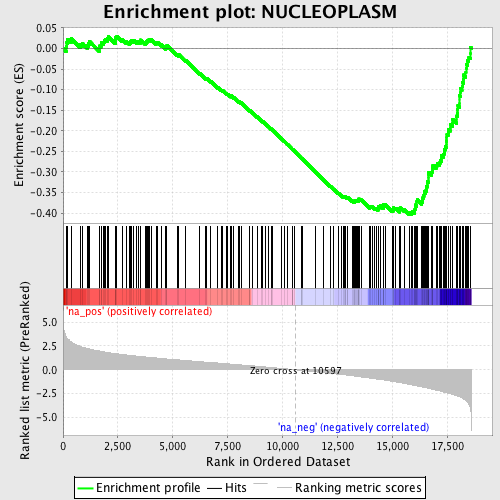
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | NUCLEOPLASM |
| Enrichment Score (ES) | -0.4032021 |
| Normalized Enrichment Score (NES) | -2.0647364 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.008474731 |
| FWER p-Value | 0.093 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SMARCB1 | 139 | 3.434 | 0.0040 | No | ||
| 2 | NUP98 | 169 | 3.340 | 0.0137 | No | ||
| 3 | HTATIP | 206 | 3.244 | 0.0226 | No | ||
| 4 | GEMIN7 | 367 | 2.923 | 0.0238 | No | ||
| 5 | POLR3H | 782 | 2.450 | 0.0096 | No | ||
| 6 | ERCC2 | 889 | 2.361 | 0.0117 | No | ||
| 7 | THRAP3 | 1116 | 2.213 | 0.0069 | No | ||
| 8 | MCRS1 | 1151 | 2.191 | 0.0124 | No | ||
| 9 | SMARCD3 | 1200 | 2.166 | 0.0171 | No | ||
| 10 | PML | 1644 | 1.966 | -0.0003 | No | ||
| 11 | TAF10 | 1650 | 1.962 | 0.0060 | No | ||
| 12 | FMR1 | 1729 | 1.931 | 0.0083 | No | ||
| 13 | HMGA1 | 1742 | 1.926 | 0.0141 | No | ||
| 14 | ISG20 | 1839 | 1.890 | 0.0153 | No | ||
| 15 | ACTB | 1873 | 1.871 | 0.0198 | No | ||
| 16 | GLIS2 | 1941 | 1.846 | 0.0224 | No | ||
| 17 | SMARCD1 | 2029 | 1.813 | 0.0237 | No | ||
| 18 | GCM1 | 2050 | 1.807 | 0.0287 | No | ||
| 19 | RBM14 | 2381 | 1.692 | 0.0165 | No | ||
| 20 | POLR2E | 2385 | 1.691 | 0.0220 | No | ||
| 21 | HES6 | 2393 | 1.689 | 0.0273 | No | ||
| 22 | RUVBL2 | 2450 | 1.673 | 0.0299 | No | ||
| 23 | POLR2B | 2703 | 1.604 | 0.0216 | No | ||
| 24 | ATXN1 | 2880 | 1.552 | 0.0173 | No | ||
| 25 | HDAC6 | 3020 | 1.516 | 0.0149 | No | ||
| 26 | TAF7 | 3075 | 1.503 | 0.0170 | No | ||
| 27 | IFI16 | 3128 | 1.490 | 0.0192 | No | ||
| 28 | NUFIP1 | 3206 | 1.467 | 0.0199 | No | ||
| 29 | PMF1 | 3364 | 1.430 | 0.0162 | No | ||
| 30 | NKX2-5 | 3448 | 1.408 | 0.0165 | No | ||
| 31 | HDAC5 | 3516 | 1.394 | 0.0175 | No | ||
| 32 | TAF6 | 3546 | 1.388 | 0.0206 | No | ||
| 33 | HDAC3 | 3770 | 1.334 | 0.0130 | No | ||
| 34 | EPAS1 | 3791 | 1.330 | 0.0164 | No | ||
| 35 | APTX | 3829 | 1.323 | 0.0188 | No | ||
| 36 | HDAC7A | 3879 | 1.310 | 0.0206 | No | ||
| 37 | LRCH4 | 3925 | 1.302 | 0.0225 | No | ||
| 38 | WDR4 | 4022 | 1.284 | 0.0216 | No | ||
| 39 | FOXE3 | 4238 | 1.238 | 0.0141 | No | ||
| 40 | MPG | 4294 | 1.226 | 0.0153 | No | ||
| 41 | NAP1L3 | 4473 | 1.183 | 0.0096 | No | ||
| 42 | TAF12 | 4687 | 1.139 | 0.0018 | No | ||
| 43 | MCM7 | 4727 | 1.130 | 0.0035 | No | ||
| 44 | BNIP3 | 4730 | 1.129 | 0.0072 | No | ||
| 45 | SPTBN4 | 5201 | 1.042 | -0.0148 | No | ||
| 46 | OGG1 | 5278 | 1.028 | -0.0155 | No | ||
| 47 | TIMM50 | 5588 | 0.966 | -0.0290 | No | ||
| 48 | HIF1A | 6233 | 0.849 | -0.0611 | No | ||
