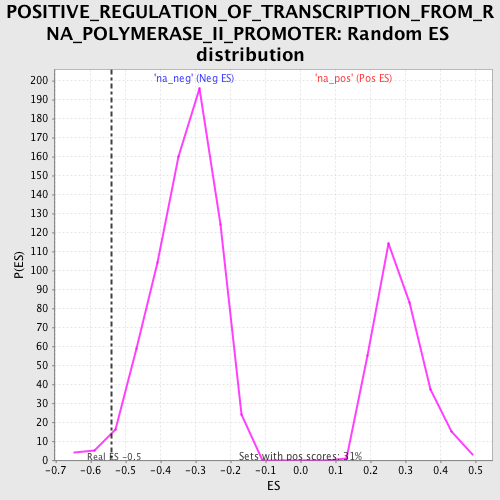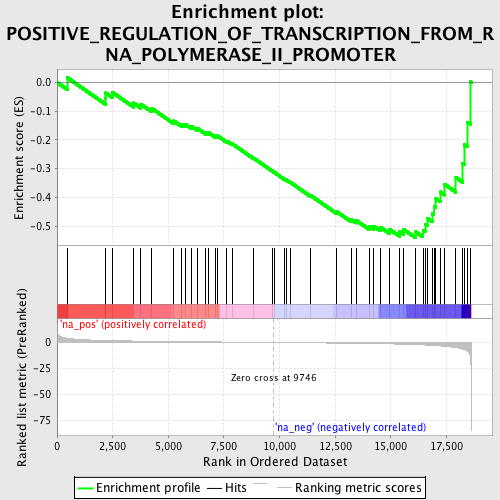
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | POSITIVE_REGULATION_OF_TRANSCRIPTION_FROM_RNA_POLYMERASE_II_PROMOTER |
| Enrichment Score (ES) | -0.5392461 |
| Normalized Enrichment Score (NES) | -1.6357237 |
| Nominal p-value | 0.01300578 |
| FDR q-value | 0.57174265 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCNH | 453 | 3.967 | 0.0167 | No | ||
| 2 | GTF2H1 | 2175 | 1.914 | -0.0561 | No | ||
| 3 | ATF4 | 2184 | 1.910 | -0.0367 | No | ||
| 4 | CLOCK | 2497 | 1.756 | -0.0353 | No | ||
| 5 | NPAS2 | 3437 | 1.384 | -0.0715 | No | ||
| 6 | GLIS1 | 3767 | 1.280 | -0.0760 | No | ||
| 7 | MKL2 | 4240 | 1.144 | -0.0895 | No | ||
| 8 | NRIP1 | 5242 | 0.899 | -0.1341 | No | ||
| 9 | ELL3 | 5610 | 0.819 | -0.1454 | No | ||
| 10 | MNAT1 | 5756 | 0.788 | -0.1450 | No | ||
| 11 | IL4 | 6034 | 0.730 | -0.1523 | No | ||
| 12 | PPARGC1B | 6312 | 0.674 | -0.1603 | No | ||
| 13 | GLI2 | 6683 | 0.598 | -0.1740 | No | ||
| 14 | TP53 | 6813 | 0.567 | -0.1751 | No | ||
| 15 | FOXD3 | 7106 | 0.510 | -0.1855 | No | ||
| 16 | CDK7 | 7199 | 0.492 | -0.1854 | No | ||
| 17 | THRAP3 | 7624 | 0.409 | -0.2039 | No | ||
| 18 | NUFIP1 | 7876 | 0.359 | -0.2137 | No | ||
| 19 | MED6 | 8834 | 0.183 | -0.2634 | No | ||
| 20 | EPAS1 | 9692 | 0.010 | -0.3094 | No | ||
| 21 | MYO6 | 9775 | -0.008 | -0.3138 | No | ||
| 22 | RBM14 | 10213 | -0.104 | -0.3362 | No | ||
| 23 | BMP6 | 10289 | -0.123 | -0.3390 | No | ||
| 24 | RXRA | 10474 | -0.163 | -0.3472 | No | ||
| 25 | UTF1 | 11372 | -0.344 | -0.3919 | No | ||
| 26 | SP1 | 12543 | -0.615 | -0.4486 | No | ||
| 27 | ELF1 | 13220 | -0.790 | -0.4768 | No | ||
| 28 | FOXH1 | 13448 | -0.853 | -0.4802 | No | ||
| 29 | FOXO3 | 14022 | -1.039 | -0.5002 | No | ||
| 30 | GLI1 | 14225 | -1.110 | -0.4996 | No | ||
| 31 | HIF1A | 14529 | -1.213 | -0.5033 | No | ||
| 32 | PLAGL1 | 14924 | -1.368 | -0.5104 | No | ||
| 33 | HNF4A | 15389 | -1.590 | -0.5189 | Yes | ||
| 34 | ERCC3 | 15553 | -1.669 | -0.5103 | Yes | ||
| 35 | SMAD2 | 16091 | -1.995 | -0.5186 | Yes | ||
| 36 | RUNX1 | 16446 | -2.312 | -0.5136 | Yes | ||
| 37 | ELF4 | 16551 | -2.416 | -0.4942 | Yes | ||
| 38 | NCOA6 | 16646 | -2.503 | -0.4733 | Yes | ||
| 39 | SMAD3 | 16869 | -2.768 | -0.4565 | Yes | ||
| 40 | MED12 | 16940 | -2.853 | -0.4307 | Yes | ||
| 41 | SQSTM1 | 17027 | -2.961 | -0.4046 | Yes | ||
| 42 | SUPT4H1 | 17216 | -3.254 | -0.3810 | Yes | ||
| 43 | SUPT5H | 17415 | -3.614 | -0.3542 | Yes | ||
| 44 | EPC1 | 17925 | -4.900 | -0.3308 | Yes | ||
| 45 | TNFRSF1A | 18237 | -6.230 | -0.2829 | Yes | ||
| 46 | MAML1 | 18306 | -6.748 | -0.2166 | Yes | ||
| 47 | ARNTL | 18452 | -8.287 | -0.1384 | Yes | ||
| 48 | ERCC2 | 18575 | -14.195 | 0.0022 | Yes |