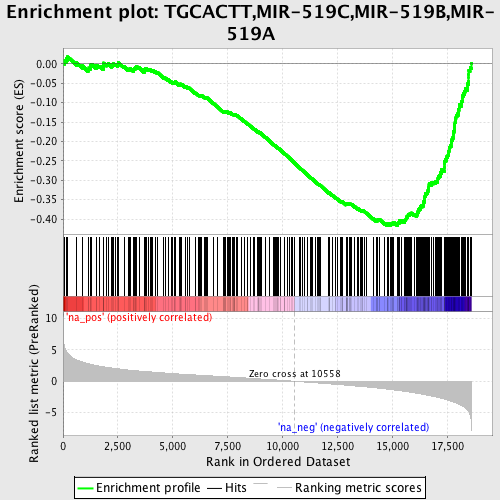
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_absentNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | TGCACTT,MIR-519C,MIR-519B,MIR-519A |
| Enrichment Score (ES) | -0.41765675 |
| Normalized Enrichment Score (NES) | -2.1545396 |
| Nominal p-value | 0.0 |
| FDR q-value | 1.6333714E-4 |
| FWER p-Value | 0.0020 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PIGS | 48 | 5.699 | 0.0087 | No | ||
| 2 | TIMP2 | 137 | 4.817 | 0.0134 | No | ||
| 3 | LIMK1 | 213 | 4.419 | 0.0180 | No | ||
| 4 | SASH1 | 614 | 3.408 | 0.0030 | No | ||
| 5 | PIK4CB | 872 | 3.073 | -0.0050 | No | ||
| 6 | NEUROG1 | 1138 | 2.802 | -0.0139 | No | ||
| 7 | SH3BP5 | 1178 | 2.763 | -0.0106 | No | ||
| 8 | PTHLH | 1236 | 2.723 | -0.0083 | No | ||
| 9 | PPP1R10 | 1237 | 2.722 | -0.0029 | No | ||
| 10 | SUV39H1 | 1291 | 2.683 | -0.0005 | No | ||
| 11 | TGFBR2 | 1508 | 2.516 | -0.0073 | No | ||
| 12 | RNF31 | 1528 | 2.503 | -0.0034 | No | ||
| 13 | TESK2 | 1656 | 2.422 | -0.0055 | No | ||
| 14 | ARHGAP1 | 1829 | 2.317 | -0.0103 | No | ||
| 15 | HOXB3 | 1837 | 2.314 | -0.0061 | No | ||
| 16 | HAS2 | 1844 | 2.306 | -0.0019 | No | ||
| 17 | CREB5 | 1852 | 2.299 | 0.0023 | No | ||
| 18 | NAGK | 1997 | 2.222 | -0.0012 | No | ||
| 19 | LRRIQ2 | 2046 | 2.197 | 0.0005 | No | ||
| 20 | KCNQ2 | 2213 | 2.119 | -0.0043 | No | ||
| 21 | PCDHA1 | 2247 | 2.100 | -0.0020 | No | ||
| 22 | HIF1A | 2285 | 2.081 | 0.0001 | No | ||
| 23 | RAB5B | 2373 | 2.042 | -0.0006 | No | ||
| 24 | LAPTM4A | 2499 | 1.988 | -0.0035 | No | ||
| 25 | CSNK1G1 | 2505 | 1.987 | 0.0002 | No | ||
| 26 | ZFHX4 | 2523 | 1.979 | 0.0032 | No | ||
| 27 | CIC | 2775 | 1.875 | -0.0068 | No | ||
| 28 | KLF12 | 2985 | 1.785 | -0.0147 | No | ||
| 29 | ABHD2 | 3021 | 1.773 | -0.0131 | No | ||
| 30 | TRIOBP | 3065 | 1.758 | -0.0120 | No | ||
| 31 | UBE3A | 3207 | 1.719 | -0.0163 | No | ||
| 32 | FASTK | 3228 | 1.712 | -0.0140 | No | ||
| 33 | GRM7 | 3246 | 1.707 | -0.0115 | No | ||
| 34 | ENPP5 | 3282 | 1.696 | -0.0101 | No | ||
| 35 | BRWD1 | 3316 | 1.684 | -0.0085 | No | ||
| 36 | DCBLD2 | 3350 | 1.673 | -0.0070 | No | ||
| 37 | DPYSL2 | 3459 | 1.641 | -0.0097 | No | ||
| 38 | PTGFRN | 3694 | 1.571 | -0.0194 | No | ||
| 39 | SOX5 | 3699 | 1.569 | -0.0165 | No | ||
