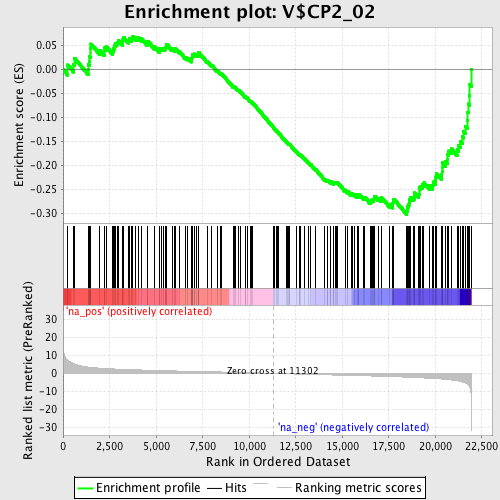
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_SCID |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | V$CP2_02 |
| Enrichment Score (ES) | -0.30199793 |
| Normalized Enrichment Score (NES) | -1.4485755 |
| Nominal p-value | 0.01058201 |
| FDR q-value | 0.40952557 |
| FWER p-Value | 0.979 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RAB10 | 252 | 7.194 | 0.0096 | No | ||
| 2 | AIM1L | 576 | 5.219 | 0.0101 | No | ||
| 3 | YWHAG | 608 | 5.096 | 0.0237 | No | ||
| 4 | MORF4L2 | 1347 | 3.499 | 0.0001 | No | ||
| 5 | HIF1A | 1360 | 3.474 | 0.0098 | No | ||
| 6 | GSK3B | 1406 | 3.424 | 0.0178 | No | ||
| 7 | CRK | 1421 | 3.408 | 0.0272 | No | ||
| 8 | NRXN3 | 1468 | 3.349 | 0.0349 | No | ||
| 9 | DYRK2 | 1469 | 3.348 | 0.0448 | No | ||
| 10 | PTHLH | 1496 | 3.318 | 0.0533 | No | ||
| 11 | TGFBR2 | 1969 | 2.868 | 0.0401 | No | ||
| 12 | RYR1 | 2206 | 2.693 | 0.0372 | No | ||
| 13 | PAK7 | 2208 | 2.693 | 0.0451 | No | ||
| 14 | NTRK3 | 2328 | 2.630 | 0.0473 | No | ||
| 15 | ARHGAP5 | 2666 | 2.449 | 0.0391 | No | ||
| 16 | CHD4 | 2724 | 2.423 | 0.0436 | No | ||
| 17 | KIF1C | 2753 | 2.408 | 0.0494 | No | ||
| 18 | TXNL5 | 2802 | 2.387 | 0.0542 | No | ||
| 19 | CS | 2897 | 2.343 | 0.0568 | No | ||
| 20 | TLX3 | 2959 | 2.318 | 0.0608 | No | ||
| 21 | HOXB6 | 3191 | 2.223 | 0.0567 | No | ||
| 22 | PPP1R16B | 3211 | 2.216 | 0.0624 | No | ||
| 23 | NFATC4 | 3239 | 2.205 | 0.0676 | No | ||
| 24 | PLXNA3 | 3510 | 2.098 | 0.0614 | No | ||
| 25 | CREB1 | 3579 | 2.070 | 0.0644 | No | ||
| 26 | ADNP | 3699 | 2.027 | 0.0649 | No | ||
| 27 | DHH | 3724 | 2.017 | 0.0697 | No | ||
| 28 | DSCAM | 3910 | 1.956 | 0.0670 | No | ||
| 29 | MMP11 | 4049 | 1.910 | 0.0663 | No | ||
| 30 | COX8C | 4206 | 1.861 | 0.0646 | No | ||
| 31 | ZFHX1B | 4532 | 1.759 | 0.0548 | No | ||
| 32 | AQP6 | 4554 | 1.750 | 0.0590 | No | ||
| 33 | PODN | 4914 | 1.630 | 0.0473 | No | ||
| 34 | PCDHB4 | 5176 | 1.549 | 0.0399 | No | ||
| 35 | NR2F1 | 5195 | 1.544 | 0.0436 | No | ||
| 36 | ALDH3B1 | 5270 | 1.524 | 0.0447 | No | ||
| 37 | H3F3B | 5398 | 1.490 | 0.0433 | No | ||
| 38 | GSH1 | 5475 | 1.466 | 0.0441 | No | ||
| 39 | RTN1 | 5510 | 1.457 | 0.0468 | No | ||
