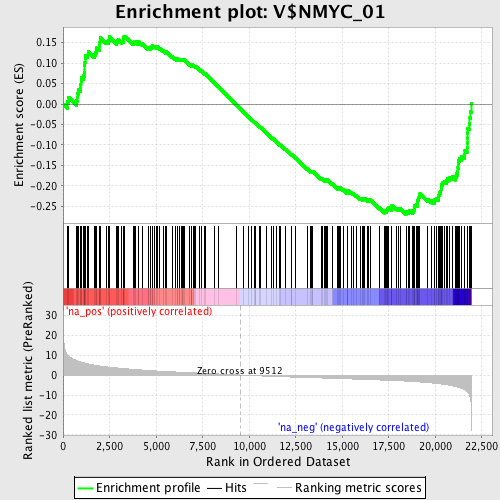
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_BT_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$NMYC_01 |
| Enrichment Score (ES) | -0.26973143 |
| Normalized Enrichment Score (NES) | -1.3223139 |
| Nominal p-value | 0.01904762 |
| FDR q-value | 0.38611683 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ARF6 | 219 | 10.345 | 0.0055 | No | ||
| 2 | RAD9A | 304 | 9.493 | 0.0159 | No | ||
| 3 | GADD45B | 712 | 7.204 | 0.0080 | No | ||
| 4 | SLC43A1 | 763 | 7.006 | 0.0162 | No | ||
| 5 | IQGAP2 | 787 | 6.930 | 0.0255 | No | ||
| 6 | SEMA7A | 817 | 6.836 | 0.0345 | No | ||
| 7 | FOXO3A | 929 | 6.515 | 0.0392 | No | ||
| 8 | CCDC6 | 941 | 6.492 | 0.0484 | No | ||
| 9 | MXD4 | 962 | 6.423 | 0.0571 | No | ||
| 10 | WBP2 | 992 | 6.356 | 0.0653 | No | ||
| 11 | WWP2 | 1105 | 6.072 | 0.0693 | No | ||
| 12 | USP34 | 1141 | 6.001 | 0.0767 | No | ||
| 13 | DNMT3A | 1150 | 5.973 | 0.0853 | No | ||
| 14 | BCL2 | 1151 | 5.973 | 0.0943 | No | ||
| 15 | STAT3 | 1171 | 5.916 | 0.1023 | No | ||
| 16 | PFN1 | 1183 | 5.900 | 0.1106 | No | ||
| 17 | UBE1 | 1195 | 5.864 | 0.1189 | No | ||
| 18 | PLAG1 | 1336 | 5.534 | 0.1208 | No | ||
| 19 | AKAP10 | 1357 | 5.486 | 0.1281 | No | ||
| 20 | TESK2 | 1692 | 4.905 | 0.1202 | No | ||
| 21 | TLE3 | 1727 | 4.860 | 0.1259 | No | ||
| 22 | TEF | 1791 | 4.765 | 0.1301 | No | ||
| 23 | KCNK5 | 1800 | 4.750 | 0.1369 | No | ||
| 24 | ANXA6 | 1937 | 4.556 | 0.1375 | No | ||
| 25 | MYST3 | 1939 | 4.555 | 0.1443 | No | ||
| 26 | AP1GBP1 | 1972 | 4.501 | 0.1496 | No | ||
| 27 | GRK6 | 1983 | 4.483 | 0.1559 | No | ||
| 28 | MFHAS1 | 1993 | 4.467 | 0.1622 | No | ||
| 29 | ZFP91 | 2331 | 4.083 | 0.1528 | No | ||
| 30 | NR5A1 | 2423 | 3.980 | 0.1546 | No | ||
| 31 | VPS16 | 2473 | 3.917 | 0.1582 | No | ||
| 32 | RUNX2 | 2474 | 3.916 | 0.1641 | No | ||
| 33 | SCYL1 | 2868 | 3.546 | 0.1513 | No | ||
| 34 | ADCY3 | 2916 | 3.504 | 0.1544 | No | ||
| 35 | HOXC13 | 2966 | 3.464 | 0.1574 | No | ||
| 36 | U2AF2 | 3156 | 3.302 | 0.1537 | No | ||
| 37 | MYCL1 | 3237 | 3.234 | 0.1548 | No | ||
| 38 | NRXN1 | 3253 | 3.223 | 0.1590 | No | ||
| 39 | SC5DL | 3258 | 3.220 | 0.1636 | No | ||
| 40 | GPD1 | 3317 | 3.167 | 0.1657 | No | ||
| 41 | FOXJ3 | 3768 | 2.823 | 0.1493 | No | ||
| 42 | PDK2 | 3808 | 2.796 | 0.1517 | No | ||
| 43 | ATP6V0B | 3887 | 2.734 | 0.1522 | No | ||
| 44 | RXRB | 4050 | 2.615 | 0.1487 | No | ||
| 45 | HBP1 | 4067 | 2.607 | 0.1519 | No | ||
| 46 | ZHX2 | 4253 | 2.468 | 0.1471 | No | ||
| 47 | NUP153 | 4569 | 2.279 | 0.1360 | No | ||
