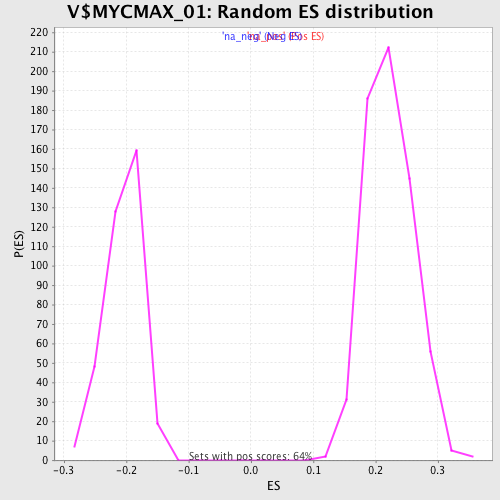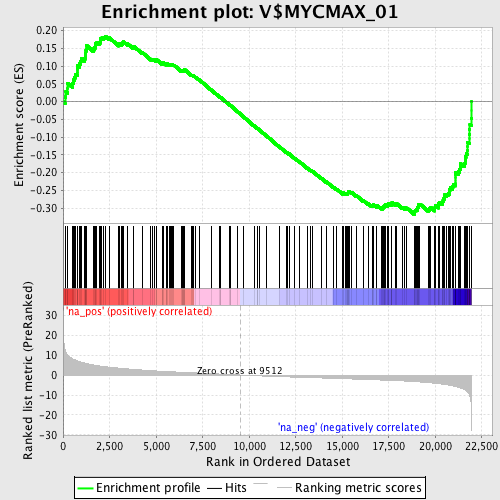
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_BT_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$MYCMAX_01 |
| Enrichment Score (ES) | -0.318241 |
| Normalized Enrichment Score (NES) | -1.567318 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.14239371 |
| FWER p-Value | 0.613 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RORC | 108 | 12.686 | 0.0131 | No | ||
| 2 | DIRC2 | 148 | 11.521 | 0.0277 | No | ||
| 3 | HSPBAP1 | 245 | 10.027 | 0.0376 | No | ||
| 4 | ISGF3G | 251 | 9.973 | 0.0515 | No | ||
| 5 | TUBA1 | 511 | 8.088 | 0.0511 | No | ||
| 6 | ATF7IP | 544 | 7.955 | 0.0610 | No | ||
| 7 | H3F3A | 611 | 7.649 | 0.0688 | No | ||
| 8 | WDR23 | 670 | 7.418 | 0.0767 | No | ||
| 9 | SLC43A1 | 763 | 7.006 | 0.0825 | No | ||
| 10 | CRMP1 | 768 | 6.999 | 0.0922 | No | ||
| 11 | IQGAP2 | 787 | 6.930 | 0.1013 | No | ||
| 12 | SFRS14 | 884 | 6.627 | 0.1063 | No | ||
| 13 | FOXO3A | 929 | 6.515 | 0.1136 | No | ||
| 14 | MXD4 | 962 | 6.423 | 0.1212 | No | ||
| 15 | USP34 | 1141 | 6.001 | 0.1216 | No | ||
| 16 | PFN1 | 1183 | 5.900 | 0.1281 | No | ||
| 17 | UBE1 | 1195 | 5.864 | 0.1359 | No | ||
| 18 | CSK | 1223 | 5.821 | 0.1430 | No | ||
| 19 | CBX5 | 1242 | 5.764 | 0.1504 | No | ||
| 20 | HIRA | 1246 | 5.757 | 0.1584 | No | ||
| 21 | KCNN4 | 1620 | 5.016 | 0.1484 | No | ||
| 22 | TESK2 | 1692 | 4.905 | 0.1521 | No | ||
| 23 | ILF3 | 1726 | 4.861 | 0.1575 | No | ||
| 24 | USP15 | 1732 | 4.851 | 0.1642 | No | ||
| 25 | CHD4 | 1814 | 4.734 | 0.1672 | No | ||
| 26 | UBE2B | 1974 | 4.499 | 0.1663 | No | ||
| 27 | PER1 | 2010 | 4.448 | 0.1710 | No | ||
| 28 | FKBP11 | 2017 | 4.437 | 0.1771 | No | ||
| 29 | EIF4E | 2088 | 4.365 | 0.1801 | No | ||
| 30 | BCAS3 | 2191 | 4.231 | 0.1814 | No | ||
| 31 | LZTS2 | 2258 | 4.161 | 0.1843 | No | ||
| 32 | RUNX2 | 2474 | 3.916 | 0.1800 | No | ||
| 33 | TUBA4 | 2990 | 3.445 | 0.1612 | No | ||
| 34 | ESRRA | 3039 | 3.404 | 0.1639 | No | ||
| 35 | U2AF2 | 3156 | 3.302 | 0.1632 | No | ||
| 36 | DNAJB5 | 3210 | 3.252 | 0.1654 | No | ||
| 37 | MYCL1 | 3237 | 3.234 | 0.1688 | No | ||
| 38 | STXBP2 | 3467 | 3.043 | 0.1627 | No | ||
| 39 | FOXJ3 | 3768 | 2.823 | 0.1529 | No | ||
| 40 | ASXL1 | 3780 | 2.814 | 0.1564 | No | ||
| 41 | ZHX2 | 4253 | 2.468 | 0.1382 | No | ||
| 42 | CHRM1 | 4720 | 2.189 | 0.1199 | No | ||
