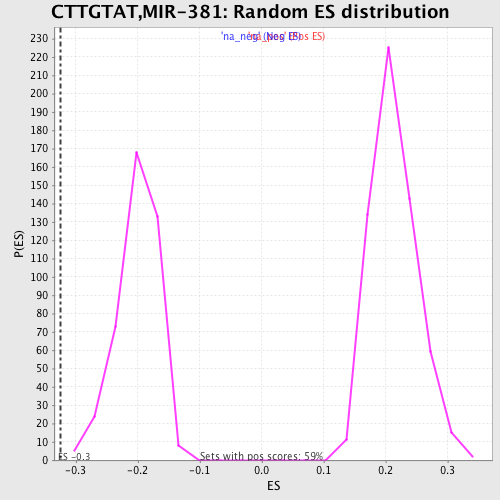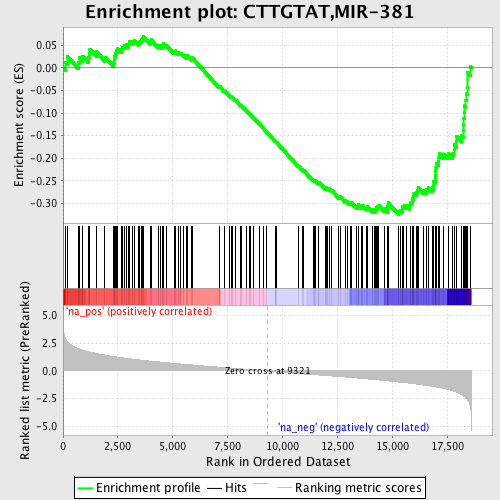
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_wtNotch_versus_absentNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | CTTGTAT,MIR-381 |
| Enrichment Score (ES) | -0.3241633 |
| Normalized Enrichment Score (NES) | -1.6095963 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.5515413 |
| FWER p-Value | 0.745 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RALGPS1 | 123 | 2.882 | 0.0120 | No | ||
| 2 | CACNA1C | 196 | 2.639 | 0.0252 | No | ||
| 3 | NRXN1 | 697 | 2.006 | 0.0111 | No | ||
| 4 | SIPA1L2 | 732 | 1.986 | 0.0221 | No | ||
| 5 | RBM16 | 895 | 1.876 | 0.0254 | No | ||
| 6 | PAPOLG | 1141 | 1.741 | 0.0234 | No | ||
| 7 | HNRPH2 | 1202 | 1.712 | 0.0312 | No | ||
| 8 | STX12 | 1221 | 1.704 | 0.0413 | No | ||
| 9 | PURG | 1523 | 1.585 | 0.0352 | No | ||
| 10 | UBN1 | 1905 | 1.431 | 0.0239 | No | ||
| 11 | ROCK2 | 2293 | 1.303 | 0.0113 | No | ||
| 12 | GORASP2 | 2328 | 1.290 | 0.0178 | No | ||
| 13 | MBNL2 | 2332 | 1.288 | 0.0260 | No | ||
| 14 | ACTG1 | 2384 | 1.274 | 0.0315 | No | ||
| 15 | WNT5A | 2425 | 1.262 | 0.0375 | No | ||
| 16 | CCNT2 | 2492 | 1.242 | 0.0419 | No | ||
| 17 | AGPAT3 | 2660 | 1.202 | 0.0407 | No | ||
| 18 | UBE3A | 2704 | 1.190 | 0.0460 | No | ||
| 19 | GAS2 | 2782 | 1.167 | 0.0494 | No | ||
| 20 | ROD1 | 2878 | 1.140 | 0.0516 | No | ||
| 21 | MDS1 | 3002 | 1.109 | 0.0522 | No | ||
| 22 | CRSP2 | 3033 | 1.100 | 0.0577 | No | ||
| 23 | GRM8 | 3144 | 1.072 | 0.0586 | No | ||
| 24 | ATRN | 3242 | 1.050 | 0.0602 | No | ||
| 25 | BRD7 | 3453 | 1.001 | 0.0553 | No | ||
| 26 | LRRC4 | 3501 | 0.991 | 0.0591 | No | ||
| 27 | EPHA4 | 3563 | 0.974 | 0.0621 | No | ||
| 28 | PHF15 | 3621 | 0.962 | 0.0653 | No | ||
| 29 | QKI | 3646 | 0.957 | 0.0702 | No | ||
| 30 | NDUFAB1 | 3970 | 0.885 | 0.0584 | No | ||
| 31 | NFIA | 4013 | 0.878 | 0.0618 | No | ||
| 32 | BCL11A | 4328 | 0.819 | 0.0501 | No | ||
| 33 | XPO7 | 4452 | 0.797 | 0.0486 | No | ||
| 34 | DHX40 | 4545 | 0.779 | 0.0487 | No | ||
| 35 | YTHDF1 | 4552 | 0.779 | 0.0534 | No | ||
| 36 | NIN | 4697 | 0.748 | 0.0504 | No | ||
| 37 | NR5A2 | 5058 | 0.686 | 0.0354 | No | ||
| 38 | RAB11FIP2 | 5101 | 0.676 | 0.0375 | No | ||
| 39 | PSIP1 | 5249 | 0.649 | 0.0337 | No | ||
| 40 | NLK | 5345 | 0.634 | 0.0326 | No | ||
| 41 | DDX6 | 5497 | 0.603 | 0.0284 | No | ||
| 42 | GFRA1 | 5638 | 0.582 | 0.0246 | No | ||
| 43 | VSNL1 | 5665 | 0.577 | 0.0269 | No | ||
| 44 | SNRK | 5863 | 0.547 | 0.0197 | No | ||
| 45 | PBEF1 | 5886 | 0.543 | 0.0221 | No | ||
| 46 | SHANK2 | 7116 | 0.340 | -0.0423 | No | ||
