 9588 | 0.010 | 0.0338 | No | ||
| 106 | YWHAE | 9858 | -0.041 | 0.0193 | No | ||
| 107 | KCND1 | 9894 | -0.048 | 0.0176 | No | ||
| 108 | MTSS1 | 9994 | -0.070 | 0.0125 | No | ||
| 109 | PTGDS | 10182 | -0.107 | 0.0027 | No | ||
| 110 | ETV5 | 10212 | -0.112 | 0.0015 | No | ||
| 111 | NRG1 | 10265 | -0.122 | -0.0009 | No | ||
| 112 | IGFBP5 | 10363 | -0.140 | -0.0057 | No | ||
| 113 | KREMEN2 | 10454 | -0.159 | -0.0100 | No | ||
| 114 | MAGEL2 | 10474 | -0.162 | -0.0105 | No | ||
| 115 | TUSC3 | 10481 | -0.163 | -0.0102 | No | ||
| 116 | STMN4 | 10899 | -0.257 | -0.0319 | No | ||
| 117 | BOC | 10939 | -0.263 | -0.0331 | No | ||
| 118 | LHX9 | 11015 | -0.281 | -0.0362 | No | ||
| 119 | HIF1A | 11096 | -0.298 | -0.0395 | No | ||
| 120 | FARP1 | 11124 | -0.304 | -0.0399 | No | ||
| 121 | PI15 | 11139 | -0.307 | -0.0396 | No | ||
| 122 | PORCN | 11716 | -0.433 | -0.0693 | No | ||
| 123 | RTN2 | 11861 | -0.464 | -0.0755 | No | ||
| 124 | CRLF1 | 11990 | -0.485 | -0.0808 | No | ||
| 125 | HOXB5 | 12281 | -0.553 | -0.0946 | No | ||
| 126 | PRDM16 | 12454 | -0.595 | -0.1019 | No | ||
| 127 | TBL1X | 12729 | -0.658 | -0.1144 | No | ||
| 128 | STK16 | 12834 | -0.682 | -0.1177 | No | ||
| 129 | MLLT7 | 12884 | -0.694 | -0.1179 | No | ||
| 130 | ZHX2 | 12888 | -0.694 | -0.1157 | No | ||
| 131 | GNB2 | 12900 | -0.697 | -0.1138 | No | ||
| 132 | CALCR | 12994 | -0.717 | -0.1164 | No | ||
| 133 | ADRA2C | 13129 | -0.755 | -0.1210 | No | ||
| 134 | PABPN1 | 13250 | -0.788 | -0.1248 | No | ||
| 135 | FOXF2 | 13300 | -0.801 | -0.1246 | No | ||
| 136 | CHD4 | 13352 | -0.812 | -0.1246 | No | ||
| 137 | RARB | 13689 | -0.896 | -0.1397 | No | ||
| 138 | HPN | 13803 | -0.929 | -0.1426 | No | ||
| 139 | HOXB9 | 13908 | -0.960 | -0.1449 | No | ||
| 140 | JUP | 14055 | -1.000 | -0.1493 | No | ||
| 141 | GRM3 | 14085 | -1.010 | -0.1474 | No | ||
| 142 | C1S | 14120 | -1.019 | -0.1457 | No | ||
| 143 | LIN7B | 14162 | -1.034 | -0.1443 | No | ||
| 144 | GNAO1 | 14595 | -1.164 | -0.1637 | No | ||
| 145 | USP25 | 14608 | -1.167 | -0.1602 | No | ||
| 146 | SHH | 14695 | -1.197 | -0.1607 | No | ||
| 147 | RBMS3 | 14849 | -1.256 | -0.1647 | No | ||
| 148 | ADCY5 | 15243 | -1.417 | -0.1810 | No | ||
| 149 | SEMA3B | 15355 | -1.475 | -0.1819 | No | ||
| 150 | MNT | 15615 | -1.616 | -0.1903 | No | ||
| 151 | LHX4 | 15624 | -1.620 | -0.1851 | No | ||
| 152 | TRPV3 | 15867 | -1.757 | -0.1921 | No | ||
| 153 | SLC9A3R1 | 15998 | -1.842 | -0.1928 | No | ||
| 154 | MEIS1 | 16251 | -2.021 | -0.1994 | No | ||
| 155 | SP4 | 16413 | -2.149 | -0.2006 | No | ||
| 156 | CNTFR | 16423 | -2.159 | -0.1936 | No | ||
| 157 | ARL6IP2 | 16531 | -2.238 | -0.1916 | No | ||
| 158 | CAMTA2 | 16588 | -2.291 | -0.1867 | No | ||
| 159 | LIMK2 | 16635 | -2.331 | -0.1811 | No | ||
| 160 | CITED2 | 16757 | -2.461 | -0.1790 | No | ||
| 161 | SPAG9 | 16798 | -2.502 | -0.1725 | No | ||
| 162 | DCTN1 | 17172 | -2.957 | -0.1824 | No | ||
| 163 | LDB1 | 17198 | -2.997 | -0.1733 | No | ||
| 164 | PIP5K2B | 17272 | -3.111 | -0.1665 | No | ||
| 165 | SPRY4 | 17419 | -3.351 | -0.1627 | No | ||
| 166 | ITGA5 | 17440 | -3.396 | -0.1520 | No | ||
| 167 | ASB7 | 17511 | -3.528 | -0.1435 | No | ||
| 168 | DIABLO | 17625 | -3.729 | -0.1366 | No | ||
| 169 | CREM | 17966 | -4.594 | -0.1391 | No | ||
| 170 | PHF12 | 17991 | -4.670 | -0.1241 | No | ||
| 171 | KLF13 | 18072 | -4.938 | -0.1112 | No | ||
| 172 | FBS1 | 18099 | -5.050 | -0.0951 | No | ||
| 173 | ITGB4 | 18137 | -5.231 | -0.0788 | No | ||
| 174 | IL2RG | 18158 | -5.351 | -0.0613 | No | ||
| 175 | BRD2 | 18187 | -5.490 | -0.0437 | No | ||
| 176 | BCL11A | 18279 | -6.014 | -0.0277 | No | ||
| 177 | DYRK1A | 18303 | -6.167 | -0.0074 | No | ||
| 178 | RGS14 | 18400 | -6.996 | 0.0117 | No |