| 43 | TOP1 | 4795 | 2.149 | 0.1196 | No | ||
| 44 | BTBD5 | 4899 | 2.097 | 0.1178 | No | ||
| 45 | ARL3 | 4991 | 2.043 | 0.1166 | No | ||
| 46 | SNCAIP | 5007 | 2.028 | 0.1187 | No | ||
| 47 | DIABLO | 5333 | 1.861 | 0.1065 | No | ||
| 48 | TRIM46 | 5361 | 1.850 | 0.1079 | No | ||
| 49 | KCNQ5 | 5382 | 1.841 | 0.1096 | No | ||
| 50 | GPS1 | 5542 | 1.756 | 0.1048 | No | ||
| 51 | ADAMTS19 | 5563 | 1.743 | 0.1063 | No | ||
| 52 | RPS6KA5 | 5605 | 1.724 | 0.1069 | No | ||
| 53 | SLC1A7 | 5740 | 1.668 | 0.1031 | No | ||
| 54 | ETV4 | 5767 | 1.656 | 0.1043 | No | ||
| 55 | RFX5 | 5807 | 1.633 | 0.1048 | No | ||
| 56 | IPO13 | 5878 | 1.600 | 0.1039 | No | ||
| 57 | LRP8 | 5950 | 1.569 | 0.1028 | No | ||
| 58 | NRAS | 6347 | 1.384 | 0.0866 | No | ||
| 59 | POGK | 6397 | 1.362 | 0.0863 | No | ||
| 60 | KRTCAP2 | 6436 | 1.344 | 0.0864 | No | ||
| 61 | RAMP2 | 6466 | 1.328 | 0.0870 | No | ||
| 62 | BMP4 | 6472 | 1.325 | 0.0887 | No | ||
| 63 | MPP3 | 6502 | 1.308 | 0.0892 | No | ||
| 64 | MNT | 6512 | 1.303 | 0.0906 | No | ||
| 65 | JMJD1A | 6903 | 1.121 | 0.0743 | No | ||
| 66 | NTN2L | 6931 | 1.109 | 0.0746 | No | ||
| 67 | ABCA1 | 7002 | 1.080 | 0.0730 | No | ||
| 68 | SLC12A5 | 7123 | 1.028 | 0.0689 | No | ||
| 69 | PPARGC1B | 7326 | 0.939 | 0.0610 | No | ||
| 70 | RPS19 | 7962 | 0.677 | 0.0328 | No | ||
| 71 | CLN3 | 8386 | 0.503 | 0.0140 | No | ||
| 72 | MEOX2 | 8437 | 0.480 | 0.0124 | No | ||
| 73 | PITX3 | 8452 | 0.474 | 0.0125 | No | ||
| 74 | ATOH8 | 8916 | 0.275 | -0.0084 | No | ||
| 75 | STMN1 | 8975 | 0.246 | -0.0107 | No | ||
| 76 | DAZAP1 | 8991 | 0.236 | -0.0111 | No | ||
| 77 | PPRC1 | 9386 | 0.064 | -0.0291 | No | ||
| 78 | RTN4RL2 | 9691 | -0.081 | -0.0430 | No | ||
| 79 | HOXA11 | 10259 | -0.319 | -0.0686 | No | ||
| 80 | EIF2C2 | 10446 | -0.387 | -0.0766 | No | ||
| 81 | EEF1B2 | 10449 | -0.388 | -0.0761 | No | ||
| 82 | GADD45G | 10557 | -0.424 | -0.0804 | No | ||
| 83 | CDC14A | 10918 | -0.555 | -0.0962 | No | ||
| 84 | EFNB1 | 11615 | -0.789 | -0.1270 | No | ||
| 85 | GPRC5C | 12001 | -0.922 | -0.1434 | No | ||
| 86 | SLC6A15 | 12049 | -0.935 | -0.1442 | No | ||
| 87 | SATB2 | 12145 | -0.960 | -0.1472 | No | ||
| 88 | FOXD3 | 12457 | -1.052 | -0.1600 | No | ||
| 89 | ZBTB10 | 12688 | -1.120 | -0.1690 | No | ||
| 90 | FLT3 | 13124 | -1.253 | -0.1872 | No | ||
| 91 | GIT1 | 13309 | -1.304 | -0.1938 | No | ||
| 92 | LIN28 | 13412 | -1.335 | -0.1966 | No | ||
