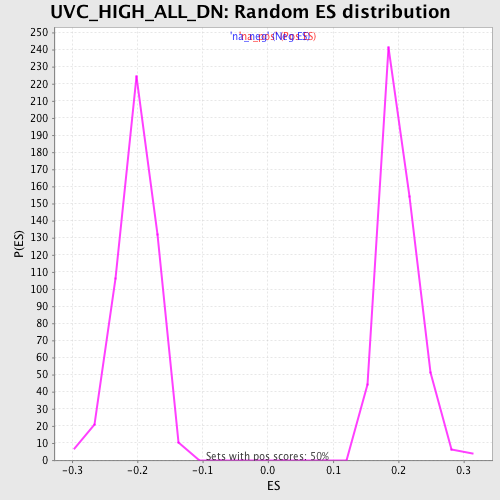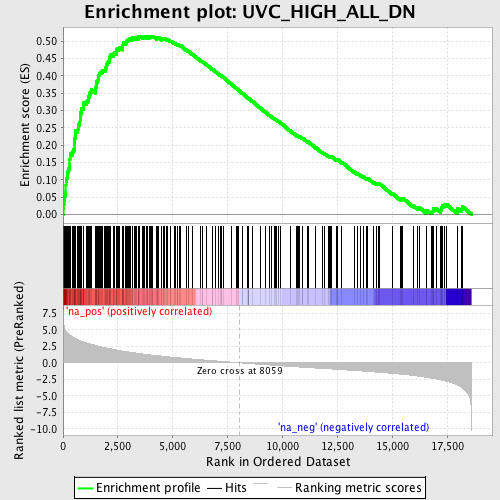
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_truncNotch |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | UVC_HIGH_ALL_DN |
| Enrichment Score (ES) | 0.51475036 |
| Normalized Enrichment Score (NES) | 2.568672 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PTK2 | 5 | 7.263 | 0.0167 | Yes | ||
| 2 | CDYL | 20 | 6.035 | 0.0300 | Yes | ||
| 3 | LBR | 43 | 5.600 | 0.0419 | Yes | ||
| 4 | SS18 | 56 | 5.427 | 0.0539 | Yes | ||
| 5 | EMP1 | 107 | 4.902 | 0.0627 | Yes | ||
| 6 | THUMPD1 | 127 | 4.780 | 0.0728 | Yes | ||
| 7 | NFYB | 128 | 4.777 | 0.0840 | Yes | ||
| 8 | RAB14 | 132 | 4.754 | 0.0949 | Yes | ||
| 9 | DUSP11 | 157 | 4.673 | 0.1045 | Yes | ||
| 10 | RNF4 | 182 | 4.588 | 0.1139 | Yes | ||
| 11 | NASP | 217 | 4.468 | 0.1225 | Yes | ||
| 12 | CDK8 | 229 | 4.436 | 0.1323 | Yes | ||
| 13 | MAPK6 | 289 | 4.244 | 0.1390 | Yes | ||
| 14 | SEC24B | 306 | 4.204 | 0.1479 | Yes | ||
| 15 | PAFAH1B1 | 309 | 4.189 | 0.1576 | Yes | ||
| 16 | UBE2G1 | 315 | 4.175 | 0.1671 | Yes | ||
| 17 | ATP2A2 | 317 | 4.171 | 0.1768 | Yes | ||
| 18 | BNIP2 | 420 | 3.922 | 0.1804 | Yes | ||
| 19 | RAF1 | 468 | 3.836 | 0.1868 | Yes | ||
| 20 | PITPNB | 508 | 3.764 | 0.1935 | Yes | ||
| 21 | TLK1 | 510 | 3.763 | 0.2022 | Yes | ||
| 22 | MAT2A | 518 | 3.752 | 0.2106 | Yes | ||
| 23 | SPOP | 529 | 3.732 | 0.2188 | Yes | ||
| 24 | CHD1 | 549 | 3.690 | 0.2264 | Yes | ||
| 25 | RAB1A | 560 | 3.680 | 0.2344 | Yes | ||
| 26 | RARS | 582 | 3.648 | 0.2418 | Yes | ||
| 27 | CRKL | 677 | 3.501 | 0.2449 | Yes | ||
| 28 | GTF2E1 | 683 | 3.496 | 0.2528 | Yes | ||
| 29 | BMI1 | 691 | 3.473 | 0.2605 | Yes | ||
| 30 | PARP1 | 762 | 3.364 | 0.2646 | Yes | ||
| 31 | MPHOSPH6 | 769 | 3.353 | 0.2721 | Yes | ||
| 32 | KLF9 | 775 | 3.344 | 0.2796 | Yes | ||
| 33 | KPNA2 | 779 | 3.339 | 0.2872 | Yes | ||
| 34 | BRD8 | 793 | 3.322 | 0.2943 | Yes | ||
| 35 | DDX3X | 840 | 3.265 | 0.2994 | Yes | ||
| 36 | CNOT2 | 843 | 3.262 | 0.3069 | Yes | ||
| 37 | RBM5 | 914 | 3.169 | 0.3105 | Yes | ||
| 38 | PPP1R12A | 918 | 3.166 | 0.3178 | Yes | ||
| 39 | REV3L | 948 | 3.128 | 0.3235 | Yes | ||
| 40 | BMPR1A | 1049 | 3.031 | 0.3251 | Yes | ||
| 41 | STX7 | 1109 | 2.959 | 0.3288 | Yes | ||
