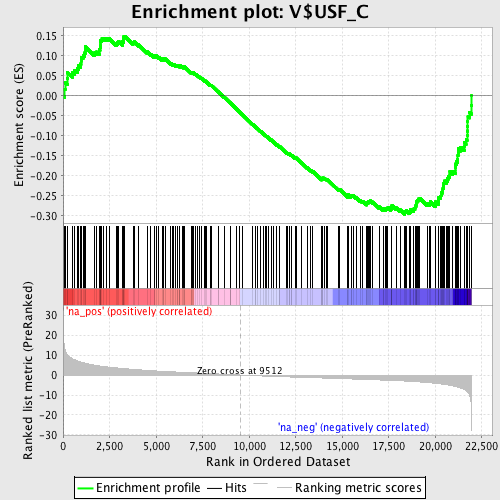
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_BT_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$USF_C |
| Enrichment Score (ES) | -0.29622185 |
| Normalized Enrichment Score (NES) | -1.4948487 |
| Nominal p-value | 0.003076923 |
| FDR q-value | 0.14463234 |
| FWER p-Value | 0.845 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BCL6 | 77 | 13.547 | 0.0165 | No | ||
| 2 | RORC | 108 | 12.686 | 0.0340 | No | ||
| 3 | ARF6 | 219 | 10.345 | 0.0443 | No | ||
| 4 | HSPBAP1 | 245 | 10.027 | 0.0580 | No | ||
| 5 | GABARAP | 487 | 8.239 | 0.0591 | No | ||
| 6 | H3F3A | 611 | 7.649 | 0.0648 | No | ||
| 7 | CRMP1 | 768 | 6.999 | 0.0680 | No | ||
| 8 | SEMA7A | 817 | 6.836 | 0.0759 | No | ||
| 9 | CCDC6 | 941 | 6.492 | 0.0799 | No | ||
| 10 | MXD4 | 962 | 6.423 | 0.0885 | No | ||
| 11 | WBP2 | 992 | 6.356 | 0.0966 | No | ||
| 12 | CNOT4 | 1072 | 6.146 | 0.1021 | No | ||
| 13 | DNMT3A | 1150 | 5.973 | 0.1074 | No | ||
| 14 | PFN1 | 1183 | 5.900 | 0.1147 | No | ||
| 15 | UBE1 | 1195 | 5.864 | 0.1228 | No | ||
| 16 | CAMK4 | 1688 | 4.913 | 0.1075 | No | ||
| 17 | TEF | 1791 | 4.765 | 0.1099 | No | ||
| 18 | ANXA6 | 1937 | 4.556 | 0.1100 | No | ||
| 19 | MYST3 | 1939 | 4.555 | 0.1167 | No | ||
| 20 | GRK6 | 1983 | 4.483 | 0.1213 | No | ||
| 21 | PB1 | 2000 | 4.460 | 0.1272 | No | ||
| 22 | PER1 | 2010 | 4.448 | 0.1334 | No | ||
| 23 | FKBP11 | 2017 | 4.437 | 0.1397 | No | ||
| 24 | EIF4E | 2088 | 4.365 | 0.1429 | No | ||
| 25 | BCAS3 | 2191 | 4.231 | 0.1445 | No | ||
| 26 | ZFP91 | 2331 | 4.083 | 0.1442 | No | ||
| 27 | RUNX2 | 2474 | 3.916 | 0.1435 | No | ||
| 28 | SIRT1 | 2856 | 3.557 | 0.1312 | No | ||
| 29 | ADCY3 | 2916 | 3.504 | 0.1337 | No | ||
| 30 | HOXC13 | 2966 | 3.464 | 0.1366 | No | ||
| 31 | RAPGEF6 | 3189 | 3.272 | 0.1312 | No | ||
| 32 | SLC26A2 | 3203 | 3.257 | 0.1355 | No | ||
| 33 | MYCL1 | 3237 | 3.234 | 0.1387 | No | ||
| 34 | BCL9L | 3239 | 3.231 | 0.1435 | No | ||
| 35 | SC5DL | 3258 | 3.220 | 0.1474 | No | ||
| 36 | GPD1 | 3317 | 3.167 | 0.1494 | No | ||
| 37 | SLC39A7 | 3757 | 2.835 | 0.1335 | No | ||
| 38 | PDK2 | 3808 | 2.796 | 0.1353 | No | ||
| 39 | RXRB | 4050 | 2.615 | 0.1281 | No | ||
| 40 | FCHSD2 | 4507 | 2.321 | 0.1106 | No | ||
| 41 | CHRM1 | 4720 | 2.189 | 0.1041 | No | ||
| 42 | BTBD5 | 4899 | 2.097 | 0.0990 | No | ||
| 43 | ELK1 | 4900 | 2.097 | 0.1021 | No | ||
| 44 | SNCAIP | 5007 | 2.028 | 0.1002 | No | ||
| 45 | TMEM15 | 5136 | 1.960 | 0.0972 | No | ||
| 46 | DIABLO | 5333 | 1.861 | 0.0910 | No | ||
| 47 | SLC20A1 | 5392 | 1.837 | 0.0910 | No | ||
| 48 | PFDN2 | 5407 | 1.830 | 0.0931 | No | ||
| 49 | AVP | 5478 | 1.787 | 0.0925 | No | ||
