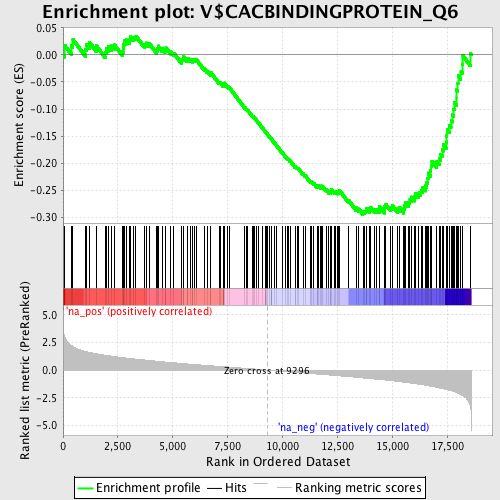
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_wtNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$CACBINDINGPROTEIN_Q6 |
| Enrichment Score (ES) | -0.2944885 |
| Normalized Enrichment Score (NES) | -1.4015098 |
| Nominal p-value | 0.015025042 |
| FDR q-value | 0.5214206 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EPC1 | 76 | 2.994 | 0.0173 | No | ||
| 2 | NCAM1 | 374 | 2.217 | 0.0170 | No | ||
| 3 | ITPR1 | 447 | 2.124 | 0.0282 | No | ||
| 4 | DMD | 1017 | 1.693 | 0.0095 | No | ||
| 5 | NFIX | 1050 | 1.676 | 0.0197 | No | ||
| 6 | CHL1 | 1210 | 1.609 | 0.0226 | No | ||
| 7 | HNRPR | 1504 | 1.479 | 0.0173 | No | ||
| 8 | LRP2 | 1914 | 1.343 | 0.0047 | No | ||
| 9 | EMID1 | 1962 | 1.327 | 0.0116 | No | ||
| 10 | ORC6L | 2051 | 1.299 | 0.0161 | No | ||
| 11 | ATF3 | 2208 | 1.257 | 0.0166 | No | ||
| 12 | NDUFS2 | 2339 | 1.227 | 0.0183 | No | ||
| 13 | POU2F1 | 2703 | 1.133 | 0.0068 | No | ||
| 14 | NEUROD2 | 2736 | 1.124 | 0.0130 | No | ||
| 15 | PIM1 | 2766 | 1.112 | 0.0194 | No | ||
| 16 | GNAO1 | 2792 | 1.105 | 0.0259 | No | ||
| 17 | KIRREL2 | 2903 | 1.079 | 0.0277 | No | ||
| 18 | LASP1 | 3042 | 1.050 | 0.0277 | No | ||
| 19 | PBX1 | 3075 | 1.043 | 0.0334 | No | ||
| 20 | RTN2 | 3219 | 1.013 | 0.0329 | No | ||
| 21 | PIP5K2B | 3319 | 0.992 | 0.0346 | No | ||
| 22 | SYT6 | 3709 | 0.914 | 0.0200 | No | ||
| 23 | MAP1A | 3792 | 0.896 | 0.0220 | No | ||
| 24 | ZNF385 | 3923 | 0.868 | 0.0211 | No | ||
| 25 | RAB6A | 4264 | 0.803 | 0.0084 | No | ||
| 26 | HOXA10 | 4315 | 0.794 | 0.0114 | No | ||
| 27 | LRFN5 | 4327 | 0.791 | 0.0165 | No | ||
| 28 | YWHAQ | 4513 | 0.753 | 0.0118 | No | ||
| 29 | NR1H3 | 4646 | 0.730 | 0.0099 | No | ||
| 30 | LRP1 | 4688 | 0.723 | 0.0128 | No | ||
| 31 | RAB11B | 4897 | 0.687 | 0.0064 | No | ||
| 32 | JPH4 | 5042 | 0.661 | 0.0033 | No | ||
| 33 | AHR | 5393 | 0.597 | -0.0114 | No | ||
| 34 | ARHGAP5 | 5409 | 0.595 | -0.0079 | No | ||
| 35 | CSDA | 5471 | 0.585 | -0.0071 | No | ||
| 36 | KPNB1 | 5484 | 0.583 | -0.0035 | No | ||
| 37 | HOXB9 | 5662 | 0.557 | -0.0092 | No | ||
| 38 | PDGFB | 5688 | 0.553 | -0.0066 | No | ||