| 42 | RYBP | 1158 | 2.912 | 0.3330 | Yes | ||
| 43 | TLK2 | 1162 | 2.905 | 0.3397 | Yes | ||
| 44 | PLK4 | 1191 | 2.890 | 0.3449 | Yes | ||
| 45 | TNFAIP3 | 1211 | 2.873 | 0.3506 | Yes | ||
| 46 | TLE4 | 1269 | 2.814 | 0.3540 | Yes | ||
| 47 | PUM2 | 1273 | 2.810 | 0.3604 | Yes | ||
| 48 | CBX3 | 1470 | 2.632 | 0.3559 | Yes | ||
| 49 | TAF11 | 1472 | 2.628 | 0.3620 | Yes | ||
| 50 | MARCKS | 1488 | 2.603 | 0.3673 | Yes | ||
| 51 | IFNGR2 | 1529 | 2.572 | 0.3711 | Yes | ||
| 52 | TBL1X | 1534 | 2.569 | 0.3769 | Yes | ||
| 53 | PHTF2 | 1535 | 2.567 | 0.3829 | Yes | ||
| 54 | ZBED4 | 1559 | 2.546 | 0.3876 | Yes | ||
| 55 | CDR2 | 1593 | 2.517 | 0.3917 | Yes | ||
| 56 | SFRS3 | 1608 | 2.503 | 0.3968 | Yes | ||
| 57 | USP1 | 1620 | 2.495 | 0.4020 | Yes | ||
| 58 | SQLE | 1644 | 2.477 | 0.4065 | Yes | ||
| 59 | CSTF3 | 1681 | 2.449 | 0.4103 | Yes | ||
| 60 | RAD21 | 1763 | 2.389 | 0.4115 | Yes | ||
| 61 | EIF4E | 1784 | 2.371 | 0.4159 | Yes | ||
| 62 | FOSL2 | 1868 | 2.308 | 0.4168 | Yes | ||
| 63 | KIF23 | 1941 | 2.259 | 0.4182 | Yes | ||
| 64 | WEE1 | 1943 | 2.258 | 0.4234 | Yes | ||
| 65 | PRKCD | 1974 | 2.241 | 0.4270 | Yes | ||
| 66 | EMG1 | 1984 | 2.233 | 0.4317 | Yes | ||
| 67 | HMGCR | 2000 | 2.222 | 0.4361 | Yes | ||
| 68 | E2F3 | 2042 | 2.193 | 0.4390 | Yes | ||
| 69 | RUNX1 | 2071 | 2.175 | 0.4425 | Yes | ||
| 70 | MAPK14 | 2121 | 2.137 | 0.4449 | Yes | ||
| 71 | TOP2A | 2126 | 2.130 | 0.4496 | Yes | ||
| 72 | CCNA2 | 2128 | 2.129 | 0.4545 | Yes | ||
| 73 | TNFAIP8 | 2168 | 2.104 | 0.4573 | Yes | ||
| 74 | AMD1 | 2180 | 2.096 | 0.4616 | Yes | ||
| 75 | RIPK1 | 2288 | 2.022 | 0.4605 | Yes | ||
| 76 | CYR61 | 2314 | 2.003 | 0.4639 | Yes | ||
| 77 | UGCG | 2342 | 1.982 | 0.4670 | Yes | ||
| 78 | FYN | 2423 | 1.940 | 0.4672 | Yes | ||
| 79 | PFTK1 | 2426 | 1.937 | 0.4716 | Yes | ||
| 80 | SERTAD2 | 2428 | 1.935 | 0.4761 | Yes | ||
| 81 | ATP6V1B2 | 2461 | 1.916 | 0.4788 | Yes | ||
| 82 | PPAP2B | 2543 | 1.868 | 0.4788 | Yes | ||
| 83 | SNX4 | 2557 | 1.858 | 0.4824 | Yes | ||
| 84 | UBTF | 2702 | 1.772 | 0.4787 | Yes | ||
| 85 | PKP4 | 2708 | 1.767 | 0.4826 | Yes | ||
| 86 | SLC20A2 | 2710 | 1.766 | 0.4867 | Yes | ||
| 87 | CUL1 | 2729 | 1.757 | 0.4898 | Yes | ||
