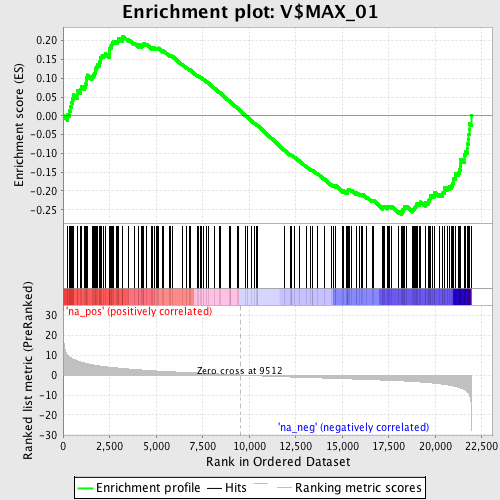
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_BT_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$MAX_01 |
| Enrichment Score (ES) | -0.26263213 |
| Normalized Enrichment Score (NES) | -1.2968086 |
| Nominal p-value | 0.03125 |
| FDR q-value | 0.37314093 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ISGF3G | 251 | 9.973 | 0.0039 | No | ||
| 2 | DUSP1 | 351 | 9.094 | 0.0134 | No | ||
| 3 | DVL2 | 380 | 8.902 | 0.0259 | No | ||
| 4 | MTCH2 | 455 | 8.423 | 0.0355 | No | ||
| 5 | TUBA1 | 511 | 8.088 | 0.0455 | No | ||
| 6 | ATF7IP | 544 | 7.955 | 0.0563 | No | ||
| 7 | GNAS | 780 | 6.945 | 0.0562 | No | ||
| 8 | IQGAP2 | 787 | 6.930 | 0.0667 | No | ||
| 9 | FOXO3A | 929 | 6.515 | 0.0703 | No | ||
| 10 | MXD4 | 962 | 6.423 | 0.0787 | No | ||
| 11 | DNMT3A | 1150 | 5.973 | 0.0794 | No | ||
| 12 | CSK | 1223 | 5.821 | 0.0851 | No | ||
| 13 | CBX5 | 1242 | 5.764 | 0.0932 | No | ||
| 14 | HIRA | 1246 | 5.757 | 0.1019 | No | ||
| 15 | ZFYVE26 | 1288 | 5.638 | 0.1088 | No | ||
| 16 | ANKHD1 | 1553 | 5.123 | 0.1045 | No | ||
| 17 | LAMP1 | 1611 | 5.028 | 0.1097 | No | ||
| 18 | TESK2 | 1692 | 4.905 | 0.1136 | No | ||
| 19 | ILF3 | 1726 | 4.861 | 0.1196 | No | ||
| 20 | USP15 | 1732 | 4.851 | 0.1269 | No | ||
| 21 | CHD4 | 1814 | 4.734 | 0.1305 | No | ||
| 22 | RLF | 1852 | 4.680 | 0.1360 | No | ||
| 23 | UBE4B | 1934 | 4.560 | 0.1393 | No | ||
| 24 | UBE2B | 1974 | 4.499 | 0.1445 | No | ||
| 25 | PER1 | 2010 | 4.448 | 0.1498 | No | ||
| 26 | FKBP11 | 2017 | 4.437 | 0.1564 | No | ||
| 27 | EIF4E | 2088 | 4.365 | 0.1599 | No | ||
| 28 | BCAS3 | 2191 | 4.231 | 0.1617 | No | ||
| 29 | LZTS2 | 2258 | 4.161 | 0.1651 | No | ||
| 30 | VPS16 | 2473 | 3.917 | 0.1614 | No | ||
| 31 | RUNX2 | 2474 | 3.916 | 0.1674 | No | ||
| 32 | PICALM | 2482 | 3.908 | 0.1731 | No | ||
| 33 | ZNF318 | 2497 | 3.894 | 0.1785 | No | ||
| 34 | SLC38A2 | 2520 | 3.874 | 0.1835 | No | ||
| 35 | NIT1 | 2599 | 3.808 | 0.1858 | No | ||
| 36 | ANKRD12 | 2622 | 3.790 | 0.1907 | No | ||
| 37 | CDK5R1 | 2644 | 3.758 | 0.1955 | No | ||
| 38 | EPC1 | 2723 | 3.682 | 0.1976 | No | ||
| 39 | SIRT1 | 2856 | 3.557 | 0.1970 | No | ||
| 40 | BCOR | 2930 | 3.491 | 0.1991 | No | ||
| 41 | AP1S2 | 2987 | 3.447 | 0.2018 | No | ||
| 42 | TUBA4 | 2990 | 3.445 | 0.2071 | No | ||
| 43 | PPM1A | 3168 | 3.292 | 0.2040 | No | ||
| 44 | SLC26A2 | 3203 | 3.257 | 0.2075 | No | ||