| 93 | DAZL | 13883 | -1.474 | -0.2161 | No | ||
| 94 | COL2A1 | 14127 | -1.543 | -0.2250 | No | ||
| 95 | ELAVL3 | 14529 | -1.656 | -0.2411 | No | ||
| 96 | KCTD15 | 14702 | -1.713 | -0.2466 | No | ||
| 97 | QTRTD1 | 15030 | -1.808 | -0.2590 | No | ||
| 98 | PCDHA10 | 15046 | -1.813 | -0.2571 | No | ||
| 99 | CEBPA | 15154 | -1.854 | -0.2594 | No | ||
| 100 | SMAD2 | 15223 | -1.872 | -0.2599 | No | ||
| 101 | GJA1 | 15279 | -1.888 | -0.2597 | No | ||
| 102 | MAT2A | 15303 | -1.894 | -0.2581 | No | ||
| 103 | ACY1 | 15309 | -1.896 | -0.2556 | No | ||
| 104 | EWSR1 | 15336 | -1.905 | -0.2541 | No | ||
| 105 | HOXB5 | 15387 | -1.922 | -0.2536 | No | ||
| 106 | SYT3 | 15512 | -1.964 | -0.2566 | No | ||
| 107 | ETV1 | 15743 | -2.044 | -0.2642 | No | ||
| 108 | BCL7C | 16142 | -2.164 | -0.2794 | No | ||
| 109 | KIAA0427 | 16400 | -2.248 | -0.2880 | No | ||
| 110 | NRIP3 | 16597 | -2.322 | -0.2937 | No | ||
| 111 | PPP1R9B | 16637 | -2.334 | -0.2922 | No | ||
| 112 | PTGES2 | 16681 | -2.350 | -0.2908 | No | ||
| 113 | NR6A1 | 16861 | -2.417 | -0.2956 | No | ||
| 114 | POU3F2 | 16862 | -2.419 | -0.2922 | No | ||
| 115 | TRIB1 | 17100 | -2.499 | -0.2995 | No | ||
| 116 | TGFB2 | 17183 | -2.526 | -0.2997 | No | ||
| 117 | FGF14 | 17196 | -2.532 | -0.2966 | No | ||
| 118 | KCMF1 | 17249 | -2.553 | -0.2954 | No | ||
| 119 | APOA5 | 17281 | -2.566 | -0.2932 | No | ||
| 120 | IPO7 | 17313 | -2.578 | -0.2909 | No | ||
| 121 | NEUROD1 | 17432 | -2.624 | -0.2926 | No | ||
| 122 | BOK | 17462 | -2.635 | -0.2902 | No | ||
| 123 | GPC3 | 17497 | -2.647 | -0.2880 | No | ||
| 124 | RHEBL1 | 17626 | -2.692 | -0.2900 | No | ||
| 125 | SRP72 | 17662 | -2.710 | -0.2878 | No | ||
| 126 | HHIP | 17668 | -2.713 | -0.2841 | No | ||
| 127 | ZADH2 | 17870 | -2.796 | -0.2894 | No | ||
| 128 | GPM6B | 17903 | -2.815 | -0.2869 | No | ||
| 129 | HMGA1 | 18236 | -2.950 | -0.2979 | No | ||
| 130 | PTMA | 18360 | -3.006 | -0.2993 | No | ||
| 131 | NDUFS1 | 18468 | -3.057 | -0.2999 | No | ||
| 132 | SYNCRIP | 18869 | -3.273 | -0.3136 | Yes | ||
| 133 | HTF9C | 18872 | -3.275 | -0.3090 | Yes | ||
| 134 | OPRS1 | 18908 | -3.303 | -0.3059 | Yes | ||
| 135 | AMPD2 | 18964 | -3.332 | -0.3037 | Yes | ||
| 136 | FGF6 | 19039 | -3.374 | -0.3023 | Yes | ||
| 137 | FKBP10 | 19051 | -3.380 | -0.2980 | Yes | ||
| 138 | RFX4 | 19091 | -3.410 | -0.2949 | Yes | ||