| 48 | MAPKAPK3 | 4580 | 2.275 | 0.1390 | No | ||
| 49 | RBBP6 | 4707 | 2.195 | 0.1365 | No | ||
| 50 | CHRM1 | 4720 | 2.189 | 0.1392 | No | ||
| 51 | PANK3 | 4778 | 2.155 | 0.1398 | No | ||
| 52 | TOP1 | 4795 | 2.149 | 0.1423 | No | ||
| 53 | BTBD5 | 4899 | 2.097 | 0.1407 | No | ||
| 54 | SNCAIP | 5007 | 2.028 | 0.1389 | No | ||
| 55 | SEPT3 | 5053 | 2.001 | 0.1398 | No | ||
| 56 | CUGBP1 | 5192 | 1.933 | 0.1363 | No | ||
| 57 | SLC20A1 | 5392 | 1.837 | 0.1300 | No | ||
| 58 | RBM3 | 5523 | 1.766 | 0.1266 | No | ||
| 59 | GPS1 | 5542 | 1.756 | 0.1284 | No | ||
| 60 | IPO13 | 5878 | 1.600 | 0.1154 | No | ||
| 61 | RPL13A | 6038 | 1.526 | 0.1104 | No | ||
| 62 | BMP2K | 6122 | 1.491 | 0.1088 | No | ||
| 63 | CTSF | 6132 | 1.486 | 0.1107 | No | ||
| 64 | IL1RAPL1 | 6231 | 1.436 | 0.1083 | No | ||
| 65 | HMGN2 | 6245 | 1.428 | 0.1099 | No | ||
| 66 | NRAS | 6347 | 1.384 | 0.1073 | No | ||
| 67 | POGK | 6397 | 1.362 | 0.1071 | No | ||
| 68 | KRTCAP2 | 6436 | 1.344 | 0.1074 | No | ||
| 69 | RAMP2 | 6466 | 1.328 | 0.1080 | No | ||
| 70 | MNT | 6512 | 1.303 | 0.1079 | No | ||
| 71 | ANGPT2 | 6805 | 1.172 | 0.0962 | No | ||
| 72 | DNAJB9 | 6892 | 1.125 | 0.0940 | No | ||
| 73 | JMJD1A | 6903 | 1.121 | 0.0952 | No | ||
| 74 | ALX4 | 6923 | 1.113 | 0.0960 | No | ||
| 75 | ABCA1 | 7002 | 1.080 | 0.0940 | No | ||
| 76 | HRH3 | 7084 | 1.045 | 0.0919 | No | ||
| 77 | SLC12A5 | 7123 | 1.028 | 0.0917 | No | ||
| 78 | PPARGC1B | 7326 | 0.939 | 0.0838 | No | ||
| 79 | SLC38A5 | 7412 | 0.902 | 0.0812 | No | ||
| 80 | CLCN2 | 7608 | 0.825 | 0.0735 | No | ||
| 81 | MCM8 | 7627 | 0.817 | 0.0739 | No | ||
| 82 | ZCCHC7 | 8139 | 0.602 | 0.0513 | No | ||
| 83 | HYAL2 | 8338 | 0.521 | 0.0430 | No | ||
| 84 | FHOD1 | 9336 | 0.092 | -0.0027 | No | ||
| 85 | RTN4RL2 | 9691 | -0.081 | -0.0189 | No | ||
| 86 | STMN4 | 9951 | -0.199 | -0.0305 | No | ||
| 87 | EN2 | 10148 | -0.278 | -0.0391 | No | ||
| 88 | NEUROD6 | 10287 | -0.332 | -0.0449 | No | ||
| 89 | PLA2G4A | 10313 | -0.343 | -0.0455 | No | ||
| 90 | HS3ST3A1 | 10335 | -0.354 | -0.0460 | No | ||
| 91 | GADD45G | 10557 | -0.424 | -0.0555 | No | ||
| 92 | NR1D1 | 10596 | -0.438 | -0.0566 | No | ||
| 93 | CDC14A | 10918 | -0.555 | -0.0705 | No | ||
| 94 | ARHGAP20 | 11170 | -0.645 | -0.0811 | No | ||
| 95 | ARRDC3 | 11277 | -0.676 | -0.0849 | No | ||
| 96 | RPIA | 11453 | -0.733 | -0.0919 | No | ||
| 97 | SGTB | 11640 | -0.796 | -0.0993 | No | ||
| 98 | DLC1 | 11686 | -0.820 | -0.1001 | No | ||
| 99 | B3GALT2 | 11933 | -0.897 | -0.1101 | No | ||
| 100 | VGF | 12255 | -0.993 | -0.1233 | No | ||