| 88 | NTF3 | 2741 | 1.751 | 0.4933 | Yes | ||
| 89 | NFE2L2 | 2747 | 1.748 | 0.4971 | Yes | ||
| 90 | SMURF2 | 2861 | 1.683 | 0.4949 | Yes | ||
| 91 | STRN3 | 2870 | 1.679 | 0.4984 | Yes | ||
| 92 | PDGFRA | 2899 | 1.665 | 0.5007 | Yes | ||
| 93 | STX3 | 2945 | 1.641 | 0.5021 | Yes | ||
| 94 | SSH1 | 2970 | 1.629 | 0.5046 | Yes | ||
| 95 | CDC6 | 3003 | 1.614 | 0.5067 | Yes | ||
| 96 | CCNG1 | 3051 | 1.592 | 0.5078 | Yes | ||
| 97 | NRIP1 | 3141 | 1.549 | 0.5066 | Yes | ||
| 98 | PIP5K3 | 3142 | 1.549 | 0.5102 | Yes | ||
| 99 | RAB11A | 3252 | 1.504 | 0.5078 | Yes | ||
| 100 | ITGA6 | 3288 | 1.484 | 0.5094 | Yes | ||
| 101 | HMGB2 | 3330 | 1.469 | 0.5106 | Yes | ||
| 102 | STXBP3 | 3419 | 1.427 | 0.5091 | Yes | ||
| 103 | NCOA3 | 3425 | 1.422 | 0.5122 | Yes | ||
| 104 | CITED2 | 3446 | 1.410 | 0.5144 | Yes | ||
| 105 | KRIT1 | 3501 | 1.384 | 0.5147 | Yes | ||
| 106 | TM4SF1 | 3639 | 1.322 | 0.5103 | Yes | ||
| 107 | POGZ | 3644 | 1.319 | 0.5132 | Yes | ||
| 108 | TNFRSF1A | 3701 | 1.299 | 0.5132 | Yes | ||
| 109 | AURKA | 3787 | 1.264 | 0.5115 | Yes | ||
| 110 | DICER1 | 3830 | 1.242 | 0.5121 | Yes | ||
| 111 | TPST1 | 3838 | 1.239 | 0.5146 | Yes | ||
| 112 | BLCAP | 3952 | 1.200 | 0.5113 | Yes | ||
| 113 | COIL | 3967 | 1.192 | 0.5133 | Yes | ||
| 114 | IL4R | 4022 | 1.170 | 0.5131 | Yes | ||
| 115 | ARL4C | 4091 | 1.144 | 0.5121 | Yes | ||
| 116 | GNAI1 | 4092 | 1.143 | 0.5148 | Yes | ||
| 117 | FUS | 4273 | 1.078 | 0.5075 | No | ||
| 118 | UBE2N | 4280 | 1.076 | 0.5097 | No | ||
| 119 | SWAP70 | 4323 | 1.062 | 0.5099 | No | ||
| 120 | PDCD10 | 4342 | 1.054 | 0.5113 | No | ||
| 121 | DYRK1A | 4492 | 1.003 | 0.5056 | No | ||
| 122 | FGF5 | 4493 | 1.003 | 0.5079 | No | ||
| 123 | STAM | 4560 | 0.977 | 0.5066 | No | ||
| 124 | FHL5 | 4579 | 0.970 | 0.5079 | No | ||
| 125 | BHLHB2 | 4633 | 0.952 | 0.5073 | No | ||
| 126 | TOPBP1 | 4715 | 0.926 | 0.5050 | No | ||
| 127 | DR1 | 4749 | 0.914 | 0.5054 | No | ||
| 128 | ZFAND5 | 4889 | 0.870 | 0.4998 | No | ||
| 129 | CASP8 | 4914 | 0.862 | 0.5005 | No | ||
| 130 | MET | 5064 | 0.814 | 0.4943 | No | ||
| 131 | CASP3 | 5125 | 0.796 | 0.4929 | No | ||
| 132 | GBAS | 5219 | 0.770 | 0.4897 | No | ||
| 133 | TARDBP | 5221 | 0.768 | 0.4914 | No | ||
| 134 | IDI1 | 5318 | 0.741 | 0.4879 | No | ||
