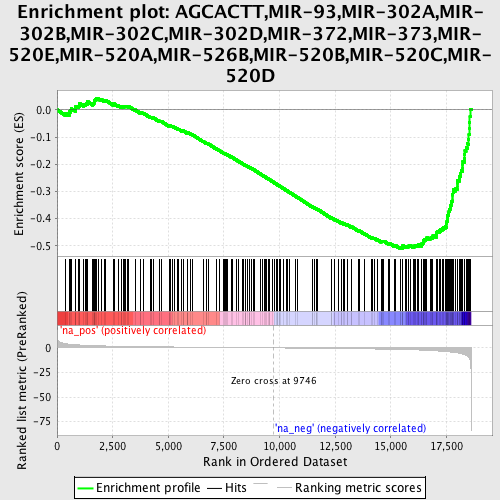
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set03_absentNotch_versus_normalThy |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | AGCACTT,MIR-93,MIR-302A,MIR-302B,MIR-302C,MIR-302D,MIR-372,MIR-373,MIR-520E,MIR-520A,MIR-526B,MIR-520B,MIR-520C,MIR-520D |
| Enrichment Score (ES) | -0.51136106 |
| Normalized Enrichment Score (NES) | -1.8793578 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.059345614 |
| FWER p-Value | 0.274 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | OSBPL5 | 387 | 4.201 | -0.0123 | No | ||
| 2 | AK2 | 542 | 3.762 | -0.0129 | No | ||
| 3 | ATAD2 | 544 | 3.756 | -0.0051 | No | ||
| 4 | SLC6A9 | 614 | 3.564 | -0.0014 | No | ||
| 5 | DNAJA2 | 624 | 3.545 | 0.0054 | No | ||
| 6 | CRK | 816 | 3.207 | 0.0017 | No | ||
| 7 | APP | 833 | 3.187 | 0.0075 | No | ||
| 8 | TRIM36 | 840 | 3.174 | 0.0138 | No | ||
| 9 | SDC1 | 954 | 2.982 | 0.0138 | No | ||
| 10 | MARCH8 | 996 | 2.933 | 0.0177 | No | ||
| 11 | LEFTY1 | 1005 | 2.923 | 0.0233 | No | ||
| 12 | ADAM9 | 1198 | 2.666 | 0.0184 | No | ||
| 13 | TRPS1 | 1265 | 2.576 | 0.0202 | No | ||
| 14 | RBBP7 | 1300 | 2.547 | 0.0236 | No | ||
| 15 | RPS6KA1 | 1354 | 2.492 | 0.0259 | No | ||
| 16 | DDHD1 | 1370 | 2.473 | 0.0303 | No | ||
| 17 | PBX3 | 1611 | 2.259 | 0.0219 | No | ||
| 18 | INHBB | 1641 | 2.237 | 0.0250 | No | ||
| 19 | PPP3R1 | 1664 | 2.223 | 0.0284 | No | ||
| 20 | VLDLR | 1667 | 2.221 | 0.0329 | No | ||
| 21 | TNFAIP1 | 1698 | 2.200 | 0.0359 | No | ||
| 22 | YWHAZ | 1722 | 2.183 | 0.0392 | No | ||
| 23 | ZFP36L2 | 1748 | 2.164 | 0.0423 | No | ||
| 24 | RPS6KA3 | 1863 | 2.086 | 0.0404 | No | ||
| 25 | MRPS25 | 1973 | 2.026 | 0.0387 | No | ||
| 26 | RGMA | 2132 | 1.936 | 0.0342 | No | ||
| 27 | ETV1 | 2183 | 1.911 | 0.0354 | No | ||
| 28 | CFL2 | 2542 | 1.735 | 0.0196 | No | ||
| 29 | NEO1 | 2566 | 1.725 | 0.0219 | No | ||
| 30 | POLQ | 2753 | 1.642 | 0.0152 | No | ||
| 31 | NR4A2 | 2905 | 1.582 | 0.0103 | No | ||
| 32 | ESR1 | 2907 | 1.581 | 0.0135 | No | ||
| 33 | CXXC6 | 2970 | 1.556 | 0.0134 | No | ||
| 34 | EDNRB | 3029 | 1.533 | 0.0134 | No | ||
| 35 | DRD1 | 3085 | 1.516 | 0.0136 | No | ||
| 36 | ANKRD13C | 3145 | 1.494 | 0.0135 | No | ||
| 37 | PHF6 | 3224 | 1.467 | 0.0123 | No | ||
| 38 | ASF1A | 3505 | 1.364 | -0.0001 | No | ||
| 39 | CUGBP2 | 3736 | 1.287 | -0.0099 | No | ||