| 40 | DUSP8 | 3710 | 1.566 | -0.0139 | No | ||
| 41 | BMPR2 | 3735 | 1.557 | -0.0121 | No | ||
| 42 | HIVEP2 | 3783 | 1.541 | -0.0117 | No | ||
| 43 | WDR1 | 3903 | 1.510 | -0.0152 | No | ||
| 44 | COL4A3 | 3963 | 1.495 | -0.0154 | No | ||
| 45 | ZDHHC1 | 4026 | 1.480 | -0.0159 | No | ||
| 46 | MAP3K3 | 4089 | 1.464 | -0.0164 | No | ||
| 47 | NRP2 | 4193 | 1.437 | -0.0191 | No | ||
| 48 | HMGA2 | 4313 | 1.407 | -0.0229 | No | ||
| 49 | NFIA | 4585 | 1.334 | -0.0350 | No | ||
| 50 | PCDHA5 | 4659 | 1.318 | -0.0364 | No | ||
| 51 | COL19A1 | 4782 | 1.292 | -0.0405 | No | ||
| 52 | KCND2 | 4938 | 1.252 | -0.0465 | No | ||
| 53 | NHLH2 | 5005 | 1.233 | -0.0477 | No | ||
| 54 | EREG | 5079 | 1.216 | -0.0492 | No | ||
| 55 | CALD1 | 5086 | 1.214 | -0.0472 | No | ||
| 56 | WNK3 | 5103 | 1.210 | -0.0456 | No | ||
| 57 | PCDHA11 | 5304 | 1.161 | -0.0543 | No | ||
| 58 | SPRY4 | 5340 | 1.154 | -0.0539 | No | ||
| 59 | KCNN2 | 5344 | 1.153 | -0.0518 | No | ||
| 60 | ARID4A | 5408 | 1.137 | -0.0530 | No | ||
| 61 | RFX3 | 5569 | 1.101 | -0.0595 | No | ||
| 62 | ZBTB4 | 5592 | 1.096 | -0.0585 | No | ||
| 63 | PANX2 | 5682 | 1.081 | -0.0613 | No | ||
| 64 | ARHGAP24 | 5737 | 1.067 | -0.0621 | No | ||
| 65 | MAP3K2 | 6019 | 1.012 | -0.0754 | No | ||
| 66 | SLC1A2 | 6048 | 1.005 | -0.0750 | No | ||
| 67 | PTPN2 | 6189 | 0.974 | -0.0807 | No | ||
| 68 | MTMR4 | 6236 | 0.967 | -0.0813 | No | ||
| 69 | TNKS1BP1 | 6273 | 0.958 | -0.0814 | No | ||
| 70 | ATP2C1 | 6310 | 0.949 | -0.0815 | No | ||
| 71 | TAF5L | 6422 | 0.924 | -0.0857 | No | ||
| 72 | MOBKL2B | 6474 | 0.914 | -0.0867 | No | ||
| 73 | SLC24A4 | 6539 | 0.901 | -0.0884 | No | ||
| 74 | EPHA4 | 6559 | 0.898 | -0.0876 | No | ||
| 75 | HABP4 | 6849 | 0.840 | -0.1018 | No | ||
| 76 | TBX3 | 7021 | 0.807 | -0.1095 | No | ||
| 77 | FGF9 | 7305 | 0.747 | -0.1235 | No | ||
| 78 | NKIRAS1 | 7310 | 0.746 | -0.1222 | No | ||
| 79 | BHLHB3 | 7339 | 0.741 | -0.1223 | No | ||
| 80 | SNIP1 | 7369 | 0.735 | -0.1224 | No | ||
| 81 | SHANK2 | 7380 | 0.733 | -0.1215 | No | ||
| 82 | DNAJB9 | 7409 | 0.726 | -0.1216 | No | ||
| 83 | HMGB3 | 7473 | 0.713 | -0.1236 | No | ||
| 84 | CRSP2 | 7537 | 0.698 | -0.1257 | No | ||
| 85 | PGM2L1 | 7588 | 0.687 | -0.1270 | No | ||
| 86 | MAP3K9 | 7610 | 0.682 | -0.1268 | No | ||