| 135 | RHOB | 5342 | 0.730 | 0.4884 | No | ||
| 136 | GEM | 5613 | 0.642 | 0.4752 | No | ||
| 137 | BACH1 | 5730 | 0.612 | 0.4703 | No | ||
| 138 | TSPAN5 | 5894 | 0.570 | 0.4628 | No | ||
| 139 | SCHIP1 | 6279 | 0.458 | 0.4430 | No | ||
| 140 | ADM | 6332 | 0.444 | 0.4412 | No | ||
| 141 | RND3 | 6352 | 0.439 | 0.4412 | No | ||
| 142 | GATA2 | 6542 | 0.390 | 0.4318 | No | ||
| 143 | SLBP | 6790 | 0.324 | 0.4192 | No | ||
| 144 | TIPARP | 6828 | 0.312 | 0.4179 | No | ||
| 145 | VEGFC | 6943 | 0.280 | 0.4123 | No | ||
| 146 | MFAP1 | 7059 | 0.248 | 0.4067 | No | ||
| 147 | PODXL | 7183 | 0.216 | 0.4005 | No | ||
| 148 | ARHGEF7 | 7199 | 0.214 | 0.4002 | No | ||
| 149 | XPO1 | 7218 | 0.210 | 0.3997 | No | ||
| 150 | EDG2 | 7313 | 0.188 | 0.3950 | No | ||
| 151 | PLEKHC1 | 7659 | 0.100 | 0.3765 | No | ||
| 152 | STK24 | 7911 | 0.038 | 0.3629 | No | ||
| 153 | PNN | 7957 | 0.025 | 0.3605 | No | ||
| 154 | RRS1 | 8001 | 0.013 | 0.3582 | No | ||
| 155 | CTGF | 8162 | -0.024 | 0.3496 | No | ||
| 156 | PDE4B | 8170 | -0.025 | 0.3492 | No | ||
| 157 | CKS2 | 8401 | -0.087 | 0.3369 | No | ||
| 158 | YY1 | 8443 | -0.100 | 0.3349 | No | ||
| 159 | DUSP1 | 8464 | -0.107 | 0.3341 | No | ||
| 160 | NUP153 | 8465 | -0.107 | 0.3344 | No | ||
| 161 | RRAS2 | 8468 | -0.107 | 0.3345 | No | ||
| 162 | GAS1 | 8615 | -0.143 | 0.3269 | No | ||
| 163 | EPS8 | 8977 | -0.225 | 0.3078 | No | ||
| 164 | SAMD4A | 9203 | -0.280 | 0.2962 | No | ||
| 165 | SATB2 | 9416 | -0.321 | 0.2854 | No | ||
| 166 | ZFP36L2 | 9483 | -0.335 | 0.2826 | No | ||
| 167 | TRIM32 | 9639 | -0.375 | 0.2751 | No | ||
| 168 | CENPA | 9683 | -0.387 | 0.2736 | No | ||
| 169 | STK38 | 9733 | -0.398 | 0.2719 | No | ||
| 170 | IRS1 | 9803 | -0.418 | 0.2691 | No | ||
| 171 | PMP22 | 9902 | -0.442 | 0.2648 | No | ||
| 172 | FZD7 | 10346 | -0.542 | 0.2420 | No | ||
| 173 | SPEN | 10640 | -0.606 | 0.2275 | No | ||
| 174 | GREM1 | 10662 | -0.612 | 0.2278 | No | ||
| 175 | AGTR1 | 10744 | -0.629 | 0.2249 | No | ||
| 176 | SMAD7 | 10786 | -0.638 | 0.2241 | No | ||
| 177 | TUBGCP3 | 10891 | -0.662 | 0.2200 | No | ||
| 178 | TRIB2 | 10916 | -0.667 | 0.2203 | No | ||
| 179 | CTDSP2 | 11159 | -0.716 | 0.2088 | No | ||
| 180 | FBXO46 | 11169 | -0.718 | 0.2100 | No | ||