| 39 | MNT | 5796 | 0.534 | -0.0086 | No | ||
| 40 | SDC1 | 5886 | 0.519 | -0.0097 | No | ||
| 41 | AKT2 | 5918 | 0.515 | -0.0077 | No | ||
| 42 | PPP1CB | 6002 | 0.500 | -0.0086 | No | ||
| 43 | KIF3C | 6064 | 0.491 | -0.0084 | No | ||
| 44 | ITGA6 | 6435 | 0.438 | -0.0254 | No | ||
| 45 | USF2 | 6585 | 0.414 | -0.0305 | No | ||
| 46 | SLC2A4 | 6719 | 0.395 | -0.0349 | No | ||
| 47 | RALBP1 | 6735 | 0.393 | -0.0329 | No | ||
| 48 | ACVRL1 | 7110 | 0.338 | -0.0507 | No | ||
| 49 | MEF2D | 7158 | 0.331 | -0.0509 | No | ||
| 50 | MYBPH | 7312 | 0.313 | -0.0570 | No | ||
| 51 | PYGO1 | 7313 | 0.313 | -0.0547 | No | ||
| 52 | KLF13 | 7314 | 0.313 | -0.0525 | No | ||
| 53 | HTATIP | 7344 | 0.308 | -0.0519 | No | ||
| 54 | DCTN1 | 7493 | 0.287 | -0.0578 | No | ||
| 55 | KCNS3 | 7560 | 0.277 | -0.0594 | No | ||
| 56 | PICALM | 8280 | 0.162 | -0.0973 | No | ||
| 57 | CLCN6 | 8356 | 0.151 | -0.1002 | No | ||
| 58 | S100A14 | 8418 | 0.143 | -0.1025 | No | ||
| 59 | PABPN1 | 8624 | 0.113 | -0.1128 | No | ||
| 60 | RTN4RL2 | 8658 | 0.109 | -0.1138 | No | ||
| 61 | MYH10 | 8722 | 0.098 | -0.1166 | No | ||
| 62 | KIF1C | 8792 | 0.085 | -0.1197 | No | ||
| 63 | MGAT3 | 8927 | 0.060 | -0.1265 | No | ||
| 64 | LIN28 | 9081 | 0.032 | -0.1346 | No | ||
| 65 | IGFBP5 | 9208 | 0.014 | -0.1413 | No | ||
| 66 | NMT1 | 9211 | 0.014 | -0.1413 | No | ||
| 67 | NXPH3 | 9220 | 0.012 | -0.1417 | No | ||
| 68 | MTHFR | 9268 | 0.005 | -0.1442 | No | ||
| 69 | PROCR | 9304 | -0.002 | -0.1461 | No | ||
| 70 | FZD4 | 9407 | -0.018 | -0.1515 | No | ||
| 71 | EIF4G1 | 9476 | -0.028 | -0.1549 | No | ||
| 72 | KNTC1 | 9479 | -0.029 | -0.1548 | No | ||
| 73 | NOL4 | 9480 | -0.029 | -0.1546 | No | ||
| 74 | CASKIN2 | 9628 | -0.050 | -0.1622 | No | ||
| 75 | NRGN | 9728 | -0.063 | -0.1672 | No | ||
| 76 | THBS3 | 9991 | -0.105 | -0.1806 | No | ||
| 77 | PRKAG1 | 10148 | -0.132 | -0.1881 | No | ||
| 78 | VPS35 | 10240 | -0.147 | -0.1920 | No | ||
| 79 | EMX2 | 10262 | -0.150 | -0.1921 | No | ||
| 80 | NEDD4L | 10350 | -0.165 | -0.1956 | No | ||
| 81 | ADAMTS9 | 10580 | -0.197 | -0.2066 | No | ||