| 40 | CCND1 | 3753 | 1.284 | -0.0081 | No | ||
| 41 | DAZAP2 | 3867 | 1.248 | -0.0117 | No | ||
| 42 | ECT2 | 4203 | 1.152 | -0.0275 | No | ||
| 43 | RABEP1 | 4246 | 1.143 | -0.0274 | No | ||
| 44 | PCDHA11 | 4311 | 1.126 | -0.0285 | No | ||
| 45 | NEK9 | 4594 | 1.057 | -0.0417 | No | ||
| 46 | SENP1 | 4621 | 1.051 | -0.0409 | No | ||
| 47 | NPAS3 | 4685 | 1.034 | -0.0422 | No | ||
| 48 | BCL2L11 | 5035 | 0.951 | -0.0592 | No | ||
| 49 | NFIB | 5052 | 0.947 | -0.0581 | No | ||
| 50 | ZFPM2 | 5090 | 0.938 | -0.0581 | No | ||
| 51 | CDCA7 | 5180 | 0.919 | -0.0611 | No | ||
| 52 | DCUN1D1 | 5259 | 0.894 | -0.0634 | No | ||
| 53 | SYNC1 | 5408 | 0.860 | -0.0697 | No | ||
| 54 | OLFM3 | 5462 | 0.849 | -0.0708 | No | ||
| 55 | LASS6 | 5611 | 0.818 | -0.0772 | No | ||
| 56 | KCNMA1 | 5658 | 0.808 | -0.0780 | No | ||
| 57 | RHOC | 5669 | 0.806 | -0.0768 | No | ||
| 58 | DPYSL5 | 5846 | 0.769 | -0.0848 | No | ||
| 59 | PLXNA1 | 5866 | 0.766 | -0.0843 | No | ||
| 60 | RBL1 | 5879 | 0.763 | -0.0833 | No | ||
| 61 | DMTF1 | 6010 | 0.735 | -0.0889 | No | ||
| 62 | RSBN1 | 6080 | 0.723 | -0.0911 | No | ||
| 63 | PCDHA12 | 6570 | 0.620 | -0.1164 | No | ||
| 64 | LHX8 | 6706 | 0.592 | -0.1225 | No | ||
| 65 | UBE2R2 | 6733 | 0.586 | -0.1227 | No | ||
| 66 | TIPARP | 6808 | 0.568 | -0.1255 | No | ||
| 67 | PCDHA10 | 7146 | 0.502 | -0.1428 | No | ||
| 68 | TAOK2 | 7288 | 0.474 | -0.1495 | No | ||
| 69 | ZKSCAN1 | 7494 | 0.434 | -0.1597 | No | ||
| 70 | ST8SIA2 | 7541 | 0.427 | -0.1614 | No | ||
| 71 | VSX1 | 7583 | 0.417 | -0.1627 | No | ||
| 72 | OSBPL8 | 7634 | 0.407 | -0.1646 | No | ||
| 73 | RDBP | 7670 | 0.401 | -0.1657 | No | ||
| 74 | MLLT6 | 7817 | 0.372 | -0.1728 | No | ||
| 75 | SSX2IP | 7865 | 0.361 | -0.1746 | No | ||
| 76 | SLC2A4 | 8044 | 0.328 | -0.1836 | No | ||
| 77 | PCDHA3 | 8138 | 0.310 | -0.1880 | No | ||
| 78 | RALGDS | 8318 | 0.280 | -0.1972 | No | ||
| 79 | MEF2C | 8378 | 0.266 | -0.1998 | No | ||
| 80 | PRDM8 | 8455 | 0.255 | -0.2034 | No | ||
| 81 | TLOC1 | 8475 | 0.249 | -0.2040 | No | ||
| 82 | WEE1 | 8488 | 0.246 | -0.2041 | No | ||
| 83 | BRMS1L | 8570 | 0.228 | -0.2080 | No | ||
| 84 | RABGAP1 | 8643 | 0.214 | -0.2115 | No | ||
| 85 | FOXL2 | 8646 | 0.214 | -0.2112 | No | ||
| 86 | KPNA2 | 8668 | 0.210 | -0.2119 | No | ||
| 87 | ZBTB4 | 8757 | 0.194 | -0.2162 | No | ||