| 87 | FOXA1 | 7615 | 0.681 | -0.1257 | No | ||
| 88 | TRIM3 | 7704 | 0.666 | -0.1292 | No | ||
| 89 | HIF1AN | 7747 | 0.657 | -0.1302 | No | ||
| 90 | KLF11 | 7780 | 0.650 | -0.1307 | No | ||
| 91 | PIK3R1 | 7817 | 0.642 | -0.1313 | No | ||
| 92 | HOXA3 | 7829 | 0.638 | -0.1307 | No | ||
| 93 | SMAD7 | 7830 | 0.638 | -0.1294 | No | ||
| 94 | CPEB2 | 7919 | 0.619 | -0.1330 | No | ||
| 95 | PKIA | 7960 | 0.611 | -0.1340 | No | ||
| 96 | PTPN4 | 8122 | 0.574 | -0.1416 | No | ||
| 97 | MAB21L1 | 8256 | 0.544 | -0.1478 | No | ||
| 98 | ZHX2 | 8396 | 0.512 | -0.1544 | No | ||
| 99 | VANGL1 | 8408 | 0.510 | -0.1540 | No | ||
| 100 | PCDHA10 | 8539 | 0.481 | -0.1601 | No | ||
| 101 | EIF2C1 | 8680 | 0.448 | -0.1669 | No | ||
| 102 | SLITRK3 | 8724 | 0.438 | -0.1684 | No | ||
| 103 | ST8SIA2 | 8851 | 0.406 | -0.1745 | No | ||
| 104 | STK38 | 8883 | 0.398 | -0.1754 | No | ||
| 105 | SYT1 | 8896 | 0.395 | -0.1752 | No | ||
| 106 | STYX | 8910 | 0.392 | -0.1752 | No | ||
| 107 | SKI | 8970 | 0.377 | -0.1776 | No | ||
| 108 | MAP3K5 | 8975 | 0.375 | -0.1771 | No | ||
| 109 | NF1 | 9054 | 0.358 | -0.1807 | No | ||
| 110 | TNFSF12 | 9210 | 0.320 | -0.1885 | No | ||
| 111 | HEY2 | 9399 | 0.283 | -0.1982 | No | ||
| 112 | FAM60A | 9573 | 0.245 | -0.2072 | No | ||
| 113 | MYT1 | 9627 | 0.228 | -0.2096 | No | ||
| 114 | RYR3 | 9657 | 0.221 | -0.2108 | No | ||
| 115 | RKHD1 | 9747 | 0.203 | -0.2152 | No | ||
| 116 | YES1 | 9778 | 0.197 | -0.2165 | No | ||
| 117 | PRKAA1 | 9787 | 0.194 | -0.2165 | No | ||
| 118 | ANKRD29 | 9808 | 0.189 | -0.2172 | No | ||
| 119 | FBXW11 | 9893 | 0.167 | -0.2215 | No | ||
| 120 | NPAT | 9921 | 0.159 | -0.2227 | No | ||
| 121 | NPAS2 | 10076 | 0.125 | -0.2308 | No | ||
| 122 | PCDHA12 | 10095 | 0.121 | -0.2316 | No | ||
| 123 | PARD6B | 10112 | 0.116 | -0.2322 | No | ||
| 124 | CCNJ | 10225 | 0.085 | -0.2382 | No | ||
| 125 | ZFPM2 | 10318 | 0.059 | -0.2431 | No | ||
| 126 | LRCH2 | 10419 | 0.032 | -0.2485 | No | ||
| 127 | PCDH20 | 10434 | 0.029 | -0.2492 | No | ||
| 128 | MAPRE3 | 10530 | 0.005 | -0.2544 | No | ||
| 129 | GNS | 10548 | 0.003 | -0.2553 | No | ||
| 130 | BIRC6 | 10752 | -0.051 | -0.2663 | No | ||
| 131 | SYN2 | 10839 | -0.073 | -0.2708 | No | ||
| 132 | NEUROG2 | 10913 | -0.089 | -0.2746 | No | ||