| 82 | ABI3 | 10583 | -0.198 | -0.2053 | No | ||
| 83 | PHACTR3 | 10678 | -0.215 | -0.2089 | No | ||
| 84 | HOXB8 | 10708 | -0.220 | -0.2089 | No | ||
| 85 | IVD | 10935 | -0.250 | -0.2193 | No | ||
| 86 | SUPT16H | 11032 | -0.265 | -0.2227 | No | ||
| 87 | UBE2B | 11267 | -0.300 | -0.2332 | No | ||
| 88 | GDPD2 | 11305 | -0.307 | -0.2330 | No | ||
| 89 | TITF1 | 11427 | -0.328 | -0.2372 | No | ||
| 90 | ZADH2 | 11575 | -0.353 | -0.2427 | No | ||
| 91 | HPCAL4 | 11600 | -0.356 | -0.2414 | No | ||
| 92 | RAB6B | 11644 | -0.362 | -0.2412 | No | ||
| 93 | MAGED2 | 11733 | -0.376 | -0.2433 | No | ||
| 94 | RALB | 11757 | -0.380 | -0.2418 | No | ||
| 95 | HOXC4 | 11823 | -0.391 | -0.2425 | No | ||
| 96 | ARPC2 | 11987 | -0.418 | -0.2484 | No | ||
| 97 | ARHGEF12 | 12112 | -0.439 | -0.2520 | No | ||
| 98 | APOA2 | 12183 | -0.448 | -0.2526 | No | ||
| 99 | NTF5 | 12206 | -0.453 | -0.2505 | No | ||
| 100 | INPP4A | 12232 | -0.457 | -0.2486 | No | ||
| 101 | MARK2 | 12372 | -0.482 | -0.2527 | No | ||
| 102 | NR4A3 | 12418 | -0.489 | -0.2517 | No | ||
| 103 | DPF1 | 12513 | -0.504 | -0.2532 | No | ||
| 104 | HEBP2 | 12563 | -0.513 | -0.2522 | No | ||
| 105 | ADAMTS4 | 12597 | -0.520 | -0.2502 | No | ||
| 106 | BCL7A | 13012 | -0.587 | -0.2685 | No | ||
| 107 | FKHL18 | 13349 | -0.645 | -0.2821 | No | ||
| 108 | CHRM1 | 13478 | -0.669 | -0.2842 | No | ||
| 109 | KRTCAP2 | 13668 | -0.705 | -0.2895 | Yes | ||
| 110 | TNNC1 | 13739 | -0.718 | -0.2881 | Yes | ||
| 111 | CACNA1D | 13806 | -0.730 | -0.2865 | Yes | ||
| 112 | PCTK2 | 13834 | -0.734 | -0.2827 | Yes | ||
| 113 | DCN | 13985 | -0.760 | -0.2854 | Yes | ||
| 114 | PHF12 | 14012 | -0.767 | -0.2814 | Yes | ||
| 115 | DLX1 | 14195 | -0.802 | -0.2855 | Yes | ||
| 116 | BCL11A | 14288 | -0.819 | -0.2846 | Yes | ||
| 117 | RNF12 | 14405 | -0.842 | -0.2849 | Yes | ||
| 118 | NCDN | 14420 | -0.846 | -0.2796 | Yes | ||
| 119 | EEF1G | 14647 | -0.885 | -0.2856 | Yes | ||
| 120 | HIF1A | 14654 | -0.887 | -0.2796 | Yes | ||
| 121 | KCNN3 | 14709 | -0.899 | -0.2761 | Yes | ||
| 122 | AP4S1 | 14925 | -0.948 | -0.2810 | Yes | ||
| 123 | RFX1 | 14999 | -0.963 | -0.2780 | Yes | ||
