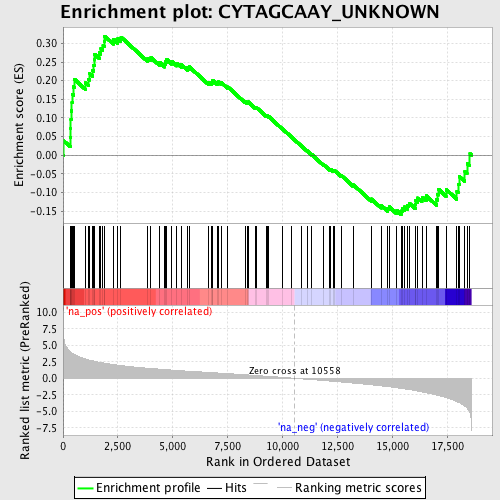
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_truncNotch_versus_absentNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | CYTAGCAAY_UNKNOWN |
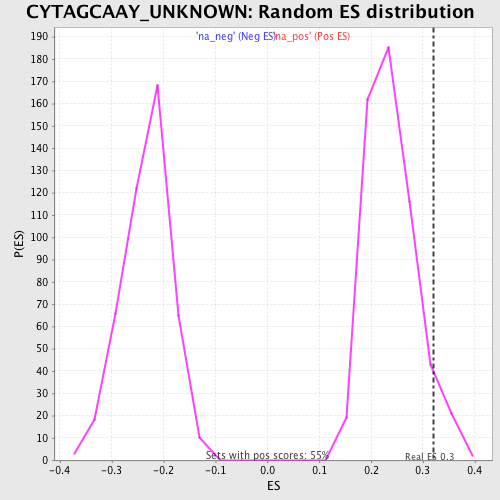
| Enrichment Score (ES) | 0.3195893 |
| Normalized Enrichment Score (NES) | 1.340376 |
| Nominal p-value | 0.06386861 |
| FDR q-value | 0.5716662 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | AP1M1 | 26 | 6.447 | 0.0387 | Yes | ||
| 2 | HRAS | 327 | 4.040 | 0.0477 | Yes | ||
| 3 | ARMC3 | 347 | 3.974 | 0.0714 | Yes | ||
| 4 | PYGB | 355 | 3.949 | 0.0956 | Yes | ||
| 5 | TPCN1 | 369 | 3.909 | 0.1192 | Yes | ||
| 6 | C19ORF20 | 403 | 3.794 | 0.1411 | Yes | ||
| 7 | USP30 | 415 | 3.773 | 0.1640 | Yes | ||
| 8 | FGR | 477 | 3.667 | 0.1835 | Yes | ||
| 9 | FKBPL | 524 | 3.570 | 0.2032 | Yes | ||
| 10 | POLR3F | 1026 | 2.896 | 0.1942 | Yes | ||
| 11 | LRP2 | 1160 | 2.779 | 0.2044 | Yes | ||
| 12 | DCTN1 | 1214 | 2.738 | 0.2185 | Yes | ||
| 13 | STRN4 | 1347 | 2.634 | 0.2278 | Yes | ||
| 14 | TRERF1 | 1394 | 2.602 | 0.2415 | Yes | ||
| 15 | GRIK5 | 1426 | 2.573 | 0.2559 | Yes | ||
| 16 | XRCC1 | 1434 | 2.566 | 0.2715 | Yes | ||
| 17 | ADCK4 | 1652 | 2.426 | 0.2749 | Yes | ||
| 18 | PRRG2 | 1722 | 2.376 | 0.2859 | Yes | ||
| 19 | RHOBTB2 | 1817 | 2.323 | 0.2953 | Yes | ||
| 20 | RGS6 | 1883 | 2.283 | 0.3060 | Yes | ||
| 21 | ASB7 | 1895 | 2.275 | 0.3196 | Yes | ||
| 22 | MRPL10 | 2293 | 2.078 | 0.3111 | No | ||
| 23 | POLR3H | 2497 | 1.990 | 0.3125 | No | ||
| 24 | HM13 | 2637 | 1.929 | 0.3170 | No | ||
| 25 | GMPR2 | 3862 | 1.522 | 0.2605 | No | ||
| 26 | SLC25A3 | 4002 | 1.487 | 0.2622 | No | ||
| 27 | GRIK2 | 4404 | 1.381 | 0.2492 | No | ||
| 28 | PARK2 | 4638 | 1.321 | 0.2448 | No | ||
| 29 | FKRP | 4678 | 1.314 | 0.2509 | No | ||
| 30 | NRGN | 4704 | 1.308 | 0.2577 | No | ||
| 31 | ATRX | 4951 | 1.248 | 0.2522 | No | ||
| 32 | GRM3 | 5189 | 1.190 | 0.2468 | No | ||
| 33 | ARL4A | 5394 | 1.141 | 0.2429 | No | ||
| 34 | TTC16 | 5674 | 1.082 | 0.2346 | No | ||
| 35 | CACNA1D | 5747 | 1.064 | 0.2373 | No | ||
| 36 | PDE1A | 6625 | 0.885 | 0.1955 | No | ||
| 37 | GABARAPL2 | 6754 | 0.861 | 0.1939 | No | ||
| 38 | TOM1L2 | 6794 | 0.852 | 0.1971 | No | ||
| 39 | MXD4 | 6811 | 0.849 | 0.2016 | No | ||
| 40 | PASK | 7023 | 0.807 | 0.1952 | No | ||