| 133 | PCDHA7 | 11006 | -0.114 | -0.2794 | No | ||
| 134 | NBL1 | 11132 | -0.145 | -0.2860 | No | ||
| 135 | BTBD7 | 11145 | -0.147 | -0.2863 | No | ||
| 136 | PTPRT | 11254 | -0.177 | -0.2919 | No | ||
| 137 | KCNJ10 | 11321 | -0.193 | -0.2951 | No | ||
| 138 | NHLH1 | 11374 | -0.205 | -0.2975 | No | ||
| 139 | USP15 | 11515 | -0.236 | -0.3047 | No | ||
| 140 | ANUBL1 | 11597 | -0.259 | -0.3086 | No | ||
| 141 | PLCB1 | 11633 | -0.268 | -0.3100 | No | ||
| 142 | BTG3 | 11698 | -0.286 | -0.3129 | No | ||
| 143 | KIAA1715 | 11723 | -0.292 | -0.3137 | No | ||
| 144 | ERBB4 | 11739 | -0.298 | -0.3139 | No | ||
| 145 | ACVR1 | 12091 | -0.393 | -0.3323 | No | ||
| 146 | KIF5A | 12122 | -0.402 | -0.3331 | No | ||
| 147 | EMX2 | 12129 | -0.403 | -0.3327 | No | ||
| 148 | PCDHA6 | 12135 | -0.404 | -0.3321 | No | ||
| 149 | GDA | 12265 | -0.439 | -0.3383 | No | ||
| 150 | MAPK4 | 12393 | -0.474 | -0.3443 | No | ||
| 151 | ABHD3 | 12425 | -0.482 | -0.3450 | No | ||
| 152 | FOXF2 | 12498 | -0.503 | -0.3480 | No | ||
| 153 | TEAD1 | 12634 | -0.542 | -0.3543 | No | ||
| 154 | SORL1 | 12697 | -0.563 | -0.3565 | No | ||
| 155 | SCAMP2 | 12713 | -0.568 | -0.3562 | No | ||
| 156 | BNC2 | 12716 | -0.569 | -0.3552 | No | ||
| 157 | SSH2 | 12718 | -0.569 | -0.3542 | No | ||
| 158 | NIPA1 | 12893 | -0.615 | -0.3624 | No | ||
| 159 | PAPOLA | 12900 | -0.615 | -0.3615 | No | ||
| 160 | GTPBP2 | 12904 | -0.616 | -0.3605 | No | ||
| 161 | RORB | 12921 | -0.620 | -0.3601 | No | ||
| 162 | ARHGAP12 | 12942 | -0.626 | -0.3600 | No | ||
| 163 | PKD2 | 12966 | -0.633 | -0.3600 | No | ||
| 164 | RTN2 | 12978 | -0.635 | -0.3593 | No | ||
| 165 | CACNB1 | 13049 | -0.657 | -0.3619 | No | ||
| 166 | STK17B | 13061 | -0.661 | -0.3611 | No | ||
| 167 | PCDHA3 | 13081 | -0.669 | -0.3609 | No | ||
| 168 | MCM7 | 13115 | -0.680 | -0.3613 | No | ||
| 169 | LMX1A | 13152 | -0.689 | -0.3619 | No | ||
| 170 | TNFRSF21 | 13300 | -0.741 | -0.3685 | No | ||
| 171 | TNFSF11 | 13408 | -0.773 | -0.3728 | No | ||
| 172 | MAP3K8 | 13443 | -0.781 | -0.3731 | No | ||
| 173 | GOPC | 13542 | -0.811 | -0.3769 | No | ||
| 174 | RASD1 | 13596 | -0.828 | -0.3781 | No | ||
| 175 | MASTL | 13659 | -0.848 | -0.3798 | No | ||
| 176 | DDX11 | 13661 | -0.849 | -0.3782 | No | ||