| 124 | STRN3 | 15244 | -1.020 | -0.2840 | Yes | ||
| 125 | ACVR1B | 15338 | -1.042 | -0.2816 | Yes | ||
| 126 | NEGR1 | 15530 | -1.088 | -0.2842 | Yes | ||
| 127 | FGF11 | 15551 | -1.094 | -0.2774 | Yes | ||
| 128 | ZFYVE1 | 15613 | -1.109 | -0.2728 | Yes | ||
| 129 | KLK12 | 15751 | -1.146 | -0.2721 | Yes | ||
| 130 | NOTCH2 | 15808 | -1.158 | -0.2669 | Yes | ||
| 131 | ACIN1 | 15879 | -1.180 | -0.2622 | Yes | ||
| 132 | EEF1A1 | 16018 | -1.216 | -0.2610 | Yes | ||
| 133 | ATP6V0C | 16070 | -1.228 | -0.2550 | Yes | ||
| 134 | DCTN2 | 16219 | -1.270 | -0.2540 | Yes | ||
| 135 | TRIM46 | 16311 | -1.298 | -0.2496 | Yes | ||
| 136 | HR | 16391 | -1.322 | -0.2445 | Yes | ||
| 137 | SIX4 | 16515 | -1.366 | -0.2414 | Yes | ||
| 138 | PRRX1 | 16578 | -1.384 | -0.2349 | Yes | ||
| 139 | DTNA | 16625 | -1.401 | -0.2274 | Yes | ||
| 140 | EFNB3 | 16636 | -1.404 | -0.2179 | Yes | ||
| 141 | ZSWIM3 | 16744 | -1.446 | -0.2134 | Yes | ||
| 142 | ARHGEF17 | 16767 | -1.456 | -0.2042 | Yes | ||
| 143 | GNB4 | 16808 | -1.470 | -0.1959 | Yes | ||
| 144 | AP1G1 | 17034 | -1.558 | -0.1969 | Yes | ||
| 145 | ARF3 | 17149 | -1.608 | -0.1916 | Yes | ||
| 146 | SALL1 | 17215 | -1.632 | -0.1835 | Yes | ||
| 147 | ROCK1 | 17278 | -1.656 | -0.1751 | Yes | ||
| 148 | MTX1 | 17324 | -1.676 | -0.1655 | Yes | ||
| 149 | MKL1 | 17463 | -1.737 | -0.1606 | Yes | ||
| 150 | TP53 | 17474 | -1.741 | -0.1487 | Yes | ||
| 151 | FBS1 | 17496 | -1.753 | -0.1374 | Yes | ||
| 152 | FOXA2 | 17616 | -1.817 | -0.1308 | Yes | ||
| 153 | ADRA1D | 17682 | -1.855 | -0.1211 | Yes | ||
| 154 | GNGT2 | 17735 | -1.889 | -0.1105 | Yes | ||
| 155 | OTX2 | 17802 | -1.935 | -0.1002 | Yes | ||
| 156 | KCNJ14 | 17829 | -1.948 | -0.0877 | Yes | ||
| 157 | ACR | 17934 | -2.017 | -0.0790 | Yes | ||
| 158 | HOXB6 | 17935 | -2.018 | -0.0646 | Yes | ||
| 159 | PKIA | 17993 | -2.070 | -0.0529 | Yes | ||
| 160 | HOXC6 | 18002 | -2.076 | -0.0385 | Yes | ||
| 161 | GRK5 | 18133 | -2.207 | -0.0298 | Yes | ||
| 162 | CACNA1G | 18205 | -2.289 | -0.0173 | Yes | ||
| 163 | FBXO36 | 18219 | -2.311 | -0.0015 | Yes | ||
| 164 | AAMP | 18558 | -3.218 | 0.0031 | Yes |