| 40 | MAPRE3 | 5535 | 1.451 | 0.0500 | No | ||
| 41 | CDC42EP3 | 5564 | 1.442 | 0.0529 | No | ||
| 42 | MID2 | 5867 | 1.366 | 0.0431 | No | ||
| 43 | KLHDC3 | 5983 | 1.336 | 0.0417 | No | ||
| 44 | TUFT1 | 6028 | 1.328 | 0.0436 | No | ||
| 45 | EFNA3 | 6250 | 1.273 | 0.0372 | No | ||
| 46 | NEK8 | 6584 | 1.200 | 0.0255 | No | ||
| 47 | EPN3 | 6707 | 1.168 | 0.0233 | No | ||
| 48 | FOXA1 | 6873 | 1.128 | 0.0190 | No | ||
| 49 | CAMK2G | 6891 | 1.124 | 0.0216 | No | ||
| 50 | ADRA1A | 6909 | 1.120 | 0.0241 | No | ||
| 51 | FOXL2 | 6926 | 1.115 | 0.0266 | No | ||
| 52 | CTNND1 | 6964 | 1.108 | 0.0282 | No | ||
| 53 | XPNPEP1 | 6968 | 1.107 | 0.0313 | No | ||
| 54 | KIFAP3 | 7075 | 1.081 | 0.0296 | No | ||
| 55 | ACTN1 | 7081 | 1.079 | 0.0326 | No | ||
| 56 | MDK | 7179 | 1.056 | 0.0312 | No | ||
| 57 | BDNF | 7264 | 1.037 | 0.0304 | No | ||
| 58 | PHPT1 | 7269 | 1.036 | 0.0333 | No | ||
| 59 | PITX2 | 7285 | 1.032 | 0.0356 | No | ||
| 60 | LDB2 | 7757 | 0.929 | 0.0167 | No | ||
| 61 | SCAMP5 | 7996 | 0.872 | 0.0084 | No | ||
| 62 | GNB3 | 8275 | 0.808 | -0.0020 | No | ||
| 63 | KCNC2 | 8475 | 0.766 | -0.0089 | No | ||
| 64 | PPFIA2 | 8527 | 0.754 | -0.0090 | No | ||
| 65 | RAB30 | 9131 | 0.607 | -0.0349 | No | ||
| 66 | ZNF385 | 9184 | 0.597 | -0.0355 | No | ||
| 67 | STARD13 | 9248 | 0.582 | -0.0367 | No | ||
| 68 | CNTN6 | 9421 | 0.543 | -0.0430 | No | ||
| 69 | PTCRA | 9508 | 0.519 | -0.0455 | No | ||
| 70 | NR1D1 | 9787 | 0.441 | -0.0569 | No | ||
| 71 | CITED1 | 9797 | 0.438 | -0.0560 | No | ||
| 72 | DDIT4 | 9891 | 0.412 | -0.0591 | No | ||
| 73 | RNF121 | 10069 | 0.364 | -0.0662 | No | ||
| 74 | HSPA1A | 10147 | 0.339 | -0.0687 | No | ||
| 75 | KIF5A | 10196 | 0.328 | -0.0699 | No | ||
| 76 | TNNC1 | 11322 | -0.005 | -0.1216 | No | ||
| 77 | GMPPB | 11376 | -0.024 | -0.1239 | No | ||
| 78 | DUSP6 | 11469 | -0.057 | -0.1280 | No | ||
| 79 | GRB7 | 11541 | -0.079 | -0.1310 | No | ||
| 80 | IRX3 | 11572 | -0.090 | -0.1321 | No | ||
| 81 | RHBDF1 | 11979 | -0.205 | -0.1502 | No | ||
| 82 | MNT | 12040 | -0.223 | -0.1523 | No | ||
| 83 | NGFRAP1 | 12096 | -0.245 | -0.1541 | No | ||
| 84 | BMP4 | 12156 | -0.262 | -0.1560 | No | ||
