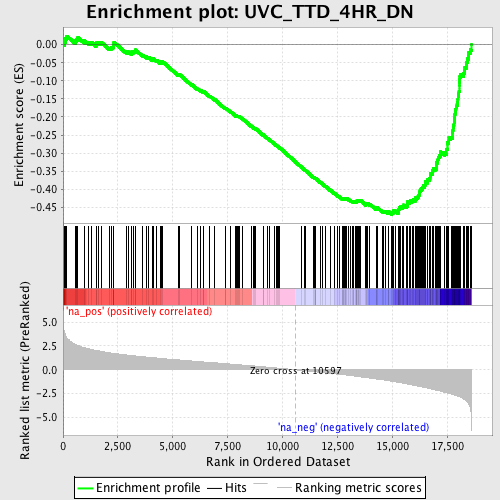
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | UVC_TTD_4HR_DN |
| Enrichment Score (ES) | -0.46982092 |
| Normalized Enrichment Score (NES) | -2.4439871 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GNAQ | 64 | 3.955 | 0.0085 | No | ||
| 2 | DLC1 | 109 | 3.570 | 0.0169 | No | ||
| 3 | NUP98 | 169 | 3.340 | 0.0238 | No | ||
| 4 | MAP4K5 | 556 | 2.658 | 0.0109 | No | ||
| 5 | LHFPL2 | 613 | 2.599 | 0.0157 | No | ||
| 6 | HAS2 | 673 | 2.542 | 0.0202 | No | ||
| 7 | IGF2BP3 | 965 | 2.304 | 0.0113 | No | ||
| 8 | EGFR | 1159 | 2.187 | 0.0075 | No | ||
| 9 | ZHX3 | 1305 | 2.107 | 0.0060 | No | ||
| 10 | PEX14 | 1503 | 2.026 | 0.0014 | No | ||
| 11 | FYN | 1528 | 2.012 | 0.0062 | No | ||
| 12 | PPP2R2A | 1627 | 1.973 | 0.0068 | No | ||
| 13 | RNF13 | 1757 | 1.920 | 0.0056 | No | ||
| 14 | EXT1 | 2128 | 1.776 | -0.0091 | No | ||
| 15 | BICD1 | 2188 | 1.754 | -0.0070 | No | ||
| 16 | CREB5 | 2274 | 1.724 | -0.0064 | No | ||
| 17 | CBLB | 2285 | 1.721 | -0.0018 | No | ||
| 18 | FGF2 | 2313 | 1.711 | 0.0019 | No | ||
| 19 | SMAD3 | 2315 | 1.711 | 0.0071 | No | ||
| 20 | ATXN1 | 2880 | 1.552 | -0.0189 | No | ||
| 21 | MTMR3 | 2984 | 1.526 | -0.0199 | No | ||
| 22 | CENPE | 3121 | 1.491 | -0.0227 | No | ||
| 23 | TLE1 | 3135 | 1.487 | -0.0190 | No | ||
| 24 | SRGAP2 | 3213 | 1.466 | -0.0187 | No | ||
| 25 | ANKS1A | 3286 | 1.445 | -0.0182 | No | ||
| 26 | GULP1 | 3296 | 1.445 | -0.0144 | No | ||
| 27 | WDFY3 | 3640 | 1.369 | -0.0289 | No | ||
| 28 | HOMER1 | 3796 | 1.330 | -0.0333 | No | ||
| 29 | MYST3 | 3903 | 1.304 | -0.0351 | No | ||
| 30 | SYNJ2 | 4068 | 1.275 | -0.0401 | No | ||
| 31 | MGAT5 | 4108 | 1.266 | -0.0384 | No | ||
| 32 | DAAM1 | 4252 | 1.235 | -0.0424 | No | ||
| 33 | ARHGEF7 | 4422 | 1.194 | -0.0480 | No | ||
| 34 | TRAM2 | 4467 | 1.185 | -0.0468 | No | ||
| 35 | INPP5F | 4542 | 1.166 | -0.0473 | No | ||
| 36 | BHLHB2 | 5252 | 1.032 | -0.0827 | No | ||
| 37 | SLIT1 | 5292 | 1.025 | -0.0817 | No | ||
| 38 | ARID5B | 5848 | 0.920 | -0.1091 | No | ||
| 39 | RGS7 | 6140 | 0.866 | -0.1223 | No | ||
| 40 | CD2AP | 6276 | 0.840 | -0.1271 | No | ||
| 41 | TGFB2 | 6387 | 0.822 | -0.1306 | No | ||
| 42 | ZCCHC11 | 6410 | 0.817 | -0.1293 | No | ||