| 177 | DDX3Y | 13721 | -0.868 | -0.3797 | No | ||
| 178 | NFAT5 | 13837 | -0.904 | -0.3842 | No | ||
| 179 | DYRK1A | 14124 | -1.003 | -0.3978 | No | ||
| 180 | E2F7 | 14266 | -1.050 | -0.4034 | No | ||
| 181 | WDR20 | 14299 | -1.064 | -0.4031 | No | ||
| 182 | LRIG1 | 14335 | -1.076 | -0.4029 | No | ||
| 183 | QKI | 14346 | -1.078 | -0.4013 | No | ||
| 184 | BICD2 | 14411 | -1.104 | -0.4026 | No | ||
| 185 | PPP3CA | 14441 | -1.117 | -0.4020 | No | ||
| 186 | BLCAP | 14664 | -1.200 | -0.4117 | No | ||
| 187 | AKT3 | 14774 | -1.238 | -0.4152 | Yes | ||
| 188 | SOX21 | 14799 | -1.248 | -0.4140 | Yes | ||
| 189 | ARID4B | 14805 | -1.250 | -0.4118 | Yes | ||
| 190 | ATRN | 14830 | -1.265 | -0.4107 | Yes | ||
| 191 | MAP3K12 | 14938 | -1.309 | -0.4139 | Yes | ||
| 192 | SP3 | 14945 | -1.311 | -0.4116 | Yes | ||
| 193 | ASF1A | 14999 | -1.333 | -0.4119 | Yes | ||
| 194 | CRKRS | 15014 | -1.341 | -0.4100 | Yes | ||
| 195 | PFN2 | 15068 | -1.367 | -0.4102 | Yes | ||
| 196 | FJX1 | 15072 | -1.368 | -0.4076 | Yes | ||
| 197 | RTN1 | 15243 | -1.447 | -0.4141 | Yes | ||
| 198 | USP3 | 15249 | -1.449 | -0.4115 | Yes | ||
| 199 | E2F1 | 15267 | -1.457 | -0.4095 | Yes | ||
| 200 | SUMF1 | 15314 | -1.478 | -0.4091 | Yes | ||
| 201 | ATG16L1 | 15320 | -1.481 | -0.4064 | Yes | ||
| 202 | SNX5 | 15329 | -1.484 | -0.4039 | Yes | ||
| 203 | SNRK | 15401 | -1.517 | -0.4048 | Yes | ||
| 204 | R3HDM1 | 15426 | -1.527 | -0.4031 | Yes | ||
| 205 | FBXL5 | 15552 | -1.588 | -0.4068 | Yes | ||
| 206 | CHD9 | 15562 | -1.591 | -0.4041 | Yes | ||
| 207 | HECA | 15582 | -1.600 | -0.4020 | Yes | ||
| 208 | DNAJB6 | 15628 | -1.620 | -0.4012 | Yes | ||
| 209 | CELSR2 | 15630 | -1.621 | -0.3981 | Yes | ||
| 210 | STAT3 | 15651 | -1.633 | -0.3959 | Yes | ||
| 211 | HOXD8 | 15669 | -1.640 | -0.3936 | Yes | ||
| 212 | ELK3 | 15714 | -1.664 | -0.3927 | Yes | ||
| 213 | PDGFRA | 15717 | -1.665 | -0.3895 | Yes | ||
| 214 | ACSL4 | 15736 | -1.676 | -0.3872 | Yes | ||
| 215 | ERBB2IP | 15790 | -1.704 | -0.3867 | Yes | ||
| 216 | DAZAP2 | 15824 | -1.723 | -0.3851 | Yes | ||
| 217 | RAB10 | 15871 | -1.750 | -0.3842 | Yes | ||
| 218 | ITCH | 16003 | -1.837 | -0.3877 | Yes | ||
| 219 | FSTL5 | 16109 | -1.889 | -0.3897 | Yes | ||
| 220 | ZDHHC9 | 16132 | -1.905 | -0.3871 | Yes | ||