| 50 | ETV4 | 5767 | 1.656 | 0.0818 | No | ||
| 51 | IPO13 | 5878 | 1.600 | 0.0791 | No | ||
| 52 | LRP8 | 5950 | 1.569 | 0.0781 | No | ||
| 53 | RPL13A | 6038 | 1.526 | 0.0764 | No | ||
| 54 | BMP2K | 6122 | 1.491 | 0.0748 | No | ||
| 55 | CTSF | 6132 | 1.486 | 0.0766 | No | ||
| 56 | IL1RAPL1 | 6231 | 1.436 | 0.0742 | No | ||
| 57 | HMGN2 | 6245 | 1.428 | 0.0757 | No | ||
| 58 | HERPUD1 | 6260 | 1.422 | 0.0772 | No | ||
| 59 | POGK | 6397 | 1.362 | 0.0729 | No | ||
| 60 | BMP4 | 6472 | 1.325 | 0.0715 | No | ||
| 61 | MPP3 | 6502 | 1.308 | 0.0721 | No | ||
| 62 | MNT | 6512 | 1.303 | 0.0736 | No | ||
| 63 | DNAJB9 | 6892 | 1.125 | 0.0579 | No | ||
| 64 | JMJD1A | 6903 | 1.121 | 0.0591 | No | ||
| 65 | NTN2L | 6931 | 1.109 | 0.0595 | No | ||
| 66 | ABCA1 | 7002 | 1.080 | 0.0579 | No | ||
| 67 | SLC12A5 | 7123 | 1.028 | 0.0539 | No | ||
| 68 | GPX1 | 7219 | 0.991 | 0.0510 | No | ||
| 69 | PPARGC1B | 7326 | 0.939 | 0.0475 | No | ||
| 70 | RPL19 | 7419 | 0.900 | 0.0446 | No | ||
| 71 | CLCN2 | 7608 | 0.825 | 0.0372 | No | ||
| 72 | MCM8 | 7627 | 0.817 | 0.0375 | No | ||
| 73 | CENTB5 | 7722 | 0.772 | 0.0344 | No | ||
| 74 | ADNP | 7904 | 0.701 | 0.0271 | No | ||
| 75 | SLC4A11 | 7929 | 0.692 | 0.0270 | No | ||
| 76 | RPS19 | 7962 | 0.677 | 0.0265 | No | ||
| 77 | HYAL2 | 8338 | 0.521 | 0.0101 | No | ||
| 78 | HOXA4 | 8349 | 0.517 | 0.0104 | No | ||
| 79 | GTF2A1 | 8677 | 0.379 | -0.0041 | No | ||
| 80 | STMN1 | 8975 | 0.246 | -0.0174 | No | ||
| 81 | GTF2H1 | 8990 | 0.237 | -0.0177 | No | ||
| 82 | DAZAP1 | 8991 | 0.236 | -0.0174 | No | ||
| 83 | FHOD1 | 9336 | 0.092 | -0.0330 | No | ||
| 84 | BCKDHA | 9489 | 0.015 | -0.0400 | No | ||
| 85 | POLR2H | 9651 | -0.067 | -0.0473 | No | ||
| 86 | PSMB3 | 10201 | -0.297 | -0.0721 | No | ||
| 87 | PLA2G4A | 10313 | -0.343 | -0.0767 | No | ||
| 88 | HS3ST3A1 | 10335 | -0.354 | -0.0772 | No | ||
| 89 | EIF2C2 | 10446 | -0.387 | -0.0817 | No | ||
| 90 | NR1D1 | 10596 | -0.438 | -0.0879 | No | ||
| 91 | TLL1 | 10780 | -0.507 | -0.0955 | No | ||
| 92 | TBX4 | 10872 | -0.537 | -0.0989 | No | ||
| 93 | CDC14A | 10918 | -0.555 | -0.1002 | No | ||
| 94 | ELOVL4 | 11035 | -0.591 | -0.1046 | No | ||
| 95 | ARHGAP20 | 11170 | -0.645 | -0.1098 | No | ||
| 96 | ARRDC3 | 11277 | -0.676 | -0.1137 | No | ||
| 97 | MYOHD1 | 11484 | -0.744 | -0.1221 | No | ||
| 98 | EFNB1 | 11615 | -0.789 | -0.1269 | No | ||
| 99 | SGTB | 11640 | -0.796 | -0.1268 | No | ||
| 100 | GPRC5C | 12001 | -0.922 | -0.1420 | No | ||
| 101 | SLC6A15 | 12049 | -0.935 | -0.1428 | No | ||
| 102 | ARX | 12137 | -0.959 | -0.1454 | No | ||
| 103 | SATB2 | 12145 | -0.960 | -0.1442 | No | ||
