Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_wtNotch_versus_absentNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$MYCMAX_03 |
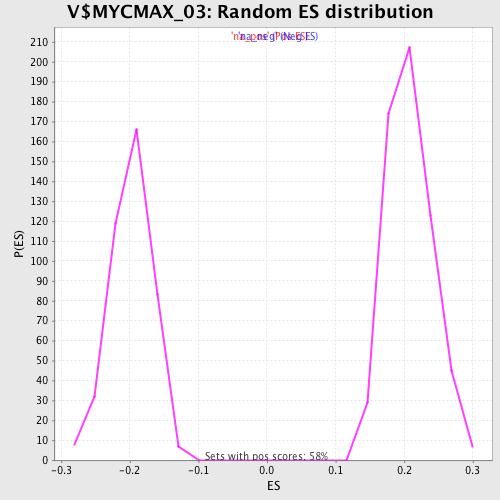
| Enrichment Score (ES) | -0.34560028 |
| Normalized Enrichment Score (NES) | -1.7472054 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.4227556 |
| FWER p-Value | 0.296 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EIF4E | 124 | 2.878 | 0.0097 | No | ||
| 2 | HOXB4 | 525 | 2.164 | 0.0003 | No | ||
| 3 | PTGES2 | 563 | 2.131 | 0.0105 | No | ||
| 4 | ANKRD17 | 664 | 2.027 | 0.0166 | No | ||
| 5 | AP1S2 | 763 | 1.959 | 0.0225 | No | ||
| 6 | LEF1 | 932 | 1.857 | 0.0239 | No | ||
| 7 | TAC1 | 1033 | 1.798 | 0.0288 | No | ||
| 8 | KCNK5 | 1185 | 1.720 | 0.0304 | No | ||
| 9 | FBXL19 | 1299 | 1.672 | 0.0338 | No | ||
| 10 | DUSP7 | 1433 | 1.619 | 0.0358 | No | ||
| 11 | ARMET | 1678 | 1.519 | 0.0312 | No | ||
| 12 | IRS4 | 1877 | 1.444 | 0.0287 | No | ||
| 13 | SOCS5 | 2401 | 1.269 | 0.0076 | No | ||
| 14 | PLCG2 | 2408 | 1.267 | 0.0145 | No | ||
| 15 | GATA5 | 2411 | 1.266 | 0.0216 | No | ||
| 16 | NR0B2 | 2727 | 1.183 | 0.0113 | No | ||
| 17 | HOXA9 | 2843 | 1.151 | 0.0116 | No | ||
| 18 | OPRS1 | 2930 | 1.127 | 0.0134 | No | ||
| 19 | SMYD4 | 2954 | 1.120 | 0.0185 | No | ||
| 20 | ZCCHC7 | 3071 | 1.092 | 0.0185 | No | ||
| 21 | TGIF2 | 3088 | 1.087 | 0.0238 | No | ||
| 22 | SEC23IP | 3124 | 1.077 | 0.0280 | No | ||
| 23 | TESK2 | 3218 | 1.057 | 0.0290 | No | ||
| 24 | USP15 | 3506 | 0.989 | 0.0191 | No | ||
| 25 | NRIP3 | 3514 | 0.987 | 0.0243 | No | ||
| 26 | NEUROD1 | 3859 | 0.909 | 0.0109 | No | ||
| 27 | TOM1 | 4085 | 0.864 | 0.0036 | No | ||
| 28 | GNAS | 4160 | 0.852 | 0.0044 | No | ||
| 29 | CCAR1 | 4166 | 0.851 | 0.0090 | No | ||
| 30 | ATF4 | 4469 | 0.794 | -0.0028 | No | ||
| 31 | SEPT3 | 4559 | 0.776 | -0.0032 | No | ||
| 32 | B3GALT2 | 4635 | 0.760 | -0.0030 | No | ||
| 33 | ISGF3G | 4702 | 0.748 | -0.0023 | No | ||
| 34 | TEF | 4825 | 0.727 | -0.0047 | No | ||
| 35 | HOXB7 | 4879 | 0.717 | -0.0035 | No | ||
| 36 | KRTCAP2 | 4948 | 0.705 | -0.0032 | No | ||
| 37 | RNF146 | 5037 | 0.689 | -0.0040 | No | ||
| 38 | VPS33A | 5044 | 0.687 | -0.0004 | No | ||
| 39 | SNX16 | 5217 | 0.655 | -0.0060 | No | ||