| 88 | BRP44L | 8815 | 0.186 | -0.2190 | No | ||
| 89 | UBE2J1 | 8880 | 0.176 | -0.2221 | No | ||
| 90 | TARDBP | 9136 | 0.129 | -0.2357 | No | ||
| 91 | ALX4 | 9226 | 0.112 | -0.2403 | No | ||
| 92 | ANK2 | 9341 | 0.088 | -0.2463 | No | ||
| 93 | BAMBI | 9379 | 0.082 | -0.2481 | No | ||
| 94 | ZBTB11 | 9430 | 0.071 | -0.2507 | No | ||
| 95 | MBNL2 | 9499 | 0.056 | -0.2543 | No | ||
| 96 | PURB | 9550 | 0.045 | -0.2569 | No | ||
| 97 | MMP24 | 9551 | 0.044 | -0.2568 | No | ||
| 98 | EPAS1 | 9692 | 0.010 | -0.2644 | No | ||
| 99 | TMEM16F | 9776 | -0.008 | -0.2689 | No | ||
| 100 | SAR1B | 9857 | -0.023 | -0.2732 | No | ||
| 101 | PURA | 9884 | -0.030 | -0.2746 | No | ||
| 102 | ABCA1 | 9904 | -0.033 | -0.2755 | No | ||
| 103 | RAD23B | 9976 | -0.054 | -0.2793 | No | ||
| 104 | RNF6 | 9985 | -0.056 | -0.2796 | No | ||
| 105 | PCDHA5 | 10034 | -0.066 | -0.2821 | No | ||
| 106 | RPS6KA5 | 10181 | -0.097 | -0.2898 | No | ||
| 107 | NEUROD1 | 10311 | -0.126 | -0.2965 | No | ||
| 108 | DHX40 | 10339 | -0.135 | -0.2977 | No | ||
| 109 | SPOP | 10432 | -0.154 | -0.3024 | No | ||
| 110 | PLEKHA3 | 10721 | -0.208 | -0.3176 | No | ||
| 111 | TRPV6 | 10826 | -0.232 | -0.3228 | No | ||
| 112 | NTN4 | 11468 | -0.366 | -0.3569 | No | ||
| 113 | MKNK2 | 11471 | -0.366 | -0.3562 | No | ||
| 114 | RGL1 | 11586 | -0.392 | -0.3616 | No | ||
| 115 | PCDHA7 | 11655 | -0.408 | -0.3645 | No | ||
| 116 | SUV39H1 | 11677 | -0.413 | -0.3648 | No | ||
| 117 | PAK7 | 11698 | -0.418 | -0.3650 | No | ||
| 118 | CYP26B1 | 12312 | -0.559 | -0.3971 | No | ||
| 119 | MYT1L | 12313 | -0.560 | -0.3960 | No | ||
| 120 | FLT1 | 12483 | -0.600 | -0.4039 | No | ||
| 121 | LHX6 | 12484 | -0.600 | -0.4027 | No | ||
| 122 | HLF | 12647 | -0.638 | -0.4101 | No | ||
| 123 | CCNJ | 12789 | -0.674 | -0.4164 | No | ||
| 124 | CREB5 | 12801 | -0.678 | -0.4156 | No | ||
| 125 | HN1 | 12890 | -0.700 | -0.4189 | No | ||
| 126 | ORMDL3 | 12927 | -0.710 | -0.4194 | No | ||
| 127 | FRMD4A | 13043 | -0.741 | -0.4241 | No | ||
| 128 | ARID4A | 13045 | -0.742 | -0.4226 | No | ||
| 129 | EPHA2 | 13246 | -0.797 | -0.4318 | No | ||
| 130 | LUC7L2 | 13251 | -0.798 | -0.4304 | No | ||
| 131 | RB1CC1 | 13561 | -0.889 | -0.4454 | No | ||
| 132 | MTUS1 | 13586 | -0.900 | -0.4448 | No | ||
| 133 | FGD5 | 13830 | -0.974 | -0.4560 | No | ||