| 43 | TSPAN5 | 6687 | 0.766 | -0.1420 | No | ||
| 44 | HIRA | 6908 | 0.727 | -0.1518 | No | ||
| 45 | ACVR1 | 7384 | 0.648 | -0.1756 | No | ||
| 46 | EHBP1 | 7417 | 0.641 | -0.1754 | No | ||
| 47 | CTGF | 7634 | 0.599 | -0.1853 | No | ||
| 48 | ARHGAP12 | 7867 | 0.555 | -0.1963 | No | ||
| 49 | SLC25A24 | 7903 | 0.548 | -0.1965 | No | ||
| 50 | TCF7L2 | 7952 | 0.538 | -0.1975 | No | ||
| 51 | ACVR2A | 7995 | 0.531 | -0.1982 | No | ||
| 52 | SEMA3C | 8031 | 0.525 | -0.1985 | No | ||
| 53 | LRIG1 | 8189 | 0.494 | -0.2055 | No | ||
| 54 | CENTG2 | 8569 | 0.422 | -0.2248 | No | ||
| 55 | FGF5 | 8688 | 0.401 | -0.2300 | No | ||
| 56 | PTPN2 | 8723 | 0.395 | -0.2307 | No | ||
| 57 | SMURF2 | 8782 | 0.384 | -0.2327 | No | ||
| 58 | RRAS2 | 9117 | 0.319 | -0.2499 | No | ||
| 59 | TUBGCP3 | 9120 | 0.318 | -0.2490 | No | ||
| 60 | PLEKHC1 | 9336 | 0.273 | -0.2599 | No | ||
| 61 | ITSN1 | 9422 | 0.250 | -0.2637 | No | ||
| 62 | EDG2 | 9623 | 0.206 | -0.2740 | No | ||
| 63 | SCHIP1 | 9714 | 0.185 | -0.2783 | No | ||
| 64 | STK39 | 9789 | 0.168 | -0.2818 | No | ||
| 65 | PRKCA | 9829 | 0.161 | -0.2834 | No | ||
| 66 | NRIP1 | 9881 | 0.150 | -0.2858 | No | ||
| 67 | FBXW11 | 10879 | -0.066 | -0.3397 | No | ||
| 68 | ROR1 | 10987 | -0.090 | -0.3453 | No | ||
| 69 | SLC16A7 | 11027 | -0.100 | -0.3471 | No | ||
| 70 | CLASP2 | 11399 | -0.184 | -0.3667 | No | ||
| 71 | FBXL7 | 11441 | -0.197 | -0.3683 | No | ||
| 72 | VEGFC | 11480 | -0.206 | -0.3698 | No | ||
| 73 | ARNT2 | 11498 | -0.209 | -0.3701 | No | ||
| 74 | DDX10 | 11510 | -0.211 | -0.3700 | No | ||
| 75 | THRAP1 | 11739 | -0.262 | -0.3816 | No | ||
| 76 | EPHA4 | 11818 | -0.279 | -0.3850 | No | ||
| 77 | CRY1 | 11974 | -0.318 | -0.3925 | No | ||
| 78 | TRIM24 | 12170 | -0.365 | -0.4020 | No | ||
| 79 | TLE4 | 12360 | -0.409 | -0.4110 | No | ||
| 80 | NCOA3 | 12515 | -0.452 | -0.4180 | No | ||
| 81 | EVI5 | 12603 | -0.474 | -0.4213 | No | ||
| 82 | GOLGA4 | 12713 | -0.499 | -0.4257 | No | ||
| 83 | GNAI1 | 12720 | -0.500 | -0.4245 | No | ||
| 84 | IFRD1 | 12779 | -0.515 | -0.4261 | No | ||
| 85 | THRAP2 | 12787 | -0.516 | -0.4249 | No | ||
| 86 | PRIM2A | 12842 | -0.531 | -0.4263 | No | ||