| 41 | NEDD8 | 7083 | 0.794 | 0.1970 | No | ||
| 42 | DSCAML1 | 7196 | 0.772 | 0.1957 | No | ||
| 43 | CCRK | 7508 | 0.705 | 0.1833 | No | ||
| 44 | PIK3R4 | 8302 | 0.535 | 0.1439 | No | ||
| 45 | EFNB3 | 8405 | 0.511 | 0.1415 | No | ||
| 46 | CDKL5 | 8432 | 0.505 | 0.1433 | No | ||
| 47 | ASNA1 | 8787 | 0.422 | 0.1268 | No | ||
| 48 | SCAMP4 | 8828 | 0.411 | 0.1272 | No | ||
| 49 | TUSC3 | 9249 | 0.312 | 0.1065 | No | ||
| 50 | NKX2-2 | 9293 | 0.304 | 0.1060 | No | ||
| 51 | GABRB3 | 9366 | 0.288 | 0.1040 | No | ||
| 52 | PACRG | 10003 | 0.142 | 0.0705 | No | ||
| 53 | ZFR | 10404 | 0.036 | 0.0491 | No | ||
| 54 | NPR3 | 10857 | -0.077 | 0.0252 | No | ||
| 55 | UBB | 11123 | -0.142 | 0.0118 | No | ||
| 56 | ANXA1 | 11337 | -0.197 | 0.0015 | No | ||
| 57 | ALMS1 | 11868 | -0.335 | -0.0250 | No | ||
| 58 | EIF2S1 | 12150 | -0.408 | -0.0376 | No | ||
| 59 | FOSB | 12199 | -0.421 | -0.0376 | No | ||
| 60 | TTLL4 | 12319 | -0.451 | -0.0412 | No | ||
| 61 | DNAJC1 | 12385 | -0.472 | -0.0418 | No | ||
| 62 | PPP1R7 | 12673 | -0.553 | -0.0538 | No | ||
| 63 | MDH1 | 13233 | -0.716 | -0.0795 | No | ||
| 64 | NME5 | 14045 | -0.974 | -0.1172 | No | ||
| 65 | KCNMA1 | 14499 | -1.134 | -0.1346 | No | ||
| 66 | SRP72 | 14793 | -1.245 | -0.1427 | No | ||
| 67 | SDCBP2 | 14858 | -1.275 | -0.1382 | No | ||
| 68 | IL20 | 15210 | -1.434 | -0.1482 | No | ||
| 69 | TBPL1 | 15409 | -1.519 | -0.1494 | No | ||
| 70 | HSPG2 | 15454 | -1.540 | -0.1422 | No | ||
| 71 | PES1 | 15571 | -1.595 | -0.1386 | No | ||
| 72 | SALL2 | 15684 | -1.647 | -0.1343 | No | ||
| 73 | MGST3 | 15764 | -1.686 | -0.1281 | No | ||
| 74 | SLC6A8 | 16058 | -1.858 | -0.1324 | No | ||
| 75 | CDC14A | 16071 | -1.868 | -0.1214 | No | ||
| 76 | PDE6C | 16168 | -1.923 | -0.1146 | No | ||
| 77 | DNAJB4 | 16359 | -2.049 | -0.1121 | No | ||
| 78 | CPD | 16549 | -2.177 | -0.1087 | No | ||
| 79 | SEC14L1 | 17020 | -2.514 | -0.1185 | No | ||
| 80 | CCNG2 | 17059 | -2.548 | -0.1046 | No | ||
| 81 | MARK3 | 17098 | -2.584 | -0.0906 | No | ||
| 82 | RNF138 | 17457 | -2.911 | -0.0918 | No | ||
| 83 | HMGCS1 | 17947 | -3.507 | -0.0964 | No | ||
| 84 | ATP6V1D | 18008 | -3.610 | -0.0771 | No | ||
| 85 | NEDD9 | 18057 | -3.680 | -0.0568 | No | ||
| 86 | CNTF | 18312 | -4.204 | -0.0444 | No | ||
| 87 | ACBD3 | 18412 | -4.515 | -0.0216 | No | ||
| 88 | CUTL1 | 18543 | -5.228 | 0.0039 | No |