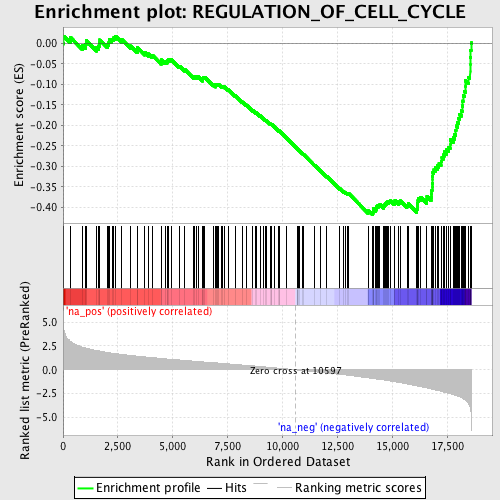
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | REGULATION_OF_CELL_CYCLE |
| Enrichment Score (ES) | -0.41760156 |
| Normalized Enrichment Score (NES) | -2.0618105 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.00827161 |
| FWER p-Value | 0.097 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CDK10 | 30 | 4.309 | 0.0177 | No | ||
| 2 | PLAGL1 | 338 | 2.962 | 0.0143 | No | ||
| 3 | ERCC2 | 889 | 2.361 | -0.0050 | No | ||
| 4 | TBRG4 | 1026 | 2.266 | -0.0022 | No | ||
| 5 | TBX3 | 1050 | 2.255 | 0.0067 | No | ||
| 6 | BMP7 | 1523 | 2.014 | -0.0099 | No | ||
| 7 | HPGD | 1623 | 1.974 | -0.0064 | No | ||
| 8 | PML | 1644 | 1.966 | 0.0013 | No | ||
| 9 | RAD9A | 1677 | 1.953 | 0.0083 | No | ||
| 10 | CDKN2C | 2038 | 1.811 | -0.0031 | No | ||
| 11 | ZBTB17 | 2086 | 1.792 | 0.0024 | No | ||
| 12 | NEK11 | 2098 | 1.788 | 0.0098 | No | ||
| 13 | MAPK12 | 2228 | 1.741 | 0.0106 | No | ||
| 14 | SMAD3 | 2315 | 1.711 | 0.0136 | No | ||
| 15 | MAD2L2 | 2384 | 1.691 | 0.0175 | No | ||
| 16 | NEK2 | 2671 | 1.615 | 0.0092 | No | ||
| 17 | CHMP1A | 3080 | 1.502 | -0.0062 | No | ||
| 18 | AIF1 | 3379 | 1.424 | -0.0160 | No | ||
| 19 | PPP1R13B | 3400 | 1.419 | -0.0107 | No | ||
| 20 | SPHK1 | 3731 | 1.344 | -0.0226 | No | ||
| 21 | UHMK1 | 3897 | 1.305 | -0.0257 | No | ||
| 22 | ING4 | 4069 | 1.275 | -0.0292 | No | ||
| 23 | CDKN1B | 4465 | 1.185 | -0.0453 | No | ||
| 24 | APBB1 | 4471 | 1.184 | -0.0403 | No | ||
| 25 | CUL2 | 4647 | 1.147 | -0.0446 | No | ||
| 26 | RASSF1 | 4735 | 1.128 | -0.0443 | No | ||
| 27 | RPRM | 4771 | 1.121 | -0.0412 | No | ||
| 28 | CDK5RAP1 | 4816 | 1.113 | -0.0386 | No | ||
| 29 | DDB1 | 4922 | 1.093 | -0.0394 | No | ||
| 30 | BTG4 | 5303 | 1.022 | -0.0554 | No | ||
| 31 | CUL5 | 5535 | 0.976 | -0.0635 | No | ||
| 32 | NEK6 | 5956 | 0.899 | -0.0823 | No | ||
| 33 | GADD45A | 5991 | 0.894 | -0.0801 | No | ||
| 34 | STK11 | 6087 | 0.876 | -0.0813 | No | ||
| 35 | TIMELESS | 6161 | 0.862 | -0.0814 | No | ||
| 36 | CITED2 | 6359 | 0.828 | -0.0884 | No | ||
| 37 | CDK5R2 | 6374 | 0.825 | -0.0855 | No | ||
| 38 | TGFB2 | 6387 | 0.822 | -0.0824 | No | ||
| 39 | CDK7 | 6460 | 0.809 | -0.0827 | No | ||
| 40 | CDKN2B | 6856 | 0.733 | -0.1008 | No | ||
| 41 | CDKN3 | 6964 | 0.719 | -0.1034 | No | ||
| 42 | CDK5RAP3 | 6970 | 0.719 | -0.1005 | No | ||
| 43 | HERC5 | 7029 | 0.709 | -0.1004 | No | ||
| 44 | RHOB | 7066 | 0.704 | -0.0992 | No | ||
| 45 | CD28 | 7225 | 0.675 | -0.1048 | No | ||
| 46 | APBB2 | 7274 | 0.667 | -0.1044 | No | ||
| 47 | FOXC1 | 7360 | 0.653 | -0.1061 | No | ||
| 48 | BMP2 | 7533 | 0.619 | -0.1126 | No | ||
| 49 | SNF1LK | 7841 | 0.559 | -0.1267 | No | ||