| 181 | CXCL12 | 11524 | -0.791 | 0.1926 | No | ||
| 182 | KIF2A | 11815 | -0.851 | 0.1788 | No | ||
| 183 | CDK7 | 11914 | -0.871 | 0.1755 | No | ||
| 184 | CEBPD | 12086 | -0.904 | 0.1683 | No | ||
| 185 | ARID5B | 12148 | -0.915 | 0.1671 | No | ||
| 186 | PPP2R5C | 12176 | -0.921 | 0.1678 | No | ||
| 187 | DDIT4 | 12211 | -0.928 | 0.1681 | No | ||
| 188 | SIAH2 | 12463 | -0.982 | 0.1568 | No | ||
| 189 | HERPUD1 | 12510 | -0.991 | 0.1566 | No | ||
| 190 | UAP1 | 12523 | -0.994 | 0.1583 | No | ||
| 191 | CD2AP | 12709 | -1.034 | 0.1506 | No | ||
| 192 | ZHX3 | 13270 | -1.152 | 0.1229 | No | ||
| 193 | POLE2 | 13417 | -1.186 | 0.1177 | No | ||
| 194 | HOXB2 | 13556 | -1.219 | 0.1130 | No | ||
| 195 | BCAR3 | 13691 | -1.256 | 0.1087 | No | ||
| 196 | ARF6 | 13830 | -1.291 | 0.1042 | No | ||
| 197 | DLX2 | 13888 | -1.303 | 0.1041 | No | ||
| 198 | SMAD3 | 14165 | -1.369 | 0.0923 | No | ||
| 199 | HMGA2 | 14303 | -1.407 | 0.0882 | No | ||
| 200 | NFIL3 | 14358 | -1.420 | 0.0886 | No | ||
| 201 | HDHD1A | 14404 | -1.433 | 0.0895 | No | ||
| 202 | THBS1 | 14996 | -1.590 | 0.0610 | No | ||
| 203 | MTMR3 | 15375 | -1.708 | 0.0445 | No | ||
| 204 | NR2F2 | 15439 | -1.727 | 0.0451 | No | ||
| 205 | ELF4 | 15483 | -1.739 | 0.0468 | No | ||
| 206 | MLX | 15956 | -1.920 | 0.0256 | No | ||
| 207 | USP10 | 16171 | -2.014 | 0.0187 | No | ||
| 208 | NCK1 | 16253 | -2.044 | 0.0191 | No | ||
| 209 | LHFPL2 | 16542 | -2.181 | 0.0085 | No | ||
| 210 | SPRED2 | 16576 | -2.203 | 0.0119 | No | ||
| 211 | HAS2 | 16772 | -2.306 | 0.0066 | No | ||
| 212 | ACVR1B | 16826 | -2.338 | 0.0092 | No | ||
| 213 | TRIP10 | 16864 | -2.359 | 0.0127 | No | ||
| 214 | TLE1 | 16890 | -2.374 | 0.0169 | No | ||
| 215 | GNE | 16996 | -2.446 | 0.0169 | No | ||
| 216 | UNG | 17203 | -2.589 | 0.0118 | No | ||
| 217 | TWIST1 | 17225 | -2.604 | 0.0167 | No | ||
| 218 | PTPN1 | 17258 | -2.625 | 0.0211 | No | ||
| 219 | SRF | 17295 | -2.658 | 0.0253 | No | ||
| 220 | FADD | 17370 | -2.718 | 0.0277 | No | ||
| 221 | FZD2 | 17482 | -2.803 | 0.0282 | No | ||
| 222 | NAP1L1 | 17970 | -3.356 | 0.0095 | No | ||
| 223 | IGF2BP3 | 17985 | -3.386 | 0.0167 | No | ||
| 224 | PHC2 | 18165 | -3.704 | 0.0156 | No | ||
| 225 | LSM2 | 18207 | -3.790 | 0.0222 | No |