| 40 | HOXA1 | 5318 | 0.639 | -0.0078 | No | ||
| 41 | ILF3 | 5441 | 0.616 | -0.0109 | No | ||
| 42 | RGL1 | 5528 | 0.599 | -0.0122 | No | ||
| 43 | NDUFA7 | 5657 | 0.578 | -0.0158 | No | ||
| 44 | MRPL27 | 5692 | 0.574 | -0.0144 | No | ||
| 45 | ZIC1 | 5736 | 0.568 | -0.0135 | No | ||
| 46 | PRPS1 | 6317 | 0.469 | -0.0422 | No | ||
| 47 | DLX2 | 6352 | 0.463 | -0.0415 | No | ||
| 48 | SLC1A7 | 6452 | 0.446 | -0.0443 | No | ||
| 49 | SNX5 | 6471 | 0.444 | -0.0427 | No | ||
| 50 | MYST3 | 6535 | 0.434 | -0.0437 | No | ||
| 51 | PLAGL1 | 6588 | 0.425 | -0.0441 | No | ||
| 52 | NFX1 | 6649 | 0.416 | -0.0449 | No | ||
| 53 | CDC14A | 6712 | 0.405 | -0.0460 | No | ||
| 54 | MEOX2 | 6877 | 0.377 | -0.0527 | No | ||
| 55 | CDKN2C | 7076 | 0.347 | -0.0615 | No | ||
| 56 | COPZ1 | 7077 | 0.347 | -0.0595 | No | ||
| 57 | PPRC1 | 7084 | 0.345 | -0.0579 | No | ||
| 58 | RELB | 7112 | 0.340 | -0.0574 | No | ||
| 59 | SCYL1 | 7247 | 0.318 | -0.0628 | No | ||
| 60 | UBE2B | 7349 | 0.300 | -0.0666 | No | ||
| 61 | EIF3S10 | 7573 | 0.267 | -0.0772 | No | ||
| 62 | RPS28 | 7659 | 0.255 | -0.0803 | No | ||
| 63 | HOXA7 | 7839 | 0.230 | -0.0887 | No | ||
| 64 | HNRPD | 7887 | 0.223 | -0.0900 | No | ||
| 65 | ELAVL3 | 8050 | 0.196 | -0.0977 | No | ||
| 66 | TAF6L | 8292 | 0.161 | -0.1098 | No | ||
| 67 | BMP6 | 8615 | 0.106 | -0.1267 | No | ||
| 68 | ODC1 | 8622 | 0.105 | -0.1264 | No | ||
| 69 | SLCO1C1 | 8807 | 0.076 | -0.1359 | No | ||
| 70 | CHST11 | 8834 | 0.073 | -0.1369 | No | ||
| 71 | HOXD10 | 9005 | 0.048 | -0.1459 | No | ||
| 72 | EIF4G1 | 9140 | 0.028 | -0.1530 | No | ||
| 73 | OPRD1 | 9257 | 0.011 | -0.1592 | No | ||
| 74 | SLC38A5 | 9314 | 0.001 | -0.1622 | No | ||
| 75 | HOXB5 | 9351 | -0.006 | -0.1642 | No | ||
| 76 | LIN28 | 9535 | -0.032 | -0.1739 | No | ||
| 77 | XPO1 | 9942 | -0.106 | -0.1953 | No | ||
| 78 | PICALM | 10336 | -0.162 | -0.2157 | No | ||
| 79 | HOXC5 | 10534 | -0.191 | -0.2253 | No | ||
| 80 | PCDHA10 | 10774 | -0.231 | -0.2369 | No | ||
| 81 | POU3F2 | 11143 | -0.290 | -0.2552 | No | ||
| 82 | SYT3 | 11209 | -0.301 | -0.2570 | No | ||
| 83 | SIRT1 | 11267 | -0.307 | -0.2584 | No | ||
| 84 | HTATIP | 11272 | -0.308 | -0.2568 | No | ||
| 85 | GPC3 | 11274 | -0.309 | -0.2551 | No | ||
