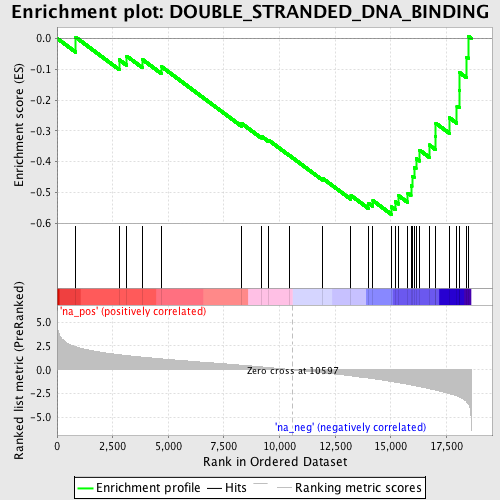
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | DOUBLE_STRANDED_DNA_BINDING |
| Enrichment Score (ES) | -0.5711651 |
| Normalized Enrichment Score (NES) | -2.0430977 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.008291105 |
| FWER p-Value | 0.13 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PNKP | 839 | 2.401 | 0.0035 | No | ||
| 2 | NTHL1 | 2792 | 1.579 | -0.0696 | No | ||
| 3 | IFI16 | 3128 | 1.490 | -0.0574 | No | ||
| 4 | APTX | 3829 | 1.323 | -0.0683 | No | ||
| 5 | HLF | 4698 | 1.136 | -0.0920 | No | ||
| 6 | XRCC5 | 8298 | 0.473 | -0.2761 | No | ||
| 7 | MSH2 | 9177 | 0.304 | -0.3172 | No | ||
| 8 | TP53 | 9507 | 0.232 | -0.3302 | No | ||
| 9 | TERF1 | 10434 | 0.034 | -0.3793 | No | ||
| 10 | CSDA | 11929 | -0.306 | -0.4535 | No | ||
| 11 | FEN1 | 13207 | -0.630 | -0.5095 | No | ||
| 12 | PURA | 14002 | -0.866 | -0.5347 | No | ||
| 13 | RBMS1 | 14194 | -0.931 | -0.5261 | No | ||
| 14 | PMS2 | 15033 | -1.218 | -0.5465 | Yes | ||
| 15 | YBX1 | 15207 | -1.289 | -0.5297 | Yes | ||
| 16 | CGGBP1 | 15351 | -1.342 | -0.5103 | Yes | ||
| 17 | ERCC5 | 15767 | -1.518 | -0.5019 | Yes | ||
| 18 | HMGB2 | 15908 | -1.582 | -0.4773 | Yes | ||
| 19 | MSH3 | 15952 | -1.600 | -0.4473 | Yes | ||
| 20 | MRE11A | 16048 | -1.648 | -0.4190 | Yes | ||
| 21 | MSH6 | 16134 | -1.685 | -0.3895 | Yes | ||
| 22 | MTERF | 16292 | -1.757 | -0.3624 | Yes | ||
| 23 | RAD51 | 16726 | -1.978 | -0.3456 | Yes | ||
| 24 | MLH1 | 17012 | -2.126 | -0.3179 | Yes | ||
| 25 | HNRPDL | 17024 | -2.135 | -0.2753 | Yes | ||
| 26 | XRCC6 | 17640 | -2.499 | -0.2578 | Yes | ||
| 27 | SMAD2 | 17972 | -2.742 | -0.2200 | Yes | ||
| 28 | ZNF638 | 18075 | -2.833 | -0.1682 | Yes | ||
| 29 | RAD51AP1 | 18097 | -2.857 | -0.1115 | Yes | ||
| 30 | SATB1 | 18414 | -3.343 | -0.0608 | Yes | ||
| 31 | KIN | 18482 | -3.536 | 0.0072 | Yes |