| 50 | PA2G4 | 8183 | 0.495 | -0.1430 | No | ||
| 51 | TGFB1 | 8358 | 0.463 | -0.1503 | No | ||
| 52 | RAD1 | 8641 | 0.409 | -0.1638 | No | ||
| 53 | CKS2 | 8780 | 0.384 | -0.1695 | No | ||
| 54 | ANAPC2 | 8811 | 0.381 | -0.1695 | No | ||
| 55 | PRMT5 | 8974 | 0.349 | -0.1767 | No | ||
| 56 | FANCG | 9112 | 0.321 | -0.1827 | No | ||
| 57 | SERTAD1 | 9221 | 0.294 | -0.1872 | No | ||
| 58 | RUNX3 | 9277 | 0.285 | -0.1889 | No | ||
| 59 | PPM1G | 9438 | 0.246 | -0.1965 | No | ||
| 60 | PPP1R9B | 9478 | 0.238 | -0.1975 | No | ||
| 61 | TP53 | 9507 | 0.232 | -0.1980 | No | ||
| 62 | RCC1 | 9622 | 0.206 | -0.2032 | No | ||
| 63 | CDKN2D | 9831 | 0.160 | -0.2138 | No | ||
| 64 | ALOX15B | 9840 | 0.159 | -0.2135 | No | ||
| 65 | INHA | 9860 | 0.156 | -0.2138 | No | ||
| 66 | BUB1 | 10189 | 0.087 | -0.2312 | No | ||
| 67 | UBE2C | 10662 | -0.017 | -0.2567 | No | ||
| 68 | CCND2 | 10685 | -0.024 | -0.2578 | No | ||
| 69 | JMY | 10699 | -0.026 | -0.2584 | No | ||
| 70 | GAS7 | 10744 | -0.037 | -0.2606 | No | ||
| 71 | DHRS2 | 10779 | -0.047 | -0.2622 | No | ||
| 72 | EREG | 10902 | -0.073 | -0.2685 | No | ||
| 73 | BTG3 | 10941 | -0.082 | -0.2702 | No | ||
| 74 | MAP2K6 | 11466 | -0.203 | -0.2977 | No | ||
| 75 | MLF1 | 11729 | -0.259 | -0.3107 | No | ||
| 76 | XPC | 12019 | -0.329 | -0.3249 | No | ||
| 77 | CCND3 | 12615 | -0.477 | -0.3550 | No | ||
| 78 | CHEK1 | 12792 | -0.517 | -0.3622 | No | ||
| 79 | BMP4 | 12869 | -0.538 | -0.3639 | No | ||
| 80 | MFN2 | 12979 | -0.570 | -0.3673 | No | ||
| 81 | CDC25A | 13018 | -0.579 | -0.3667 | No | ||
| 82 | CDC23 | 13900 | -0.834 | -0.4107 | No | ||
| 83 | CDC6 | 13931 | -0.843 | -0.4086 | No | ||
| 84 | MYC | 14099 | -0.902 | -0.4136 | Yes | ||
| 85 | ATM | 14127 | -0.911 | -0.4110 | Yes | ||
| 86 | BIRC5 | 14134 | -0.913 | -0.4072 | Yes | ||
| 87 | GAS1 | 14136 | -0.913 | -0.4032 | Yes | ||
| 88 | NPM2 | 14257 | -0.950 | -0.4054 | Yes | ||
| 89 | TTK | 14282 | -0.959 | -0.4024 | Yes | ||
| 90 | CDC37 | 14298 | -0.963 | -0.3989 | Yes | ||
| 91 | EPGN | 14336 | -0.975 | -0.3966 | Yes | ||
| 92 | CDK5R1 | 14394 | -0.995 | -0.3952 | Yes | ||
| 93 | ASNS | 14438 | -1.008 | -0.3930 | Yes | ||
| 94 | PTEN | 14609 | -1.063 | -0.3975 | Yes | ||
| 95 | CCNG1 | 14637 | -1.073 | -0.3941 | Yes | ||
| 96 | ERCC3 | 14676 | -1.088 | -0.3913 | Yes | ||
| 97 | DLG1 | 14724 | -1.106 | -0.3889 | Yes | ||
| 98 | CDC16 | 14779 | -1.128 | -0.3868 | Yes | ||
| 99 | KNTC1 | 14846 | -1.150 | -0.3852 | Yes | ||
| 100 | PCBP4 | 14926 | -1.177 | -0.3842 | Yes | ||
| 101 | TIPIN | 15091 | -1.244 | -0.3876 | Yes | ||
| 102 | ERN1 | 15123 | -1.258 | -0.3836 | Yes | ||