| 49 | ZBTB16 | 6501 | 0.800 | -0.0729 | No | ||
| 50 | FIGLA | 6533 | 0.796 | -0.0719 | No | ||
| 51 | HDAC9 | 6717 | 0.760 | -0.0793 | No | ||
| 52 | GEMIN6 | 7043 | 0.707 | -0.0945 | No | ||
| 53 | FOXF2 | 7204 | 0.679 | -0.1010 | No | ||
| 54 | NUDT21 | 7286 | 0.666 | -0.1031 | No | ||
| 55 | NDUFA13 | 7453 | 0.635 | -0.1100 | No | ||
| 56 | HIPK2 | 7510 | 0.624 | -0.1109 | No | ||
| 57 | LMO4 | 7641 | 0.599 | -0.1160 | No | ||
| 58 | TAF6L | 7652 | 0.596 | -0.1145 | No | ||
| 59 | TDG | 7781 | 0.570 | -0.1195 | No | ||
| 60 | POLR2J | 7990 | 0.533 | -0.1290 | No | ||
| 61 | U2AF1 | 8030 | 0.525 | -0.1294 | No | ||
| 62 | TFIP11 | 8111 | 0.510 | -0.1320 | No | ||
| 63 | PHB | 8480 | 0.439 | -0.1505 | No | ||
| 64 | RUVBL1 | 8510 | 0.434 | -0.1506 | No | ||
| 65 | CIB1 | 8652 | 0.407 | -0.1569 | No | ||
| 66 | POLA1 | 8844 | 0.374 | -0.1660 | No | ||
| 67 | GEMIN5 | 9038 | 0.336 | -0.1754 | No | ||
| 68 | POLR3C | 9103 | 0.322 | -0.1778 | No | ||
| 69 | CHAF1B | 9230 | 0.291 | -0.1836 | No | ||
| 70 | NCR1 | 9352 | 0.268 | -0.1893 | No | ||
| 71 | PPP1R9B | 9478 | 0.238 | -0.1953 | No | ||
| 72 | TP53 | 9507 | 0.232 | -0.1960 | No | ||
| 73 | NPAS2 | 9550 | 0.222 | -0.1976 | No | ||
| 74 | HDAC10 | 9965 | 0.134 | -0.2196 | No | ||
| 75 | PRM2 | 10096 | 0.111 | -0.2263 | No | ||
| 76 | SMARCD2 | 10234 | 0.079 | -0.2334 | No | ||
| 77 | ELP4 | 10443 | 0.033 | -0.2446 | No | ||
| 78 | ELL2 | 10453 | 0.031 | -0.2450 | No | ||
| 79 | CDK9 | 10456 | 0.030 | -0.2450 | No | ||
| 80 | POLR2A | 10546 | 0.010 | -0.2498 | No | ||
| 81 | CHAF1A | 10863 | -0.064 | -0.2668 | No | ||
| 82 | XPO1 | 10884 | -0.068 | -0.2676 | No | ||
| 83 | SP100 | 10912 | -0.075 | -0.2688 | No | ||
| 84 | TAF4 | 11511 | -0.211 | -0.3006 | No | ||
| 85 | RELA | 11860 | -0.290 | -0.3185 | No | ||
| 86 | HDAC8 | 12185 | -0.367 | -0.3349 | No | ||
| 87 | PRPF31 | 12341 | -0.405 | -0.3419 | No | ||
| 88 | CBX1 | 12541 | -0.458 | -0.3512 | No | ||
| 89 | PHF21A | 12693 | -0.494 | -0.3577 | No | ||
| 90 | GTF2F1 | 12759 | -0.510 | -0.3595 | No | ||
| 91 | PTBP1 | 12809 | -0.523 | -0.3604 | No | ||
| 92 | NAP1L2 | 12852 | -0.534 | -0.3609 | No | ||
| 93 | CPSF3 | 12862 | -0.536 | -0.3596 | No | ||
| 94 | HDAC11 | 12960 | -0.562 | -0.3630 | No | ||
| 95 | PAX8 | 12967 | -0.567 | -0.3614 | No | ||
| 96 | CLOCK | 13167 | -0.620 | -0.3701 | No | ||
| 97 | DYRK1A | 13236 | -0.637 | -0.3717 | No | ||
| 98 | ELP3 | 13274 | -0.651 | -0.3715 | No | ||
| 99 | TAF9 | 13299 | -0.660 | -0.3706 | No | ||
| 100 | GTF3C3 | 13323 | -0.667 | -0.3696 | No | ||