| 221 | RAPGEF4 | 16142 | -1.911 | -0.3838 | Yes | ||
| 222 | VAMP4 | 16171 | -1.925 | -0.3815 | Yes | ||
| 223 | SERTAD2 | 16188 | -1.935 | -0.3786 | Yes | ||
| 224 | ZFYVE9 | 16208 | -1.951 | -0.3758 | Yes | ||
| 225 | ANKRD50 | 16228 | -1.957 | -0.3729 | Yes | ||
| 226 | GOSR1 | 16279 | -1.986 | -0.3717 | Yes | ||
| 227 | SMOC1 | 16283 | -1.990 | -0.3679 | Yes | ||
| 228 | LACE1 | 16313 | -2.013 | -0.3655 | Yes | ||
| 229 | OGT | 16374 | -2.058 | -0.3647 | Yes | ||
| 230 | RBBP6 | 16408 | -2.077 | -0.3624 | Yes | ||
| 231 | PURB | 16416 | -2.082 | -0.3587 | Yes | ||
| 232 | STC1 | 16435 | -2.095 | -0.3555 | Yes | ||
| 233 | L3MBTL3 | 16455 | -2.108 | -0.3524 | Yes | ||
| 234 | POLQ | 16463 | -2.114 | -0.3486 | Yes | ||
| 235 | SOX4 | 16485 | -2.128 | -0.3455 | Yes | ||
| 236 | CUL3 | 16489 | -2.130 | -0.3415 | Yes | ||
| 237 | WDFY3 | 16508 | -2.150 | -0.3382 | Yes | ||
| 238 | TSG101 | 16520 | -2.162 | -0.3345 | Yes | ||
| 239 | SFMBT1 | 16580 | -2.197 | -0.3334 | Yes | ||
| 240 | MECP2 | 16612 | -2.221 | -0.3307 | Yes | ||
| 241 | BCL2L2 | 16626 | -2.230 | -0.3270 | Yes | ||
| 242 | ROCK2 | 16643 | -2.242 | -0.3234 | Yes | ||
| 243 | VLDLR | 16657 | -2.251 | -0.3197 | Yes | ||
| 244 | WEE1 | 16673 | -2.258 | -0.3160 | Yes | ||
| 245 | KIF23 | 16675 | -2.259 | -0.3116 | Yes | ||
| 246 | KPNA3 | 16694 | -2.272 | -0.3081 | Yes | ||
| 247 | STX6 | 16781 | -2.336 | -0.3081 | Yes | ||
| 248 | SMOC2 | 16804 | -2.354 | -0.3047 | Yes | ||
| 249 | IGF1 | 16894 | -2.417 | -0.3048 | Yes | ||
| 250 | SEMA4B | 16968 | -2.475 | -0.3038 | Yes | ||
| 251 | CNOT4 | 17025 | -2.520 | -0.3019 | Yes | ||
| 252 | PHTF2 | 17081 | -2.567 | -0.2998 | Yes | ||
| 253 | TBL1X | 17082 | -2.569 | -0.2947 | Yes | ||
| 254 | EFTUD1 | 17107 | -2.589 | -0.2909 | Yes | ||
| 255 | ASXL2 | 17137 | -2.617 | -0.2873 | Yes | ||
| 256 | RAD52 | 17191 | -2.673 | -0.2849 | Yes | ||
| 257 | LAMP2 | 17195 | -2.679 | -0.2798 | Yes | ||
| 258 | TNRC6A | 17242 | -2.721 | -0.2769 | Yes | ||
| 259 | CASP2 | 17256 | -2.732 | -0.2722 | Yes | ||
| 260 | DNM2 | 17371 | -2.834 | -0.2728 | Yes | ||
| 261 | ARHGEF11 | 17372 | -2.835 | -0.2672 | Yes | ||
| 262 | ADIPOR2 | 17380 | -2.838 | -0.2620 | Yes | ||
| 263 | YTHDF2 | 17382 | -2.842 | -0.2564 | Yes | ||