| 85 | TRPV2 | 12182 | -0.269 | -0.1564 | No | ||
| 86 | KCNE3 | 12553 | -0.380 | -0.1723 | No | ||
| 87 | GLTP | 12692 | -0.419 | -0.1774 | No | ||
| 88 | MOGAT2 | 12702 | -0.422 | -0.1765 | No | ||
| 89 | MRPL14 | 12765 | -0.442 | -0.1781 | No | ||
| 90 | LZTS2 | 12970 | -0.495 | -0.1860 | No | ||
| 91 | NNAT | 13194 | -0.558 | -0.1946 | No | ||
| 92 | ETV1 | 13308 | -0.590 | -0.1980 | No | ||
| 93 | UPK2 | 13566 | -0.667 | -0.2079 | No | ||
| 94 | NHS | 14059 | -0.794 | -0.2281 | No | ||
| 95 | RPL41 | 14180 | -0.824 | -0.2312 | No | ||
| 96 | CAPN6 | 14232 | -0.835 | -0.2311 | No | ||
| 97 | RHBDL1 | 14356 | -0.868 | -0.2342 | No | ||
| 98 | DSC2 | 14388 | -0.874 | -0.2330 | No | ||
| 99 | HSPA1L | 14541 | -0.917 | -0.2373 | No | ||
| 100 | RCOR2 | 14551 | -0.919 | -0.2350 | No | ||
| 101 | RGS8 | 14616 | -0.936 | -0.2352 | No | ||
| 102 | FEV | 14678 | -0.953 | -0.2352 | No | ||
| 103 | IL1RAPL1 | 14730 | -0.965 | -0.2347 | No | ||
| 104 | DEF6 | 15154 | -1.088 | -0.2509 | No | ||
| 105 | PPP1R9B | 15294 | -1.128 | -0.2540 | No | ||
| 106 | ARHGAP26 | 15482 | -1.177 | -0.2591 | No | ||
| 107 | GPRC5C | 15528 | -1.190 | -0.2577 | No | ||
| 108 | BACH1 | 15657 | -1.224 | -0.2600 | No | ||
| 109 | SLC25A14 | 15793 | -1.258 | -0.2625 | No | ||
| 110 | PRKCH | 15876 | -1.279 | -0.2625 | No | ||
| 111 | CAP1 | 15891 | -1.281 | -0.2593 | No | ||
| 112 | CILP2 | 16163 | -1.350 | -0.2678 | No | ||
| 113 | TRIM8 | 16203 | -1.361 | -0.2656 | No | ||
| 114 | ARX | 16515 | -1.445 | -0.2756 | No | ||
| 115 | FGF9 | 16548 | -1.454 | -0.2728 | No | ||
| 116 | FBS1 | 16604 | -1.470 | -0.2710 | No | ||
| 117 | STAC2 | 16695 | -1.493 | -0.2708 | No | ||
| 118 | WNT10B | 16710 | -1.500 | -0.2670 | No | ||
| 119 | NOL4 | 16745 | -1.511 | -0.2641 | No | ||
| 120 | LTBP1 | 16926 | -1.562 | -0.2678 | No | ||
| 121 | PAPPA | 17083 | -1.611 | -0.2702 | No | ||
| 122 | DCX | 17088 | -1.613 | -0.2656 | No | ||
| 123 | PRDM4 | 17550 | -1.768 | -0.2816 | No | ||
| 124 | SRMS | 17704 | -1.820 | -0.2833 | No | ||
| 125 | GNAQ | 17710 | -1.822 | -0.2781 | No | ||
| 126 | FOXP2 | 17726 | -1.828 | -0.2734 | No | ||
| 127 | KCTD15 | 17768 | -1.841 | -0.2699 | No | ||
| 128 | JUNB | 18468 | -2.107 | -0.2958 | Yes | ||