| 101 | GTF2H4 | 13372 | -0.683 | -0.3699 | No | ||
| 102 | PTF1A | 13427 | -0.697 | -0.3705 | No | ||
| 103 | GTF3C5 | 13465 | -0.708 | -0.3701 | No | ||
| 104 | ZNF148 | 13471 | -0.708 | -0.3680 | No | ||
| 105 | POLI | 13472 | -0.709 | -0.3656 | No | ||
| 106 | NR6A1 | 13518 | -0.723 | -0.3656 | No | ||
| 107 | COIL | 13604 | -0.753 | -0.3677 | No | ||
| 108 | NFYC | 13982 | -0.858 | -0.3852 | No | ||
| 109 | MEN1 | 14004 | -0.866 | -0.3835 | No | ||
| 110 | PPARGC1A | 14078 | -0.893 | -0.3844 | No | ||
| 111 | SNAPC4 | 14196 | -0.931 | -0.3877 | No | ||
| 112 | ARID4A | 14265 | -0.953 | -0.3881 | No | ||
| 113 | PITX2 | 14353 | -0.981 | -0.3896 | No | ||
| 114 | EDF1 | 14376 | -0.989 | -0.3874 | No | ||
| 115 | MED6 | 14380 | -0.990 | -0.3843 | No | ||
| 116 | HCLS1 | 14443 | -1.009 | -0.3842 | No | ||
| 117 | PPARGC1B | 14457 | -1.013 | -0.3815 | No | ||
| 118 | ELF4 | 14590 | -1.055 | -0.3851 | No | ||
| 119 | SHFM1 | 14593 | -1.056 | -0.3817 | No | ||
| 120 | UBE2I | 14608 | -1.062 | -0.3789 | No | ||
| 121 | ERCC3 | 14676 | -1.088 | -0.3789 | No | ||
| 122 | MYO6 | 15002 | -1.208 | -0.3924 | No | ||
| 123 | TOPBP1 | 15048 | -1.224 | -0.3908 | No | ||
| 124 | MED4 | 15055 | -1.228 | -0.3869 | No | ||
| 125 | PHF12 | 15154 | -1.268 | -0.3880 | No | ||
| 126 | FUSIP1 | 15328 | -1.334 | -0.3929 | No | ||
| 127 | SAP30 | 15343 | -1.339 | -0.3892 | No | ||
| 128 | YEATS4 | 15389 | -1.360 | -0.3870 | No | ||
| 129 | ARNTL | 15540 | -1.419 | -0.3904 | No | ||
| 130 | ERCC5 | 15767 | -1.518 | -0.3976 | No | ||
| 131 | HDAC1 | 15872 | -1.567 | -0.3979 | Yes | ||
| 132 | TAF11 | 15919 | -1.587 | -0.3951 | Yes | ||
| 133 | MKI67IP | 16014 | -1.634 | -0.3947 | Yes | ||
| 134 | IKBKAP | 16033 | -1.643 | -0.3901 | Yes | ||
| 135 | MRE11A | 16048 | -1.648 | -0.3854 | Yes | ||
| 136 | TRIM27 | 16052 | -1.651 | -0.3800 | Yes | ||
| 137 | WDHD1 | 16121 | -1.681 | -0.3780 | Yes | ||
| 138 | ASH2L | 16123 | -1.681 | -0.3724 | Yes | ||
| 139 | IVNS1ABP | 16132 | -1.685 | -0.3672 | Yes | ||
| 140 | CPSF1 | 16348 | -1.786 | -0.3728 | Yes | ||
| 141 | RTCD1 | 16377 | -1.800 | -0.3683 | Yes | ||
| 142 | ASF1A | 16393 | -1.808 | -0.3630 | Yes | ||
| 143 | NFYA | 16411 | -1.818 | -0.3578 | Yes | ||
| 144 | EZH2 | 16450 | -1.831 | -0.3537 | Yes | ||
| 145 | ACTL6A | 16453 | -1.832 | -0.3477 | Yes | ||
| 146 | NCOA6 | 16535 | -1.874 | -0.3457 | Yes | ||
| 147 | TBP | 16552 | -1.880 | -0.3403 | Yes | ||
| 148 | GTF3C1 | 16566 | -1.891 | -0.3346 | Yes | ||
| 149 | SMARCC1 | 16602 | -1.911 | -0.3301 | Yes | ||
| 150 | NAP1L4 | 16604 | -1.914 | -0.3237 | Yes | ||