| 134 | PGBD5 | 14117 | -1.069 | -0.4693 | No | ||
| 135 | PARP8 | 14187 | -1.096 | -0.4708 | No | ||
| 136 | PRRX1 | 14255 | -1.121 | -0.4721 | No | ||
| 137 | SLITRK3 | 14397 | -1.166 | -0.4773 | No | ||
| 138 | ATP2B2 | 14561 | -1.225 | -0.4836 | No | ||
| 139 | MNT | 14597 | -1.236 | -0.4830 | No | ||
| 140 | NEUROD6 | 14647 | -1.254 | -0.4830 | No | ||
| 141 | ARHGEF17 | 14689 | -1.270 | -0.4826 | No | ||
| 142 | PLAG1 | 14914 | -1.364 | -0.4920 | No | ||
| 143 | PAPOLA | 14933 | -1.372 | -0.4901 | No | ||
| 144 | MUTED | 15162 | -1.471 | -0.4994 | No | ||
| 145 | E2F5 | 15231 | -1.500 | -0.5000 | No | ||
| 146 | RAB11FIP1 | 15441 | -1.616 | -0.5080 | Yes | ||
| 147 | FNDC3A | 15455 | -1.624 | -0.5053 | Yes | ||
| 148 | MTMR3 | 15520 | -1.651 | -0.5054 | Yes | ||
| 149 | CRIM1 | 15531 | -1.657 | -0.5025 | Yes | ||
| 150 | GABPB2 | 15540 | -1.661 | -0.4995 | Yes | ||
| 151 | UBE2B | 15649 | -1.725 | -0.5017 | Yes | ||
| 152 | YPEL2 | 15699 | -1.750 | -0.5008 | Yes | ||
| 153 | PCDHA6 | 15775 | -1.801 | -0.5011 | Yes | ||
| 154 | RAB6A | 15814 | -1.824 | -0.4994 | Yes | ||
| 155 | KLHL18 | 15890 | -1.875 | -0.4996 | Yes | ||
| 156 | HIC2 | 16009 | -1.942 | -0.5019 | Yes | ||
| 157 | OXR1 | 16055 | -1.971 | -0.5003 | Yes | ||
| 158 | SMAD2 | 16091 | -1.995 | -0.4980 | Yes | ||
| 159 | LATS2 | 16196 | -2.095 | -0.4993 | Yes | ||
| 160 | ZNF385 | 16254 | -2.139 | -0.4980 | Yes | ||
| 161 | YTHDF3 | 16256 | -2.142 | -0.4936 | Yes | ||
| 162 | TUSC2 | 16393 | -2.265 | -0.4963 | Yes | ||
| 163 | LEF1 | 16397 | -2.266 | -0.4917 | Yes | ||
| 164 | RUNX1 | 16446 | -2.312 | -0.4895 | Yes | ||
| 165 | NFYA | 16453 | -2.318 | -0.4850 | Yes | ||
| 166 | ZMYND11 | 16465 | -2.332 | -0.4808 | Yes | ||
| 167 | MAP3K14 | 16495 | -2.360 | -0.4774 | Yes | ||
| 168 | RAB11A | 16568 | -2.430 | -0.4763 | Yes | ||
| 169 | NR2C2 | 16597 | -2.456 | -0.4727 | Yes | ||
| 170 | PCDHA1 | 16623 | -2.478 | -0.4689 | Yes | ||
| 171 | KLF12 | 16764 | -2.638 | -0.4711 | Yes | ||
| 172 | TAL1 | 16816 | -2.703 | -0.4682 | Yes | ||
| 173 | PPP6C | 16891 | -2.802 | -0.4664 | Yes | ||
| 174 | MAPK9 | 16893 | -2.804 | -0.4606 | Yes | ||
| 175 | DUSP2 | 17039 | -2.987 | -0.4623 | Yes | ||
| 176 | MINK1 | 17042 | -2.989 | -0.4562 | Yes | ||
| 177 | NR4A3 | 17062 | -3.004 | -0.4510 | Yes | ||
| 178 | SF4 | 17091 | -3.054 | -0.4462 | Yes | ||