| 104 | VGF | 12255 | -0.993 | -0.1478 | No | ||
| 105 | NPTX1 | 12293 | -1.004 | -0.1480 | No | ||
| 106 | CHST11 | 12496 | -1.067 | -0.1557 | No | ||
| 107 | BHLHB2 | 12513 | -1.071 | -0.1549 | No | ||
| 108 | ALDH3B1 | 12824 | -1.165 | -0.1674 | No | ||
| 109 | FLT3 | 13124 | -1.253 | -0.1793 | No | ||
| 110 | GIT1 | 13309 | -1.304 | -0.1858 | No | ||
| 111 | LIN28 | 13412 | -1.335 | -0.1885 | No | ||
| 112 | DAZL | 13883 | -1.474 | -0.2080 | No | ||
| 113 | TNFRSF21 | 13903 | -1.480 | -0.2066 | No | ||
| 114 | FGF11 | 13918 | -1.484 | -0.2051 | No | ||
| 115 | CGN | 13962 | -1.491 | -0.2048 | No | ||
| 116 | EIF4G1 | 14061 | -1.526 | -0.2071 | No | ||
| 117 | COL2A1 | 14127 | -1.543 | -0.2078 | No | ||
| 118 | NKX2-3 | 14220 | -1.573 | -0.2097 | No | ||
| 119 | DRD1 | 14808 | -1.745 | -0.2341 | No | ||
| 120 | RPL22 | 14852 | -1.756 | -0.2335 | No | ||
| 121 | MAX | 15281 | -1.888 | -0.2504 | No | ||
| 122 | MAT2A | 15303 | -1.894 | -0.2485 | No | ||
| 123 | FADS3 | 15315 | -1.899 | -0.2462 | No | ||
| 124 | LHX9 | 15468 | -1.946 | -0.2503 | No | ||
| 125 | SYT3 | 15512 | -1.964 | -0.2494 | No | ||
| 126 | GRIN2A | 15580 | -1.993 | -0.2495 | No | ||
| 127 | ETV1 | 15743 | -2.044 | -0.2539 | No | ||
| 128 | HSPE1 | 15969 | -2.115 | -0.2611 | No | ||
| 129 | PRKCE | 16103 | -2.152 | -0.2641 | No | ||
| 130 | ACVR2B | 16289 | -2.215 | -0.2693 | No | ||
| 131 | SLC31A2 | 16327 | -2.224 | -0.2677 | No | ||
| 132 | HPCA | 16348 | -2.233 | -0.2653 | No | ||
| 133 | KIAA0427 | 16400 | -2.248 | -0.2643 | No | ||
| 134 | PAX6 | 16454 | -2.272 | -0.2634 | No | ||
| 135 | RAB2 | 16500 | -2.289 | -0.2621 | No | ||
| 136 | PPP1R9B | 16637 | -2.334 | -0.2649 | No | ||
| 137 | WEE1 | 16986 | -2.462 | -0.2772 | No | ||
| 138 | FGF14 | 17196 | -2.532 | -0.2831 | No | ||
| 139 | LTBR | 17221 | -2.543 | -0.2804 | No | ||
| 140 | POLR2L | 17338 | -2.592 | -0.2819 | No | ||
| 141 | ENPP6 | 17402 | -2.613 | -0.2809 | No | ||
| 142 | USP2 | 17454 | -2.632 | -0.2794 | No | ||
| 143 | RUSC1 | 17618 | -2.689 | -0.2829 | No | ||
| 144 | RHEBL1 | 17626 | -2.692 | -0.2792 | No | ||
| 145 | SRP72 | 17662 | -2.710 | -0.2768 | No | ||
| 146 | HHIP | 17668 | -2.713 | -0.2730 | No | ||
| 147 | NXPH1 | 17906 | -2.815 | -0.2797 | No | ||
| 148 | SH3KBP1 | 18109 | -2.902 | -0.2847 | No | ||
| 149 | PTMA | 18360 | -3.006 | -0.2918 | Yes | ||
| 150 | MICAL2 | 18377 | -3.012 | -0.2880 | Yes | ||
| 151 | XRN2 | 18427 | -3.037 | -0.2858 | Yes | ||
| 152 | SLC9A5 | 18601 | -3.127 | -0.2891 | Yes | ||
| 153 | SYT6 | 18666 | -3.158 | -0.2874 | Yes | ||
| 154 | TDRD1 | 18681 | -3.166 | -0.2833 | Yes | ||
| 155 | EIF4B | 18819 | -3.244 | -0.2848 | Yes | ||