| 47 | TMEM16D | 7118 | 0.339 | -0.0401 | No | ||
| 48 | RPS6KA3 | 7352 | 0.299 | -0.0508 | No | ||
| 49 | EIF3S10 | 7573 | 0.267 | -0.0610 | No | ||
| 50 | CGGBP1 | 7656 | 0.255 | -0.0638 | No | ||
| 51 | LZTS2 | 7703 | 0.247 | -0.0647 | No | ||
| 52 | AZIN1 | 7849 | 0.229 | -0.0711 | No | ||
| 53 | ACVR2A | 8075 | 0.193 | -0.0820 | No | ||
| 54 | ITCH | 8110 | 0.188 | -0.0826 | No | ||
| 55 | NAP1L5 | 8378 | 0.147 | -0.0961 | No | ||
| 56 | RHOQ | 8490 | 0.129 | -0.1013 | No | ||
| 57 | SSH2 | 8543 | 0.119 | -0.1034 | No | ||
| 58 | UBR1 | 8686 | 0.095 | -0.1104 | No | ||
| 59 | JAG2 | 8943 | 0.056 | -0.1239 | No | ||
| 60 | ST8SIA2 | 9112 | 0.032 | -0.1328 | No | ||
| 61 | MITF | 9271 | 0.008 | -0.1413 | No | ||
| 62 | PHF2 | 9678 | -0.057 | -0.1629 | No | ||
| 63 | RPL35A | 9695 | -0.062 | -0.1634 | No | ||
| 64 | ETF1 | 9724 | -0.067 | -0.1645 | No | ||
| 65 | SOX4 | 10722 | -0.220 | -0.2171 | No | ||
| 66 | UBE4A | 10900 | -0.253 | -0.2250 | No | ||
| 67 | PARD6B | 10975 | -0.264 | -0.2273 | No | ||
| 68 | RNF2 | 11400 | -0.324 | -0.2482 | No | ||
| 69 | EIF1 | 11452 | -0.330 | -0.2488 | No | ||
| 70 | PDGFC | 11498 | -0.337 | -0.2491 | No | ||
| 71 | GRM1 | 11644 | -0.356 | -0.2546 | No | ||
| 72 | ZCCHC11 | 11646 | -0.356 | -0.2524 | No | ||
| 73 | RAP2C | 11960 | -0.403 | -0.2667 | No | ||
| 74 | ASXL2 | 11981 | -0.407 | -0.2651 | No | ||
| 75 | ABCA1 | 12059 | -0.419 | -0.2666 | No | ||
| 76 | BHLHB5 | 12132 | -0.431 | -0.2677 | No | ||
| 77 | APRIN | 12244 | -0.447 | -0.2708 | No | ||
| 78 | PAPOLA | 12570 | -0.494 | -0.2852 | No | ||
| 79 | KIF1B | 12632 | -0.503 | -0.2853 | No | ||
| 80 | BTBD7 | 12852 | -0.540 | -0.2936 | No | ||
| 81 | TRIM63 | 12967 | -0.559 | -0.2962 | No | ||
| 82 | RPS6KB1 | 13079 | -0.575 | -0.2985 | No | ||
| 83 | ETS1 | 13124 | -0.581 | -0.2971 | No | ||
| 84 | RNF111 | 13391 | -0.628 | -0.3075 | No | ||
| 85 | FLRT3 | 13440 | -0.637 | -0.3059 | No | ||
| 86 | BIRC6 | 13456 | -0.640 | -0.3026 | No | ||
| 87 | KCTD3 | 13617 | -0.667 | -0.3070 | No | ||
| 88 | GAD1 | 13643 | -0.673 | -0.3040 | No | ||
| 89 | LPHN1 | 13834 | -0.709 | -0.3097 | No | ||
| 90 | CNOT7 | 13873 | -0.713 | -0.3071 | No | ||
| 91 | R3HDM1 | 14092 | -0.750 | -0.3141 | No | ||
| 92 | WEE1 | 14185 | -0.770 | -0.3141 | No | ||
| 93 | THRAP1 | 14251 | -0.784 | -0.3125 | No | ||
| 94 | PNRC1 | 14276 | -0.788 | -0.3087 | No | ||
| 95 | ARID4B | 14341 | -0.801 | -0.3070 | No | ||
| 96 | ZFPM2 | 14396 | -0.809 | -0.3047 | No | ||
| 97 | B3GNT5 | 14628 | -0.855 | -0.3117 | No | ||
| 98 | TBC1D15 | 14772 | -0.885 | -0.3137 | No | ||
| 99 | EPHA3 | 14795 | -0.890 | -0.3091 | No | ||