| 87 | ZFYVE9 | 12847 | -0.532 | -0.4249 | No | ||
| 88 | ZHX2 | 12884 | -0.541 | -0.4252 | No | ||
| 89 | E2F3 | 12935 | -0.554 | -0.4262 | No | ||
| 90 | F3 | 13007 | -0.576 | -0.4284 | No | ||
| 91 | AUH | 13090 | -0.600 | -0.4310 | No | ||
| 92 | SIPA1L1 | 13166 | -0.620 | -0.4332 | No | ||
| 93 | DYRK1A | 13236 | -0.637 | -0.4350 | No | ||
| 94 | MTF2 | 13249 | -0.642 | -0.4337 | No | ||
| 95 | ORC2L | 13315 | -0.665 | -0.4352 | No | ||
| 96 | NEK7 | 13361 | -0.680 | -0.4356 | No | ||
| 97 | IL7R | 13373 | -0.683 | -0.4342 | No | ||
| 98 | MITF | 13381 | -0.686 | -0.4325 | No | ||
| 99 | RGL1 | 13418 | -0.695 | -0.4323 | No | ||
| 100 | CLASP1 | 13461 | -0.707 | -0.4325 | No | ||
| 101 | ZNF148 | 13471 | -0.708 | -0.4308 | No | ||
| 102 | KLF10 | 13524 | -0.725 | -0.4314 | No | ||
| 103 | PHLPP | 13546 | -0.732 | -0.4304 | No | ||
| 104 | STARD13 | 13801 | -0.811 | -0.4417 | No | ||
| 105 | PCTK2 | 13824 | -0.817 | -0.4404 | No | ||
| 106 | DNMBP | 13844 | -0.821 | -0.4390 | No | ||
| 107 | CENPA | 13875 | -0.829 | -0.4381 | No | ||
| 108 | BCAR3 | 13985 | -0.860 | -0.4414 | No | ||
| 109 | PIP5K3 | 14280 | -0.958 | -0.4545 | No | ||
| 110 | NUP153 | 14299 | -0.964 | -0.4526 | No | ||
| 111 | ANAPC10 | 14310 | -0.967 | -0.4502 | No | ||
| 112 | IRAK1BP1 | 14555 | -1.041 | -0.4603 | No | ||
| 113 | LYN | 14620 | -1.067 | -0.4605 | No | ||
| 114 | TBL1X | 14710 | -1.102 | -0.4620 | No | ||
| 115 | CASP8 | 14812 | -1.139 | -0.4641 | No | ||
| 116 | SOS1 | 14822 | -1.141 | -0.4611 | No | ||
| 117 | PTPRK | 14983 | -1.203 | -0.4662 | Yes | ||
| 118 | PPAP2B | 15021 | -1.214 | -0.4645 | Yes | ||
| 119 | TOPBP1 | 15048 | -1.224 | -0.4622 | Yes | ||
| 120 | PARD3 | 15072 | -1.235 | -0.4597 | Yes | ||
| 121 | BMP2K | 15147 | -1.265 | -0.4599 | Yes | ||
| 122 | LARS2 | 15270 | -1.315 | -0.4626 | Yes | ||
| 123 | SOCS5 | 15286 | -1.320 | -0.4594 | Yes | ||
| 124 | MNAT1 | 15294 | -1.324 | -0.4558 | Yes | ||
| 125 | SWAP70 | 15316 | -1.331 | -0.4529 | Yes | ||
| 126 | CREBBP | 15329 | -1.334 | -0.4495 | Yes | ||
| 127 | GSK3B | 15391 | -1.361 | -0.4487 | Yes | ||
| 128 | MDN1 | 15474 | -1.395 | -0.4489 | Yes | ||
| 129 | PBX3 | 15509 | -1.408 | -0.4465 | Yes | ||
| 130 | KIF2A | 15533 | -1.417 | -0.4435 | Yes | ||