| 45 | DNAJB5 | 3210 | 3.252 | 0.2123 | No | ||
| 46 | TAF6L | 3514 | 3.010 | 0.2030 | No | ||
| 47 | PDK2 | 3808 | 2.796 | 0.1938 | No | ||
| 48 | HBP1 | 4067 | 2.607 | 0.1860 | No | ||
| 49 | GNB2 | 4074 | 2.605 | 0.1898 | No | ||
| 50 | CPEB4 | 4224 | 2.491 | 0.1868 | No | ||
| 51 | ATP6V1A | 4266 | 2.462 | 0.1887 | No | ||
| 52 | FEN1 | 4325 | 2.423 | 0.1898 | No | ||
| 53 | PPP1R3C | 4340 | 2.415 | 0.1929 | No | ||
| 54 | KBTBD2 | 4487 | 2.329 | 0.1898 | No | ||
| 55 | ALDH6A1 | 4760 | 2.164 | 0.1806 | No | ||
| 56 | TOP1 | 4795 | 2.149 | 0.1824 | No | ||
| 57 | BTBD5 | 4899 | 2.097 | 0.1809 | No | ||
| 58 | SNCAIP | 5007 | 2.028 | 0.1791 | No | ||
| 59 | SEPT3 | 5053 | 2.001 | 0.1801 | No | ||
| 60 | LZTFL1 | 5109 | 1.971 | 0.1806 | No | ||
| 61 | BRDT | 5345 | 1.856 | 0.1727 | No | ||
| 62 | PFDN2 | 5407 | 1.830 | 0.1727 | No | ||
| 63 | SLC1A7 | 5740 | 1.668 | 0.1601 | No | ||
| 64 | HPS5 | 5756 | 1.659 | 0.1619 | No | ||
| 65 | IPO13 | 5878 | 1.600 | 0.1589 | No | ||
| 66 | POGK | 6397 | 1.362 | 0.1372 | No | ||
| 67 | HOXD10 | 6654 | 1.231 | 0.1273 | No | ||
| 68 | COPZ1 | 6779 | 1.182 | 0.1234 | No | ||
| 69 | GATA5 | 6824 | 1.163 | 0.1232 | No | ||
| 70 | GPX1 | 7219 | 0.991 | 0.1066 | No | ||
| 71 | PRKCH | 7283 | 0.959 | 0.1052 | No | ||
| 72 | HHEX | 7379 | 0.918 | 0.1023 | No | ||
| 73 | SLC38A5 | 7412 | 0.902 | 0.1022 | No | ||
| 74 | TOPORS | 7552 | 0.848 | 0.0971 | No | ||
| 75 | CENTB5 | 7722 | 0.772 | 0.0906 | No | ||
| 76 | GUCY1A2 | 7821 | 0.731 | 0.0872 | No | ||
| 77 | ZCCHC7 | 8139 | 0.602 | 0.0735 | No | ||
| 78 | CLN3 | 8386 | 0.503 | 0.0630 | No | ||
| 79 | MEOX2 | 8437 | 0.480 | 0.0615 | No | ||
| 80 | ATOH8 | 8916 | 0.275 | 0.0399 | No | ||
| 81 | STMN1 | 8975 | 0.246 | 0.0376 | No | ||
| 82 | GTF2H1 | 8990 | 0.237 | 0.0374 | No | ||
| 83 | PPRC1 | 9386 | 0.064 | 0.0193 | No | ||
| 84 | RPS2 | 9398 | 0.061 | 0.0189 | No | ||
| 85 | PRDM4 | 9789 | -0.122 | 0.0012 | No | ||
| 86 | ASNA1 | 9903 | -0.176 | -0.0038 | No | ||
| 87 | EME1 | 10133 | -0.272 | -0.0139 | No | ||
| 88 | HOXA11 | 10259 | -0.319 | -0.0191 | No | ||
| 89 | KCNE4 | 10405 | -0.375 | -0.0252 | No | ||
| 90 | EIF2C2 | 10446 | -0.387 | -0.0264 | No | ||
| 91 | EEF1B2 | 10449 | -0.388 | -0.0259 | No | ||
| 92 | IRS4 | 11904 | -0.890 | -0.0914 | No | ||
| 93 | SSR1 | 12209 | -0.978 | -0.1038 | No | ||
| 94 | VGF | 12255 | -0.993 | -0.1044 | No | ||