| 86 | MCM2 | 11323 | -0.316 | -0.2559 | No | ||
| 87 | TRIM37 | 11692 | -0.364 | -0.2738 | No | ||
| 88 | LIG3 | 11734 | -0.370 | -0.2739 | No | ||
| 89 | RPA1 | 11768 | -0.375 | -0.2736 | No | ||
| 90 | CBX8 | 11975 | -0.406 | -0.2824 | No | ||
| 91 | CPT1A | 12538 | -0.489 | -0.3101 | No | ||
| 92 | TOPORS | 12707 | -0.517 | -0.3163 | No | ||
| 93 | HOXA11 | 12763 | -0.524 | -0.3163 | No | ||
| 94 | SHMT1 | 12770 | -0.525 | -0.3136 | No | ||
| 95 | UTY | 12782 | -0.527 | -0.3112 | No | ||
| 96 | RBBP4 | 12833 | -0.536 | -0.3108 | No | ||
| 97 | GABARAP | 12894 | -0.548 | -0.3110 | No | ||
| 98 | PAX6 | 13085 | -0.576 | -0.3180 | No | ||
| 99 | ZNF318 | 13128 | -0.582 | -0.3169 | No | ||
| 100 | BRD2 | 13136 | -0.583 | -0.3140 | No | ||
| 101 | BMP7 | 13186 | -0.591 | -0.3133 | No | ||
| 102 | BAX | 13218 | -0.596 | -0.3116 | No | ||
| 103 | DNMT3A | 13247 | -0.602 | -0.3097 | No | ||
| 104 | ATP6V1C1 | 13283 | -0.610 | -0.3081 | No | ||
| 105 | NPTX1 | 13761 | -0.694 | -0.3300 | No | ||
| 106 | PTMA | 13857 | -0.712 | -0.3311 | No | ||
| 107 | TRIB1 | 14126 | -0.760 | -0.3413 | Yes | ||
| 108 | UBXD1 | 14135 | -0.761 | -0.3374 | Yes | ||
| 109 | CBARA1 | 14162 | -0.766 | -0.3344 | Yes | ||
| 110 | NIT1 | 14218 | -0.776 | -0.3330 | Yes | ||
| 111 | CD164 | 14336 | -0.800 | -0.3347 | Yes | ||
| 112 | TIMM8A | 14473 | -0.825 | -0.3374 | Yes | ||
| 113 | GGN | 14555 | -0.841 | -0.3370 | Yes | ||
| 114 | FMR1 | 14625 | -0.854 | -0.3359 | Yes | ||
| 115 | DRD1 | 14649 | -0.860 | -0.3322 | Yes | ||
| 116 | TBC1D15 | 14772 | -0.885 | -0.3338 | Yes | ||
| 117 | SYT6 | 14907 | -0.914 | -0.3359 | Yes | ||
| 118 | UTX | 15000 | -0.933 | -0.3355 | Yes | ||
| 119 | IGF2R | 15091 | -0.952 | -0.3350 | Yes | ||
| 120 | UBE4B | 15107 | -0.954 | -0.3303 | Yes | ||
| 121 | AKAP1 | 15335 | -1.001 | -0.3370 | Yes | ||
| 122 | HNRPA1 | 15361 | -1.006 | -0.3326 | Yes | ||
| 123 | TGFB2 | 15506 | -1.035 | -0.3345 | Yes | ||
| 124 | CUL5 | 15601 | -1.056 | -0.3336 | Yes | ||
| 125 | DDX3X | 15680 | -1.071 | -0.3317 | Yes | ||
| 126 | VPS16 | 15719 | -1.080 | -0.3276 | Yes | ||
| 127 | FKBP11 | 15858 | -1.116 | -0.3287 | Yes | ||
| 128 | NEUROD2 | 15880 | -1.124 | -0.3234 | Yes | ||
| 129 | CELSR3 | 15909 | -1.131 | -0.3185 | Yes | ||