| 129 | ATP10D | 18485 | -2.113 | -0.2903 | Yes | ||
| 130 | CPNE1 | 18502 | -2.122 | -0.2848 | Yes | ||
| 131 | GPC3 | 18559 | -2.141 | -0.2811 | Yes | ||
| 132 | DDAH2 | 18588 | -2.150 | -0.2760 | Yes | ||
| 133 | HTATIP | 18602 | -2.156 | -0.2703 | Yes | ||
| 134 | PFTK1 | 18647 | -2.182 | -0.2659 | Yes | ||
| 135 | MAPK3 | 18809 | -2.253 | -0.2666 | Yes | ||
| 136 | RBBP6 | 18884 | -2.286 | -0.2633 | Yes | ||
| 137 | EPC1 | 18893 | -2.291 | -0.2569 | Yes | ||
| 138 | TRERF1 | 19119 | -2.401 | -0.2602 | Yes | ||
| 139 | UBE1 | 19127 | -2.406 | -0.2535 | Yes | ||
| 140 | SHB | 19128 | -2.407 | -0.2464 | Yes | ||
| 141 | PRKACA | 19225 | -2.460 | -0.2435 | Yes | ||
| 142 | FRMD3 | 19303 | -2.501 | -0.2397 | Yes | ||
| 143 | SLC25A28 | 19390 | -2.542 | -0.2362 | Yes | ||
| 144 | TRAF4 | 19681 | -2.699 | -0.2416 | Yes | ||
| 145 | NRIP2 | 19864 | -2.806 | -0.2417 | Yes | ||
| 146 | PRKAG1 | 19884 | -2.818 | -0.2342 | Yes | ||
| 147 | FLOT2 | 19989 | -2.884 | -0.2305 | Yes | ||
| 148 | CALD1 | 20032 | -2.916 | -0.2239 | Yes | ||
| 149 | HPCAL1 | 20078 | -2.948 | -0.2173 | Yes | ||
| 150 | DDR1 | 20326 | -3.118 | -0.2194 | Yes | ||
| 151 | RASSF1 | 20358 | -3.144 | -0.2116 | Yes | ||
| 152 | SLC9A1 | 20386 | -3.160 | -0.2035 | Yes | ||
| 153 | BTBD14B | 20391 | -3.164 | -0.1944 | Yes | ||
| 154 | FKBP2 | 20548 | -3.305 | -0.1919 | Yes | ||
| 155 | MAST2 | 20654 | -3.411 | -0.1866 | Yes | ||
| 156 | KCNH2 | 20668 | -3.418 | -0.1772 | Yes | ||
| 157 | HCFC1 | 20728 | -3.478 | -0.1697 | Yes | ||
| 158 | TBC1D5 | 20849 | -3.627 | -0.1645 | Yes | ||
| 159 | GABARAP | 21171 | -4.086 | -0.1672 | Yes | ||
| 160 | DNAJB1 | 21242 | -4.240 | -0.1579 | Yes | ||
| 161 | HECTD1 | 21343 | -4.458 | -0.1494 | Yes | ||
| 162 | NRF1 | 21445 | -4.729 | -0.1401 | Yes | ||
| 163 | LHX2 | 21501 | -4.887 | -0.1283 | Yes | ||
| 164 | FLI1 | 21627 | -5.352 | -0.1183 | Yes | ||
| 165 | TLN1 | 21707 | -5.706 | -0.1051 | Yes | ||
| 166 | SLC2A9 | 21750 | -6.089 | -0.0891 | Yes | ||
| 167 | NISCH | 21769 | -6.206 | -0.0717 | Yes | ||
| 168 | PPP2R5C | 21843 | -7.340 | -0.0534 | Yes | ||
| 169 | BCL11B | 21858 | -7.702 | -0.0314 | Yes | ||
| 170 | CLDN4 | 21928 | -12.050 | 0.0009 | Yes |