| 95 | NPTX1 | 12293 | -1.004 | -0.1045 | No | ||
| 96 | FOXD3 | 12457 | -1.052 | -0.1104 | No | ||
| 97 | ZBTB10 | 12688 | -1.120 | -0.1192 | No | ||
| 98 | SUMF1 | 13069 | -1.237 | -0.1348 | No | ||
| 99 | GIT1 | 13309 | -1.304 | -0.1437 | No | ||
| 100 | BMP6 | 13373 | -1.323 | -0.1446 | No | ||
| 101 | LIN28 | 13412 | -1.335 | -0.1443 | No | ||
| 102 | EN1 | 13659 | -1.411 | -0.1534 | No | ||
| 103 | NMNAT2 | 14027 | -1.516 | -0.1679 | No | ||
| 104 | DDX3X | 14411 | -1.623 | -0.1830 | No | ||
| 105 | ELAVL3 | 14529 | -1.656 | -0.1858 | No | ||
| 106 | HOXC5 | 14622 | -1.689 | -0.1874 | No | ||
| 107 | GGN | 14639 | -1.693 | -0.1855 | No | ||
| 108 | QTRTD1 | 15030 | -1.808 | -0.2007 | No | ||
| 109 | PCDHA10 | 15046 | -1.813 | -0.1985 | No | ||
| 110 | SMAD2 | 15223 | -1.872 | -0.2037 | No | ||
| 111 | GJA1 | 15279 | -1.888 | -0.2033 | No | ||
| 112 | MAX | 15281 | -1.888 | -0.2005 | No | ||
| 113 | MAT2A | 15303 | -1.894 | -0.1985 | No | ||
| 114 | ACY1 | 15309 | -1.896 | -0.1958 | No | ||
| 115 | HOXB5 | 15387 | -1.922 | -0.1964 | No | ||
| 116 | SYT3 | 15512 | -1.964 | -0.1990 | No | ||
| 117 | ETV1 | 15743 | -2.044 | -0.2064 | No | ||
| 118 | ADAMTS3 | 15755 | -2.048 | -0.2038 | No | ||
| 119 | ANAPC13 | 15933 | -2.103 | -0.2087 | No | ||
| 120 | CFL1 | 16028 | -2.129 | -0.2097 | No | ||
| 121 | HOXA7 | 16112 | -2.156 | -0.2102 | No | ||
| 122 | SLC31A2 | 16327 | -2.224 | -0.2166 | No | ||
| 123 | NRIP3 | 16597 | -2.322 | -0.2253 | No | ||
| 124 | PTGES2 | 16681 | -2.350 | -0.2255 | No | ||
| 125 | TGFB2 | 17183 | -2.526 | -0.2446 | No | ||
| 126 | FGF14 | 17196 | -2.532 | -0.2413 | No | ||
| 127 | CD164 | 17269 | -2.560 | -0.2406 | No | ||
| 128 | NEUROD1 | 17432 | -2.624 | -0.2440 | No | ||
| 129 | BOK | 17462 | -2.635 | -0.2413 | No | ||
| 130 | THUMPD2 | 17535 | -2.658 | -0.2405 | No | ||
| 131 | SRP72 | 17662 | -2.710 | -0.2421 | No | ||
| 132 | ADSS | 18029 | -2.867 | -0.2545 | No | ||
| 133 | SUPT16H | 18208 | -2.939 | -0.2581 | Yes | ||
| 134 | HMGA1 | 18236 | -2.950 | -0.2548 | Yes | ||
| 135 | TCF4 | 18239 | -2.951 | -0.2503 | Yes | ||
| 136 | DZIP1 | 18314 | -2.983 | -0.2491 | Yes | ||
| 137 | DSCAM | 18339 | -2.997 | -0.2456 | Yes | ||
| 138 | PTMA | 18360 | -3.006 | -0.2418 | Yes | ||
| 139 | NDUFS1 | 18468 | -3.057 | -0.2420 | Yes | ||
| 140 | NFX1 | 18775 | -3.224 | -0.2511 | Yes | ||