| 101 | BAI2 | 12283 | -1.002 | -0.1231 | No | ||
| 102 | CHST11 | 12496 | -1.067 | -0.1312 | No | ||
| 103 | FLT3 | 13124 | -1.253 | -0.1581 | No | ||
| 104 | GIT1 | 13309 | -1.304 | -0.1646 | No | ||
| 105 | ADAM10 | 13366 | -1.321 | -0.1652 | No | ||
| 106 | LIN28 | 13412 | -1.335 | -0.1653 | No | ||
| 107 | NEF3 | 13424 | -1.340 | -0.1638 | No | ||
| 108 | GTPBP1 | 13889 | -1.476 | -0.1829 | No | ||
| 109 | FGF11 | 13918 | -1.484 | -0.1820 | No | ||
| 110 | EIF4G1 | 14061 | -1.526 | -0.1862 | No | ||
| 111 | PLAGL1 | 14112 | -1.539 | -0.1862 | No | ||
| 112 | COL2A1 | 14127 | -1.543 | -0.1845 | No | ||
| 113 | ZZZ3 | 14196 | -1.566 | -0.1853 | No | ||
| 114 | NEUROD2 | 14457 | -1.636 | -0.1948 | No | ||
| 115 | BARHL1 | 14757 | -1.729 | -0.2060 | No | ||
| 116 | DUSP4 | 14782 | -1.737 | -0.2044 | No | ||
| 117 | RPL22 | 14852 | -1.756 | -0.2050 | No | ||
| 118 | NKX2-2 | 14903 | -1.772 | -0.2046 | No | ||
| 119 | PCDHA10 | 15046 | -1.813 | -0.2084 | No | ||
| 120 | GJA1 | 15279 | -1.888 | -0.2163 | No | ||
| 121 | MAX | 15281 | -1.888 | -0.2135 | No | ||
| 122 | POU3F1 | 15297 | -1.893 | -0.2113 | No | ||
| 123 | LHX9 | 15468 | -1.946 | -0.2162 | No | ||
| 124 | GRIN2A | 15580 | -1.993 | -0.2183 | No | ||
| 125 | ETV1 | 15743 | -2.044 | -0.2227 | No | ||
| 126 | HSPE1 | 15969 | -2.115 | -0.2299 | No | ||
| 127 | PRKCE | 16103 | -2.152 | -0.2328 | No | ||
| 128 | BCL7C | 16142 | -2.164 | -0.2313 | No | ||
| 129 | HRSP12 | 16189 | -2.176 | -0.2301 | No | ||
| 130 | HPCA | 16348 | -2.233 | -0.2340 | No | ||
| 131 | KIAA0427 | 16400 | -2.248 | -0.2330 | No | ||
| 132 | RAB2 | 16500 | -2.289 | -0.2341 | No | ||
| 133 | WEE1 | 16986 | -2.462 | -0.2527 | No | ||
| 134 | APOA5 | 17281 | -2.566 | -0.2624 | No | ||
| 135 | RAB30 | 17332 | -2.587 | -0.2608 | No | ||
| 136 | ENPP6 | 17402 | -2.613 | -0.2600 | No | ||
| 137 | ADPRHL1 | 17414 | -2.618 | -0.2566 | No | ||
| 138 | USP2 | 17454 | -2.632 | -0.2544 | No | ||
| 139 | GPC3 | 17497 | -2.647 | -0.2524 | No | ||
| 140 | CTBP2 | 17648 | -2.704 | -0.2552 | No | ||
| 141 | SRP72 | 17662 | -2.710 | -0.2518 | No | ||
| 142 | HHIP | 17668 | -2.713 | -0.2479 | No | ||
| 143 | NXPH1 | 17906 | -2.815 | -0.2546 | No | ||
| 144 | ADSS | 18029 | -2.867 | -0.2559 | No | ||
| 145 | SH3KBP1 | 18109 | -2.902 | -0.2552 | No | ||
| 146 | XRN2 | 18427 | -3.037 | -0.2652 | Yes | ||
| 147 | NDUFS1 | 18468 | -3.057 | -0.2624 | Yes | ||
| 148 | SEMA3F | 18584 | -3.113 | -0.2630 | Yes | ||
| 149 | SLC9A5 | 18601 | -3.127 | -0.2591 | Yes | ||
| 150 | SOX12 | 18768 | -3.222 | -0.2619 | Yes | ||