| 130 | SNCAIP | 15949 | -1.142 | -0.3141 | Yes | ||
| 131 | STMN1 | 15992 | -1.152 | -0.3098 | Yes | ||
| 132 | NPM1 | 15993 | -1.153 | -0.3032 | Yes | ||
| 133 | CITED2 | 16062 | -1.168 | -0.3003 | Yes | ||
| 134 | MAT2A | 16104 | -1.177 | -0.2958 | Yes | ||
| 135 | RAB3IL1 | 16222 | -1.208 | -0.2952 | Yes | ||
| 136 | UBXD3 | 16312 | -1.236 | -0.2930 | Yes | ||
| 137 | UTP14A | 16367 | -1.248 | -0.2888 | Yes | ||
| 138 | SLC38A2 | 16430 | -1.262 | -0.2850 | Yes | ||
| 139 | CBX5 | 16548 | -1.294 | -0.2840 | Yes | ||
| 140 | PABPC1 | 16657 | -1.327 | -0.2822 | Yes | ||
| 141 | CEBPB | 16737 | -1.354 | -0.2788 | Yes | ||
| 142 | HEXA | 16785 | -1.368 | -0.2736 | Yes | ||
| 143 | COMMD3 | 16815 | -1.377 | -0.2673 | Yes | ||
| 144 | CHD4 | 16895 | -1.401 | -0.2636 | Yes | ||
| 145 | RPL13A | 16978 | -1.429 | -0.2599 | Yes | ||
| 146 | ALDH6A1 | 17024 | -1.445 | -0.2541 | Yes | ||
| 147 | QTRTD1 | 17145 | -1.492 | -0.2521 | Yes | ||
| 148 | GADD45G | 17175 | -1.504 | -0.2451 | Yes | ||
| 149 | PPM1A | 17318 | -1.569 | -0.2438 | Yes | ||
| 150 | ARPC5 | 17481 | -1.634 | -0.2433 | Yes | ||
| 151 | LAMP1 | 17606 | -1.697 | -0.2403 | Yes | ||
| 152 | H3F3A | 17616 | -1.701 | -0.2311 | Yes | ||
| 153 | CUGBP1 | 17737 | -1.767 | -0.2276 | Yes | ||
| 154 | DCTN4 | 17805 | -1.806 | -0.2209 | Yes | ||
| 155 | SYNCRIP | 17816 | -1.816 | -0.2111 | Yes | ||
| 156 | EIF4B | 17836 | -1.829 | -0.2017 | Yes | ||
| 157 | ADAM12 | 18040 | -1.990 | -0.2013 | Yes | ||
| 158 | PABPC4 | 18152 | -2.106 | -0.1954 | Yes | ||
| 159 | GJA1 | 18174 | -2.135 | -0.1843 | Yes | ||
| 160 | TNRC15 | 18202 | -2.165 | -0.1734 | Yes | ||
| 161 | HMOX1 | 18249 | -2.228 | -0.1632 | Yes | ||
| 162 | KBTBD2 | 18334 | -2.357 | -0.1544 | Yes | ||
| 163 | PFDN2 | 18360 | -2.392 | -0.1421 | Yes | ||
| 164 | PER1 | 18385 | -2.454 | -0.1294 | Yes | ||
| 165 | SSR1 | 18396 | -2.475 | -0.1158 | Yes | ||
| 166 | NR1D1 | 18454 | -2.653 | -0.1038 | Yes | ||
| 167 | GTF2A1 | 18458 | -2.667 | -0.0887 | Yes | ||
| 168 | UCHL1 | 18518 | -2.887 | -0.0755 | Yes | ||
| 169 | EPC1 | 18540 | -2.994 | -0.0596 | Yes | ||
| 170 | BOK | 18569 | -3.165 | -0.0430 | Yes | ||
| 171 | CTSF | 18599 | -3.667 | -0.0237 | Yes | ||
| 172 | ALDH3B1 | 18614 | -4.308 | 0.0001 | Yes |