| 141 | EIF4B | 18819 | -3.244 | -0.2481 | Yes | ||
| 142 | SYNCRIP | 18869 | -3.273 | -0.2452 | Yes | ||
| 143 | OPRS1 | 18908 | -3.303 | -0.2419 | Yes | ||
| 144 | AMPD2 | 18964 | -3.332 | -0.2393 | Yes | ||
| 145 | AKAP12 | 18976 | -3.339 | -0.2346 | Yes | ||
| 146 | FGF6 | 19039 | -3.374 | -0.2322 | Yes | ||
| 147 | SCRT2 | 19174 | -3.467 | -0.2330 | Yes | ||
| 148 | XPO1 | 19211 | -3.485 | -0.2293 | Yes | ||
| 149 | DHX35 | 19451 | -3.648 | -0.2346 | Yes | ||
| 150 | EIF3S10 | 19465 | -3.661 | -0.2296 | Yes | ||
| 151 | ATF4 | 19609 | -3.777 | -0.2303 | Yes | ||
| 152 | RAI14 | 19616 | -3.781 | -0.2247 | Yes | ||
| 153 | MRPL40 | 19681 | -3.832 | -0.2218 | Yes | ||
| 154 | CCAR1 | 19715 | -3.866 | -0.2173 | Yes | ||
| 155 | ARMET | 19739 | -3.891 | -0.2123 | Yes | ||
| 156 | PLCG2 | 19865 | -4.009 | -0.2119 | Yes | ||
| 157 | RCOR2 | 19961 | -4.086 | -0.2099 | Yes | ||
| 158 | ATP6V1C1 | 19967 | -4.092 | -0.2038 | Yes | ||
| 159 | RAB3IL1 | 20225 | -4.385 | -0.2089 | Yes | ||
| 160 | NUDC | 20381 | -4.549 | -0.2089 | Yes | ||
| 161 | PABPC1 | 20409 | -4.595 | -0.2031 | Yes | ||
| 162 | TIMM10 | 20474 | -4.692 | -0.1988 | Yes | ||
| 163 | OPRD1 | 20484 | -4.703 | -0.1919 | Yes | ||
| 164 | BAX | 20631 | -4.885 | -0.1911 | Yes | ||
| 165 | HNRPA1 | 20763 | -5.130 | -0.1892 | Yes | ||
| 166 | AKAP1 | 20869 | -5.283 | -0.1858 | Yes | ||
| 167 | UBXD3 | 20939 | -5.375 | -0.1807 | Yes | ||
| 168 | TFRC | 20958 | -5.404 | -0.1731 | Yes | ||
| 169 | COMMD3 | 20996 | -5.475 | -0.1664 | Yes | ||
| 170 | TCERG1 | 21064 | -5.636 | -0.1607 | Yes | ||
| 171 | MTHFD1 | 21089 | -5.684 | -0.1531 | Yes | ||
| 172 | NPM1 | 21234 | -6.089 | -0.1503 | Yes | ||
| 173 | RCL1 | 21292 | -6.273 | -0.1432 | Yes | ||
| 174 | PRPS1 | 21340 | -6.411 | -0.1354 | Yes | ||
| 175 | CPT1A | 21349 | -6.433 | -0.1259 | Yes | ||
| 176 | SNX5 | 21360 | -6.467 | -0.1163 | Yes | ||
| 177 | SUCLG2 | 21556 | -7.140 | -0.1142 | Yes | ||
| 178 | MRPL27 | 21560 | -7.154 | -0.1033 | Yes | ||
| 179 | QTRT1 | 21616 | -7.430 | -0.0944 | Yes | ||
| 180 | PA2G4 | 21722 | -8.235 | -0.0864 | Yes | ||
| 181 | SNTB2 | 21730 | -8.300 | -0.0739 | Yes | ||
| 182 | NOTCH1 | 21772 | -8.813 | -0.0622 | Yes | ||
| 183 | AATF | 21801 | -9.220 | -0.0492 | Yes | ||
| 184 | PSME3 | 21823 | -9.439 | -0.0356 | Yes | ||
| 185 | TCOF1 | 21838 | -9.974 | -0.0208 | Yes | ||
| 186 | CAD | 21940 | -16.671 | 0.0003 | Yes |