| 139 | PPAT | 19093 | -3.411 | -0.2901 | Yes | ||
| 140 | SCRT2 | 19174 | -3.467 | -0.2889 | Yes | ||
| 141 | RAI14 | 19616 | -3.781 | -0.3038 | Yes | ||
| 142 | MRPL40 | 19681 | -3.832 | -0.3012 | Yes | ||
| 143 | ARMET | 19739 | -3.891 | -0.2983 | Yes | ||
| 144 | RCOR2 | 19961 | -4.086 | -0.3027 | Yes | ||
| 145 | SFXN2 | 19978 | -4.105 | -0.2976 | Yes | ||
| 146 | IVNS1ABP | 19995 | -4.130 | -0.2924 | Yes | ||
| 147 | CNNM1 | 20163 | -4.303 | -0.2940 | Yes | ||
| 148 | CGREF1 | 20168 | -4.310 | -0.2880 | Yes | ||
| 149 | RAB3IL1 | 20225 | -4.385 | -0.2843 | Yes | ||
| 150 | NUDC | 20381 | -4.549 | -0.2850 | Yes | ||
| 151 | PABPC1 | 20409 | -4.595 | -0.2797 | Yes | ||
| 152 | ENO3 | 20436 | -4.643 | -0.2743 | Yes | ||
| 153 | TIMM10 | 20474 | -4.692 | -0.2693 | Yes | ||
| 154 | OPRD1 | 20484 | -4.703 | -0.2630 | Yes | ||
| 155 | G6PC3 | 20596 | -4.833 | -0.2612 | Yes | ||
| 156 | PAICS | 20688 | -4.990 | -0.2583 | Yes | ||
| 157 | IFRD2 | 20755 | -5.117 | -0.2541 | Yes | ||
| 158 | HNRPA1 | 20763 | -5.130 | -0.2471 | Yes | ||
| 159 | HNRPDL | 20813 | -5.210 | -0.2419 | Yes | ||
| 160 | UBXD3 | 20939 | -5.375 | -0.2400 | Yes | ||
| 161 | TFRC | 20958 | -5.404 | -0.2332 | Yes | ||
| 162 | TCERG1 | 21064 | -5.636 | -0.2300 | Yes | ||
| 163 | RANBP1 | 21068 | -5.642 | -0.2221 | Yes | ||
| 164 | HOXA3 | 21069 | -5.643 | -0.2140 | Yes | ||
| 165 | PUS1 | 21072 | -5.646 | -0.2061 | Yes | ||
| 166 | MTHFD1 | 21089 | -5.684 | -0.1987 | Yes | ||
| 167 | NPM1 | 21234 | -6.089 | -0.1967 | Yes | ||
| 168 | RCL1 | 21292 | -6.273 | -0.1904 | Yes | ||
| 169 | PRPS1 | 21340 | -6.411 | -0.1834 | Yes | ||
| 170 | SNX5 | 21360 | -6.467 | -0.1751 | Yes | ||
| 171 | SUCLG2 | 21556 | -7.140 | -0.1739 | Yes | ||
| 172 | NCL | 21602 | -7.341 | -0.1655 | Yes | ||
| 173 | QTRT1 | 21616 | -7.430 | -0.1555 | Yes | ||
| 174 | LYAR | 21651 | -7.674 | -0.1462 | Yes | ||
| 175 | TIMM9 | 21713 | -8.175 | -0.1373 | Yes | ||
| 176 | PA2G4 | 21722 | -8.235 | -0.1260 | Yes | ||
| 177 | PRDX4 | 21727 | -8.270 | -0.1144 | Yes | ||
| 178 | ATAD3A | 21821 | -9.429 | -0.1053 | Yes | ||
| 179 | PSME3 | 21823 | -9.439 | -0.0919 | Yes | ||
| 180 | TCOF1 | 21838 | -9.974 | -0.0783 | Yes | ||
| 181 | NOL5A | 21859 | -10.573 | -0.0642 | Yes | ||
| 182 | LAP3 | 21929 | -14.156 | -0.0472 | Yes | ||
| 183 | CAD | 21940 | -16.671 | -0.0240 | Yes | ||
| 184 | SDC1 | 21941 | -17.025 | 0.0003 | Yes |