| 264 | MYB | 17394 | -2.858 | -0.2514 | Yes | ||
| 265 | SNX16 | 17415 | -2.878 | -0.2467 | Yes | ||
| 266 | GOLGA1 | 17481 | -2.932 | -0.2445 | Yes | ||
| 267 | ACTL6A | 17491 | -2.939 | -0.2392 | Yes | ||
| 268 | EPC2 | 17538 | -3.007 | -0.2357 | Yes | ||
| 269 | MIB1 | 17553 | -3.016 | -0.2305 | Yes | ||
| 270 | THRA | 17565 | -3.030 | -0.2251 | Yes | ||
| 271 | NFIX | 17591 | -3.058 | -0.2204 | Yes | ||
| 272 | RAP2C | 17595 | -3.061 | -0.2145 | Yes | ||
| 273 | CUGBP2 | 17636 | -3.096 | -0.2106 | Yes | ||
| 274 | RB1 | 17687 | -3.152 | -0.2071 | Yes | ||
| 275 | HBP1 | 17690 | -3.155 | -0.2009 | Yes | ||
| 276 | DDX5 | 17717 | -3.189 | -0.1960 | Yes | ||
| 277 | SFRS5 | 17737 | -3.212 | -0.1907 | Yes | ||
| 278 | DDX3X | 17776 | -3.265 | -0.1863 | Yes | ||
| 279 | SFRS2 | 17780 | -3.269 | -0.1800 | Yes | ||
| 280 | NR1D1 | 17796 | -3.281 | -0.1743 | Yes | ||
| 281 | MBNL1 | 17821 | -3.320 | -0.1691 | Yes | ||
| 282 | MAPRE1 | 17831 | -3.327 | -0.1630 | Yes | ||
| 283 | KLF9 | 17841 | -3.344 | -0.1569 | Yes | ||
| 284 | PKNOX1 | 17860 | -3.371 | -0.1512 | Yes | ||
| 285 | TOPORS | 17875 | -3.404 | -0.1452 | Yes | ||
| 286 | EFNB1 | 17893 | -3.422 | -0.1393 | Yes | ||
| 287 | PLEKHA3 | 17907 | -3.450 | -0.1332 | Yes | ||
| 288 | EIF4G2 | 17956 | -3.525 | -0.1289 | Yes | ||
| 289 | MAML1 | 18015 | -3.619 | -0.1249 | Yes | ||
| 290 | AHCTF1 | 18022 | -3.627 | -0.1180 | Yes | ||
| 291 | PIGA | 18045 | -3.660 | -0.1120 | Yes | ||
| 292 | CNOT7 | 18059 | -3.681 | -0.1054 | Yes | ||
| 293 | MMP14 | 18142 | -3.834 | -0.1023 | Yes | ||
| 294 | SOCS6 | 18167 | -3.868 | -0.0959 | Yes | ||
| 295 | ANKFY1 | 18202 | -3.933 | -0.0900 | Yes | ||
| 296 | SLC4A7 | 18208 | -3.959 | -0.0824 | Yes | ||
| 297 | CDK2 | 18234 | -4.014 | -0.0758 | Yes | ||
| 298 | PAFAH1B1 | 18307 | -4.189 | -0.0715 | Yes | ||
| 299 | MAPK6 | 18327 | -4.244 | -0.0641 | Yes | ||
| 300 | NDEL1 | 18422 | -4.555 | -0.0602 | Yes | ||
| 301 | KBTBD2 | 18430 | -4.581 | -0.0515 | Yes | ||
| 302 | SLC40A1 | 18471 | -4.701 | -0.0444 | Yes | ||
| 303 | SMAD5 | 18483 | -4.747 | -0.0356 | Yes | ||
| 304 | BRMS1L | 18487 | -4.772 | -0.0263 | Yes | ||
| 305 | CFL2 | 18494 | -4.838 | -0.0171 | Yes | ||
| 306 | ITPR1 | 18547 | -5.257 | -0.0095 | Yes | ||
| 307 | M6PR | 18604 | -6.683 | 0.0007 | Yes |