| 131 | JMJD2C | 15658 | -1.472 | -0.4458 | Yes | ||
| 132 | UGCG | 15685 | -1.484 | -0.4427 | Yes | ||
| 133 | UST | 15697 | -1.489 | -0.4388 | Yes | ||
| 134 | KLF7 | 15710 | -1.494 | -0.4349 | Yes | ||
| 135 | ARFGEF1 | 15799 | -1.532 | -0.4351 | Yes | ||
| 136 | SOCS6 | 15822 | -1.541 | -0.4316 | Yes | ||
| 137 | PTPN12 | 15902 | -1.580 | -0.4311 | Yes | ||
| 138 | MSH3 | 15952 | -1.600 | -0.4289 | Yes | ||
| 139 | SAMD4A | 16042 | -1.646 | -0.4288 | Yes | ||
| 140 | RBM16 | 16043 | -1.646 | -0.4238 | Yes | ||
| 141 | WDHD1 | 16121 | -1.681 | -0.4229 | Yes | ||
| 142 | MID1 | 16173 | -1.701 | -0.4205 | Yes | ||
| 143 | CTBP2 | 16208 | -1.718 | -0.4172 | Yes | ||
| 144 | XRCC4 | 16226 | -1.726 | -0.4129 | Yes | ||
| 145 | MELK | 16238 | -1.730 | -0.4083 | Yes | ||
| 146 | FNDC3A | 16264 | -1.741 | -0.4043 | Yes | ||
| 147 | ATF2 | 16286 | -1.753 | -0.4002 | Yes | ||
| 148 | ORC5L | 16341 | -1.784 | -0.3977 | Yes | ||
| 149 | RUNX1 | 16383 | -1.803 | -0.3945 | Yes | ||
| 150 | STK3 | 16407 | -1.816 | -0.3902 | Yes | ||
| 151 | LRP6 | 16492 | -1.852 | -0.3892 | Yes | ||
| 152 | SLC4A7 | 16499 | -1.856 | -0.3839 | Yes | ||
| 153 | MPP6 | 16505 | -1.859 | -0.3786 | Yes | ||
| 154 | BARD1 | 16590 | -1.904 | -0.3774 | Yes | ||
| 155 | MYST4 | 16623 | -1.922 | -0.3733 | Yes | ||
| 156 | PPP2R5E | 16688 | -1.953 | -0.3709 | Yes | ||
| 157 | TCF12 | 16722 | -1.970 | -0.3667 | Yes | ||
| 158 | MYO10 | 16725 | -1.977 | -0.3608 | Yes | ||
| 159 | ARIH1 | 16744 | -1.986 | -0.3558 | Yes | ||
| 160 | UBL3 | 16825 | -2.031 | -0.3540 | Yes | ||
| 161 | NCOA1 | 16840 | -2.036 | -0.3486 | Yes | ||
| 162 | MAPK6 | 16859 | -2.048 | -0.3434 | Yes | ||
| 163 | CUTL1 | 16954 | -2.094 | -0.3421 | Yes | ||
| 164 | SPOP | 17008 | -2.123 | -0.3386 | Yes | ||
| 165 | PPP1R12A | 17014 | -2.128 | -0.3324 | Yes | ||
| 166 | HNRPDL | 17024 | -2.135 | -0.3265 | Yes | ||
| 167 | MAP2K4 | 17047 | -2.144 | -0.3212 | Yes | ||
| 168 | PAFAH1B1 | 17084 | -2.165 | -0.3166 | Yes | ||
| 169 | PUM2 | 17116 | -2.184 | -0.3117 | Yes | ||
| 170 | DAPK1 | 17149 | -2.205 | -0.3067 | Yes | ||
| 171 | PTK2 | 17189 | -2.224 | -0.3021 | Yes | ||
| 172 | CTAGE5 | 17212 | -2.241 | -0.2965 | Yes | ||