| 156 | EIF5A | 18853 | -3.265 | -0.2815 | Yes | ||
| 157 | OPRS1 | 18908 | -3.303 | -0.2791 | Yes | ||
| 158 | FARP1 | 18923 | -3.308 | -0.2748 | Yes | ||
| 159 | AMPD2 | 18964 | -3.332 | -0.2717 | Yes | ||
| 160 | AKAP12 | 18976 | -3.339 | -0.2673 | Yes | ||
| 161 | HPCAL4 | 19005 | -3.357 | -0.2636 | Yes | ||
| 162 | FGF6 | 19039 | -3.374 | -0.2601 | Yes | ||
| 163 | PPAT | 19093 | -3.411 | -0.2575 | Yes | ||
| 164 | SCRT2 | 19174 | -3.467 | -0.2560 | Yes | ||
| 165 | EXOSC5 | 19572 | -3.745 | -0.2687 | Yes | ||
| 166 | MORF4L2 | 19680 | -3.832 | -0.2680 | Yes | ||
| 167 | ZNRF2 | 19718 | -3.868 | -0.2639 | Yes | ||
| 168 | IVNS1ABP | 19995 | -4.130 | -0.2705 | Yes | ||
| 169 | SET | 20004 | -4.145 | -0.2647 | Yes | ||
| 170 | CNNM1 | 20163 | -4.303 | -0.2656 | Yes | ||
| 171 | B4GALT2 | 20173 | -4.315 | -0.2596 | Yes | ||
| 172 | ABCB6 | 20189 | -4.339 | -0.2539 | Yes | ||
| 173 | FBL | 20264 | -4.415 | -0.2507 | Yes | ||
| 174 | SEZ6L | 20304 | -4.450 | -0.2459 | Yes | ||
| 175 | TIAL1 | 20346 | -4.505 | -0.2411 | Yes | ||
| 176 | NUDC | 20381 | -4.549 | -0.2360 | Yes | ||
| 177 | PABPC1 | 20409 | -4.595 | -0.2304 | Yes | ||
| 178 | ENO3 | 20436 | -4.643 | -0.2247 | Yes | ||
| 179 | EEF1E1 | 20455 | -4.662 | -0.2186 | Yes | ||
| 180 | TIMM10 | 20474 | -4.692 | -0.2125 | Yes | ||
| 181 | G6PC3 | 20596 | -4.833 | -0.2109 | Yes | ||
| 182 | SCFD2 | 20660 | -4.930 | -0.2065 | Yes | ||
| 183 | PAICS | 20688 | -4.990 | -0.2003 | Yes | ||
| 184 | PSEN2 | 20782 | -5.170 | -0.1969 | Yes | ||
| 185 | AK3 | 20784 | -5.174 | -0.1893 | Yes | ||
| 186 | UBXD3 | 20939 | -5.375 | -0.1884 | Yes | ||
| 187 | HSPA4 | 21067 | -5.641 | -0.1859 | Yes | ||
| 188 | HOXA3 | 21069 | -5.643 | -0.1776 | Yes | ||
| 189 | MTHFD1 | 21089 | -5.684 | -0.1700 | Yes | ||
| 190 | IPO4 | 21131 | -5.788 | -0.1633 | Yes | ||
| 191 | POLR3E | 21203 | -5.991 | -0.1577 | Yes | ||
| 192 | EIF3S1 | 21214 | -6.022 | -0.1492 | Yes | ||
| 193 | GNA13 | 21224 | -6.051 | -0.1407 | Yes | ||
| 194 | NPM1 | 21234 | -6.089 | -0.1321 | Yes | ||
| 195 | SNX5 | 21360 | -6.467 | -0.1282 | Yes | ||
| 196 | SOCS5 | 21548 | -7.116 | -0.1263 | Yes | ||
| 197 | SUCLG2 | 21556 | -7.140 | -0.1160 | Yes | ||
| 198 | LYAR | 21651 | -7.674 | -0.1089 | Yes | ||
| 199 | HMOX1 | 21705 | -8.120 | -0.0993 | Yes | ||
| 200 | TIMM9 | 21713 | -8.175 | -0.0875 | Yes | ||
| 201 | PA2G4 | 21722 | -8.235 | -0.0757 | Yes | ||
| 202 | CACNA1D | 21745 | -8.426 | -0.0642 | Yes | ||
| 203 | EXOSC2 | 21754 | -8.613 | -0.0518 | Yes | ||
| 204 | NLN | 21828 | -9.662 | -0.0409 | Yes | ||
| 205 | LAP3 | 21929 | -14.156 | -0.0245 | Yes | ||
| 206 | SDC1 | 21941 | -17.025 | 0.0003 | Yes |