| 151 | EIF4B | 18819 | -3.244 | -0.2593 | Yes | ||
| 152 | JPH1 | 18858 | -3.267 | -0.2561 | Yes | ||
| 153 | HTF9C | 18872 | -3.275 | -0.2518 | Yes | ||
| 154 | RNF128 | 18899 | -3.297 | -0.2481 | Yes | ||
| 155 | AMPD2 | 18964 | -3.332 | -0.2460 | Yes | ||
| 156 | SLC17A2 | 19036 | -3.373 | -0.2442 | Yes | ||
| 157 | FGF6 | 19039 | -3.374 | -0.2392 | Yes | ||
| 158 | FKBP10 | 19051 | -3.380 | -0.2346 | Yes | ||
| 159 | PPAT | 19093 | -3.411 | -0.2314 | Yes | ||
| 160 | LMNB1 | 19119 | -3.431 | -0.2274 | Yes | ||
| 161 | PFKFB3 | 19165 | -3.460 | -0.2243 | Yes | ||
| 162 | SCRT2 | 19174 | -3.467 | -0.2194 | Yes | ||
| 163 | FKBP5 | 19597 | -3.763 | -0.2332 | Yes | ||
| 164 | DLX1 | 19783 | -3.932 | -0.2358 | Yes | ||
| 165 | RCOR2 | 19961 | -4.086 | -0.2378 | Yes | ||
| 166 | SFXN2 | 19978 | -4.105 | -0.2324 | Yes | ||
| 167 | COPS7A | 20058 | -4.200 | -0.2297 | Yes | ||
| 168 | B4GALT2 | 20173 | -4.315 | -0.2285 | Yes | ||
| 169 | TPM2 | 20185 | -4.335 | -0.2224 | Yes | ||
| 170 | LIG3 | 20211 | -4.370 | -0.2170 | Yes | ||
| 171 | FBL | 20264 | -4.415 | -0.2128 | Yes | ||
| 172 | SEZ6L | 20304 | -4.450 | -0.2079 | Yes | ||
| 173 | ELOVL6 | 20321 | -4.475 | -0.2019 | Yes | ||
| 174 | TIAL1 | 20346 | -4.505 | -0.1963 | Yes | ||
| 175 | PABPC1 | 20409 | -4.595 | -0.1922 | Yes | ||
| 176 | TIMM10 | 20474 | -4.692 | -0.1881 | Yes | ||
| 177 | G6PC3 | 20596 | -4.833 | -0.1864 | Yes | ||
| 178 | SCFD2 | 20660 | -4.930 | -0.1819 | Yes | ||
| 179 | PSEN2 | 20782 | -5.170 | -0.1797 | Yes | ||
| 180 | HSPD1 | 20908 | -5.331 | -0.1774 | Yes | ||
| 181 | MTHFD1 | 21089 | -5.684 | -0.1772 | Yes | ||
| 182 | IPO4 | 21131 | -5.788 | -0.1704 | Yes | ||
| 183 | POLR3E | 21203 | -5.991 | -0.1646 | Yes | ||
| 184 | EIF3S1 | 21214 | -6.022 | -0.1561 | Yes | ||
| 185 | GNA13 | 21224 | -6.051 | -0.1474 | Yes | ||
| 186 | NPM1 | 21234 | -6.089 | -0.1387 | Yes | ||
| 187 | RCL1 | 21292 | -6.273 | -0.1319 | Yes | ||
| 188 | ATP5G1 | 21410 | -6.613 | -0.1273 | Yes | ||
| 189 | SOCS5 | 21548 | -7.116 | -0.1229 | Yes | ||
| 190 | PES1 | 21591 | -7.294 | -0.1139 | Yes | ||
| 191 | HMOX1 | 21705 | -8.120 | -0.1069 | Yes | ||
| 192 | TIMM9 | 21713 | -8.175 | -0.0949 | Yes | ||
| 193 | PA2G4 | 21722 | -8.235 | -0.0829 | Yes | ||
| 194 | PRDX4 | 21727 | -8.270 | -0.0707 | Yes | ||
| 195 | CACNA1D | 21745 | -8.426 | -0.0588 | Yes | ||
| 196 | ATAD3A | 21821 | -9.429 | -0.0481 | Yes | ||
| 197 | NOL5A | 21859 | -10.573 | -0.0339 | Yes | ||
| 198 | CEBPB | 21876 | -11.074 | -0.0180 | Yes | ||
| 199 | LAP3 | 21929 | -14.156 | 0.0008 | Yes |