| 100 | RNF13 | 14805 | -0.892 | -0.3038 | No | ||
| 101 | PMP22 | 14811 | -0.893 | -0.2983 | No | ||
| 102 | CSTF3 | 15289 | -0.990 | -0.3178 | Yes | ||
| 103 | OSBPL3 | 15373 | -1.009 | -0.3157 | Yes | ||
| 104 | FOXF2 | 15452 | -1.025 | -0.3133 | Yes | ||
| 105 | ZMYND11 | 15466 | -1.028 | -0.3074 | Yes | ||
| 106 | PDAP1 | 15535 | -1.041 | -0.3043 | Yes | ||
| 107 | SRRM1 | 15666 | -1.067 | -0.3045 | Yes | ||
| 108 | THRAP2 | 15810 | -1.102 | -0.3051 | Yes | ||
| 109 | KHDRBS1 | 15820 | -1.104 | -0.2984 | Yes | ||
| 110 | EIF4G2 | 15903 | -1.129 | -0.2956 | Yes | ||
| 111 | SATB1 | 15911 | -1.132 | -0.2886 | Yes | ||
| 112 | SLC6A8 | 15971 | -1.148 | -0.2844 | Yes | ||
| 113 | EIF4G3 | 15990 | -1.152 | -0.2779 | Yes | ||
| 114 | UBE2E2 | 16086 | -1.174 | -0.2755 | Yes | ||
| 115 | INSIG1 | 16140 | -1.187 | -0.2706 | Yes | ||
| 116 | ZFAND3 | 16175 | -1.196 | -0.2648 | Yes | ||
| 117 | SLC38A2 | 16430 | -1.262 | -0.2703 | Yes | ||
| 118 | SLC24A3 | 16561 | -1.298 | -0.2690 | Yes | ||
| 119 | ACSL3 | 16630 | -1.319 | -0.2641 | Yes | ||
| 120 | BAT2 | 16814 | -1.376 | -0.2651 | Yes | ||
| 121 | SEMA6D | 16884 | -1.398 | -0.2598 | Yes | ||
| 122 | CHD4 | 16895 | -1.401 | -0.2513 | Yes | ||
| 123 | JPH1 | 16961 | -1.422 | -0.2456 | Yes | ||
| 124 | COL3A1 | 16974 | -1.426 | -0.2371 | Yes | ||
| 125 | APPBP2 | 16985 | -1.431 | -0.2284 | Yes | ||
| 126 | NFKBIA | 16992 | -1.433 | -0.2194 | Yes | ||
| 127 | KPNA3 | 17033 | -1.448 | -0.2122 | Yes | ||
| 128 | HNRPR | 17112 | -1.479 | -0.2069 | Yes | ||
| 129 | CLCF1 | 17130 | -1.486 | -0.1982 | Yes | ||
| 130 | BCL11B | 17148 | -1.494 | -0.1894 | Yes | ||
| 131 | STAG1 | 17357 | -1.586 | -0.1904 | Yes | ||
| 132 | KCTD15 | 17547 | -1.669 | -0.1899 | Yes | ||
| 133 | RNF144 | 17741 | -1.770 | -0.1889 | Yes | ||
| 134 | SYNCRIP | 17816 | -1.816 | -0.1811 | Yes | ||
| 135 | DLC1 | 17823 | -1.821 | -0.1697 | Yes | ||
| 136 | MTSS1 | 17929 | -1.892 | -0.1631 | Yes | ||
| 137 | HBP1 | 17938 | -1.898 | -0.1513 | Yes | ||
| 138 | GJA1 | 18174 | -2.135 | -0.1502 | Yes | ||
| 139 | ELOVL7 | 18230 | -2.203 | -0.1389 | Yes | ||
| 140 | CXCR4 | 18240 | -2.216 | -0.1251 | Yes | ||
| 141 | SEMA3C | 18266 | -2.253 | -0.1119 | Yes | ||
| 142 | KLF4 | 18281 | -2.277 | -0.0979 | Yes | ||
| 143 | KLF12 | 18302 | -2.309 | -0.0840 | Yes | ||
| 144 | ZFYVE20 | 18359 | -2.392 | -0.0716 | Yes | ||
| 145 | HIPK1 | 18386 | -2.457 | -0.0571 | Yes | ||
| 146 | GTF2I | 18423 | -2.530 | -0.0427 | Yes | ||
| 147 | CEBPA | 18434 | -2.555 | -0.0267 | Yes | ||
| 148 | SLMAP | 18435 | -2.557 | -0.0102 | Yes | ||
| 149 | ANKRD50 | 18561 | -3.082 | 0.0030 | Yes |