| 173 | ZNF292 | 17383 | -2.339 | -0.2987 | Yes | ||
| 174 | RBM39 | 17472 | -2.390 | -0.2962 | Yes | ||
| 175 | CNOT4 | 17473 | -2.390 | -0.2890 | Yes | ||
| 176 | KIF23 | 17511 | -2.417 | -0.2837 | Yes | ||
| 177 | ZZZ3 | 17512 | -2.418 | -0.2764 | Yes | ||
| 178 | SEC24B | 17516 | -2.420 | -0.2692 | Yes | ||
| 179 | TRIM33 | 17582 | -2.459 | -0.2653 | Yes | ||
| 180 | ATBF1 | 17585 | -2.461 | -0.2580 | Yes | ||
| 181 | RBM6 | 17679 | -2.526 | -0.2554 | Yes | ||
| 182 | TLK1 | 17754 | -2.574 | -0.2516 | Yes | ||
| 183 | RNGTT | 17757 | -2.575 | -0.2439 | Yes | ||
| 184 | STRN3 | 17765 | -2.578 | -0.2365 | Yes | ||
| 185 | WDR7 | 17774 | -2.584 | -0.2291 | Yes | ||
| 186 | ABI1 | 17790 | -2.597 | -0.2221 | Yes | ||
| 187 | TLK2 | 17816 | -2.616 | -0.2155 | Yes | ||
| 188 | BLM | 17825 | -2.619 | -0.2081 | Yes | ||
| 189 | PKN2 | 17845 | -2.641 | -0.2011 | Yes | ||
| 190 | EIF4G3 | 17856 | -2.652 | -0.1936 | Yes | ||
| 191 | FCHSD2 | 17866 | -2.659 | -0.1861 | Yes | ||
| 192 | MARCH7 | 17877 | -2.667 | -0.1785 | Yes | ||
| 193 | CNOT2 | 17907 | -2.690 | -0.1720 | Yes | ||
| 194 | OGT | 17951 | -2.718 | -0.1661 | Yes | ||
| 195 | FUBP1 | 17981 | -2.750 | -0.1593 | Yes | ||
| 196 | CDK8 | 17994 | -2.758 | -0.1516 | Yes | ||
| 197 | STAG1 | 17999 | -2.762 | -0.1435 | Yes | ||
| 198 | PARG | 18020 | -2.785 | -0.1362 | Yes | ||
| 199 | ITSN2 | 18042 | -2.802 | -0.1288 | Yes | ||
| 200 | ATXN2 | 18057 | -2.820 | -0.1211 | Yes | ||
| 201 | DHX15 | 18058 | -2.822 | -0.1125 | Yes | ||
| 202 | AQR | 18063 | -2.826 | -0.1042 | Yes | ||
| 203 | ZNF638 | 18075 | -2.833 | -0.0962 | Yes | ||
| 204 | EIF4E | 18085 | -2.842 | -0.0881 | Yes | ||
| 205 | ZRF1 | 18124 | -2.885 | -0.0814 | Yes | ||
| 206 | UPF2 | 18242 | -3.039 | -0.0786 | Yes | ||
| 207 | SRPK2 | 18286 | -3.115 | -0.0715 | Yes | ||
| 208 | REV3L | 18312 | -3.147 | -0.0633 | Yes | ||
| 209 | MPHOSPH1 | 18370 | -3.248 | -0.0566 | Yes | ||
| 210 | ROCK2 | 18399 | -3.300 | -0.0482 | Yes | ||
| 211 | ANKRD17 | 18423 | -3.382 | -0.0392 | Yes | ||
| 212 | CHD1 | 18463 | -3.482 | -0.0308 | Yes | ||
| 213 | SSBP2 | 18492 | -3.556 | -0.0215 | Yes | ||
| 214 | CDYL | 18552 | -3.934 | -0.0128 | Yes | ||
| 215 | IL1RAP | 18610 | -5.370 | 0.0003 | Yes |