| 103 | MNAT1 | 15294 | -1.324 | -0.3869 | Yes | ||
| 104 | CCNG2 | 15359 | -1.345 | -0.3843 | Yes | ||
| 105 | CDT1 | 15674 | -1.479 | -0.3947 | Yes | ||
| 106 | GMNN | 15735 | -1.506 | -0.3912 | Yes | ||
| 107 | PIN1 | 16127 | -1.683 | -0.4049 | Yes | ||
| 108 | CDKN1C | 16139 | -1.686 | -0.3980 | Yes | ||
| 109 | CCNT2 | 16150 | -1.693 | -0.3909 | Yes | ||
| 110 | ZWINT | 16172 | -1.700 | -0.3845 | Yes | ||
| 111 | FBXO5 | 16196 | -1.714 | -0.3780 | Yes | ||
| 112 | CDC25C | 16298 | -1.762 | -0.3756 | Yes | ||
| 113 | CCNK | 16572 | -1.893 | -0.3819 | Yes | ||
| 114 | CDC45L | 16582 | -1.897 | -0.3739 | Yes | ||
| 115 | UHRF2 | 16772 | -2.003 | -0.3752 | Yes | ||
| 116 | KHDRBS1 | 16794 | -2.018 | -0.3673 | Yes | ||
| 117 | CDKN2A | 16805 | -2.022 | -0.3588 | Yes | ||
| 118 | ZW10 | 16818 | -2.028 | -0.3504 | Yes | ||
| 119 | EIF4G2 | 16823 | -2.030 | -0.3415 | Yes | ||
| 120 | CUL1 | 16836 | -2.035 | -0.3331 | Yes | ||
| 121 | TBRG1 | 16839 | -2.036 | -0.3241 | Yes | ||
| 122 | CCNE2 | 16850 | -2.041 | -0.3155 | Yes | ||
| 123 | NOTCH2 | 16901 | -2.067 | -0.3089 | Yes | ||
| 124 | CHFR | 16958 | -2.095 | -0.3026 | Yes | ||
| 125 | RAD17 | 17065 | -2.152 | -0.2987 | Yes | ||
| 126 | PTPRC | 17131 | -2.196 | -0.2924 | Yes | ||
| 127 | NBN | 17226 | -2.246 | -0.2875 | Yes | ||
| 128 | SESN1 | 17265 | -2.267 | -0.2794 | Yes | ||
| 129 | RB1 | 17323 | -2.301 | -0.2722 | Yes | ||
| 130 | BCCIP | 17378 | -2.335 | -0.2646 | Yes | ||
| 131 | CDK2 | 17488 | -2.401 | -0.2598 | Yes | ||
| 132 | HCFC1 | 17565 | -2.449 | -0.2530 | Yes | ||
| 133 | CCNT1 | 17639 | -2.498 | -0.2457 | Yes | ||
| 134 | GTF2H1 | 17644 | -2.502 | -0.2348 | Yes | ||
| 135 | ATR | 17777 | -2.586 | -0.2304 | Yes | ||
| 136 | BUB1B | 17842 | -2.637 | -0.2220 | Yes | ||
| 137 | MAD2L1 | 17898 | -2.685 | -0.2130 | Yes | ||
| 138 | CUL3 | 17909 | -2.691 | -0.2015 | Yes | ||
| 139 | NUSAP1 | 17980 | -2.749 | -0.1930 | Yes | ||
| 140 | EGF | 18031 | -2.793 | -0.1832 | Yes | ||
| 141 | CHEK2 | 18071 | -2.831 | -0.1726 | Yes | ||
| 142 | CDC7 | 18162 | -2.928 | -0.1644 | Yes | ||
| 143 | GTPBP4 | 18189 | -2.963 | -0.1526 | Yes | ||
| 144 | HUS1 | 18218 | -3.005 | -0.1406 | Yes | ||
| 145 | CCNA2 | 18230 | -3.023 | -0.1277 | Yes | ||
| 146 | CUL4A | 18284 | -3.111 | -0.1167 | Yes | ||
| 147 | SMC1A | 18339 | -3.191 | -0.1053 | Yes | ||
| 148 | APC | 18346 | -3.202 | -0.0913 | Yes | ||
| 149 | LATS2 | 18471 | -3.501 | -0.0824 | Yes | ||
| 150 | CCND1 | 18545 | -3.865 | -0.0690 | Yes | ||
| 151 | ANLN | 18546 | -3.873 | -0.0517 | Yes | ||
| 152 | PCAF | 18550 | -3.924 | -0.0343 | Yes | ||
| 153 | TGFA | 18567 | -4.065 | -0.0170 | Yes | ||
| 154 | MADD | 18595 | -4.378 | 0.0011 | Yes |