| 151 | MED12 | 16648 | -1.932 | -0.3195 | Yes | ||
| 152 | TOP2A | 16650 | -1.933 | -0.3131 | Yes | ||
| 153 | SMARCA4 | 16653 | -1.934 | -0.3067 | Yes | ||
| 154 | HNRPK | 16662 | -1.937 | -0.3006 | Yes | ||
| 155 | CDKN2A | 16805 | -2.022 | -0.3015 | Yes | ||
| 156 | SMARCE1 | 16820 | -2.028 | -0.2954 | Yes | ||
| 157 | NAP1L1 | 16827 | -2.032 | -0.2889 | Yes | ||
| 158 | POLQ | 16854 | -2.044 | -0.2835 | Yes | ||
| 159 | POLR3K | 17001 | -2.120 | -0.2843 | Yes | ||
| 160 | GTF3C2 | 17041 | -2.142 | -0.2792 | Yes | ||
| 161 | ARID1A | 17156 | -2.208 | -0.2779 | Yes | ||
| 162 | RPA1 | 17181 | -2.221 | -0.2718 | Yes | ||
| 163 | RBBP5 | 17242 | -2.254 | -0.2674 | Yes | ||
| 164 | ATXN3 | 17252 | -2.261 | -0.2603 | Yes | ||
| 165 | RB1 | 17323 | -2.301 | -0.2564 | Yes | ||
| 166 | INTS6 | 17365 | -2.327 | -0.2508 | Yes | ||
| 167 | WRN | 17392 | -2.343 | -0.2443 | Yes | ||
| 168 | SAP18 | 17437 | -2.371 | -0.2387 | Yes | ||
| 169 | RBM39 | 17472 | -2.390 | -0.2325 | Yes | ||
| 170 | GIT2 | 17475 | -2.391 | -0.2246 | Yes | ||
| 171 | ING2 | 17490 | -2.402 | -0.2172 | Yes | ||
| 172 | TOPORS | 17494 | -2.407 | -0.2093 | Yes | ||
| 173 | SFRS2 | 17548 | -2.439 | -0.2040 | Yes | ||
| 174 | OIP5 | 17566 | -2.449 | -0.1966 | Yes | ||
| 175 | XRCC6 | 17640 | -2.499 | -0.1922 | Yes | ||
| 176 | GTF2H1 | 17644 | -2.502 | -0.1839 | Yes | ||
| 177 | NUP54 | 17751 | -2.571 | -0.1810 | Yes | ||
| 178 | KPNA2 | 17761 | -2.577 | -0.1729 | Yes | ||
| 179 | HDAC2 | 17934 | -2.708 | -0.1731 | Yes | ||
| 180 | SFRS2IP | 17936 | -2.710 | -0.1640 | Yes | ||
| 181 | SMAD2 | 17972 | -2.742 | -0.1567 | Yes | ||
| 182 | NFYB | 17985 | -2.753 | -0.1481 | Yes | ||
| 183 | CDK8 | 17994 | -2.758 | -0.1392 | Yes | ||
| 184 | CHEK2 | 18071 | -2.831 | -0.1338 | Yes | ||
| 185 | RSF1 | 18074 | -2.833 | -0.1244 | Yes | ||
| 186 | ZNF638 | 18075 | -2.833 | -0.1149 | Yes | ||
| 187 | TOP1 | 18114 | -2.872 | -0.1073 | Yes | ||
| 188 | SMARCA5 | 18120 | -2.882 | -0.0978 | Yes | ||
| 189 | TAF13 | 18182 | -2.948 | -0.0912 | Yes | ||
| 190 | TOP2B | 18210 | -2.984 | -0.0826 | Yes | ||
| 191 | NARG1 | 18243 | -3.040 | -0.0742 | Yes | ||
| 192 | SMAD4 | 18254 | -3.068 | -0.0644 | Yes | ||
| 193 | MDM2 | 18357 | -3.228 | -0.0590 | Yes | ||
| 194 | GTF3C4 | 18363 | -3.239 | -0.0484 | Yes | ||
| 195 | MPHOSPH1 | 18370 | -3.248 | -0.0378 | Yes | ||
| 196 | EED | 18439 | -3.421 | -0.0300 | Yes | ||
| 197 | POLH | 18493 | -3.557 | -0.0209 | Yes | ||
| 198 | TAF5 | 18566 | -4.062 | -0.0111 | Yes | ||
| 199 | EPC1 | 18571 | -4.100 | 0.0024 | Yes |