| 179 | MAP3K11 | 17183 | -3.201 | -0.4444 | Yes | ||
| 180 | ASF1B | 17233 | -3.287 | -0.4403 | Yes | ||
| 181 | PCAF | 17305 | -3.404 | -0.4371 | Yes | ||
| 182 | PERQ1 | 17370 | -3.536 | -0.4332 | Yes | ||
| 183 | ZDHHC9 | 17441 | -3.665 | -0.4294 | Yes | ||
| 184 | MECP2 | 17498 | -3.820 | -0.4245 | Yes | ||
| 185 | ARID4B | 17499 | -3.822 | -0.4165 | Yes | ||
| 186 | IRF2 | 17520 | -3.862 | -0.4096 | Yes | ||
| 187 | RRAGD | 17537 | -3.908 | -0.4023 | Yes | ||
| 188 | FOXJ2 | 17549 | -3.921 | -0.3948 | Yes | ||
| 189 | CCND2 | 17556 | -3.932 | -0.3869 | Yes | ||
| 190 | ULK1 | 17590 | -3.987 | -0.3804 | Yes | ||
| 191 | ATXN1 | 17611 | -4.029 | -0.3731 | Yes | ||
| 192 | CREBL1 | 17643 | -4.123 | -0.3663 | Yes | ||
| 193 | IQWD1 | 17667 | -4.174 | -0.3588 | Yes | ||
| 194 | TOX | 17686 | -4.236 | -0.3510 | Yes | ||
| 195 | FEM1C | 17714 | -4.300 | -0.3435 | Yes | ||
| 196 | SNF1LK | 17729 | -4.328 | -0.3353 | Yes | ||
| 197 | WDR37 | 17771 | -4.472 | -0.3282 | Yes | ||
| 198 | COL23A1 | 17773 | -4.476 | -0.3190 | Yes | ||
| 199 | RAB22A | 17775 | -4.485 | -0.3097 | Yes | ||
| 200 | ARHGEF10 | 17794 | -4.514 | -0.3013 | Yes | ||
| 201 | AEBP2 | 17820 | -4.582 | -0.2931 | Yes | ||
| 202 | MKRN1 | 17922 | -4.891 | -0.2884 | Yes | ||
| 203 | WDR45 | 17982 | -5.054 | -0.2811 | Yes | ||
| 204 | PLAGL2 | 17986 | -5.060 | -0.2708 | Yes | ||
| 205 | KLF13 | 17992 | -5.096 | -0.2605 | Yes | ||
| 206 | PLEKHM1 | 18071 | -5.381 | -0.2535 | Yes | ||
| 207 | MCL1 | 18097 | -5.495 | -0.2434 | Yes | ||
| 208 | E2F7 | 18127 | -5.625 | -0.2333 | Yes | ||
| 209 | RASSF2 | 18154 | -5.765 | -0.2228 | Yes | ||
| 210 | ARHGEF18 | 18224 | -6.140 | -0.2137 | Yes | ||
| 211 | SS18L1 | 18232 | -6.212 | -0.2012 | Yes | ||
| 212 | TOB2 | 18239 | -6.235 | -0.1886 | Yes | ||
| 213 | MAML1 | 18306 | -6.748 | -0.1781 | Yes | ||
| 214 | IHPK1 | 18319 | -6.870 | -0.1645 | Yes | ||
| 215 | PFN2 | 18332 | -6.976 | -0.1506 | Yes | ||
| 216 | BCL11A | 18381 | -7.407 | -0.1378 | Yes | ||
| 217 | ZFYVE26 | 18439 | -8.137 | -0.1240 | Yes | ||
| 218 | HRBL | 18494 | -9.033 | -0.1082 | Yes | ||
| 219 | BCL6 | 18511 | -9.435 | -0.0894 | Yes | ||
| 220 | BTG1 | 18531 | -10.306 | -0.0690 | Yes | ||
| 221 | PHF2 | 18539 | -10.641 | -0.0473 | Yes | ||
| 222 | GNB5 | 18557 | -11.559 | -0.0242 | Yes | ||
| 223 | TLE4 | 18572 | -13.147 | 0.0024 | Yes |
