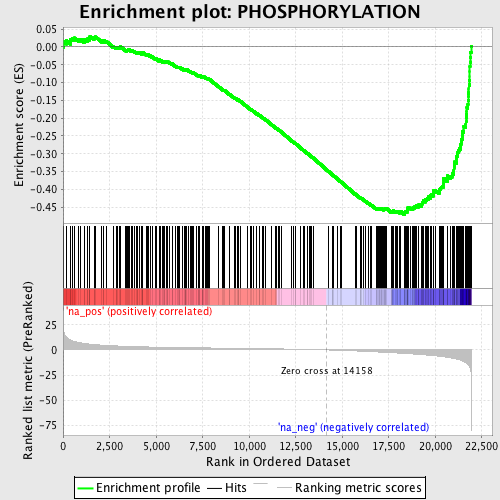
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set01_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | PHOSPHORYLATION |
| Enrichment Score (ES) | -0.47057378 |
| Normalized Enrichment Score (NES) | -2.074751 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.010483386 |
| FWER p-Value | 0.056 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IL5 | 11 | 22.605 | 0.0164 | No | ||
| 2 | PRDX4 | 193 | 12.146 | 0.0172 | No | ||
| 3 | MCM7 | 398 | 9.851 | 0.0152 | No | ||
| 4 | DYRK3 | 410 | 9.697 | 0.0219 | No | ||
| 5 | EPHB2 | 502 | 9.013 | 0.0245 | No | ||
| 6 | CDK2AP1 | 622 | 8.326 | 0.0252 | No | ||
| 7 | AURKA | 833 | 7.454 | 0.0211 | No | ||
| 8 | IKBKAP | 954 | 7.011 | 0.0209 | No | ||
| 9 | RIPK4 | 1133 | 6.476 | 0.0175 | No | ||
| 10 | FGR | 1144 | 6.449 | 0.0219 | No | ||
| 11 | PLK4 | 1306 | 6.023 | 0.0190 | No | ||
| 12 | UQCRC1 | 1327 | 5.967 | 0.0225 | No | ||
| 13 | F2 | 1399 | 5.815 | 0.0236 | No | ||
| 14 | CDC42BPA | 1415 | 5.784 | 0.0272 | No | ||
| 15 | EGFR | 1441 | 5.720 | 0.0304 | No | ||
| 16 | CCL2 | 1663 | 5.299 | 0.0241 | No | ||
| 17 | TSSK4 | 1673 | 5.287 | 0.0277 | No | ||
| 18 | CDKN2C | 1747 | 5.171 | 0.0282 | No | ||
| 19 | FES | 2066 | 4.731 | 0.0171 | No | ||
| 20 | CDKN2A | 2159 | 4.589 | 0.0163 | No | ||
| 21 | GMFB | 2187 | 4.553 | 0.0184 | No | ||
| 22 | EPHA8 | 2318 | 4.394 | 0.0157 | No | ||
| 23 | MAPKAPK2 | 2720 | 3.998 | 0.0002 | No | ||
| 24 | TSSK6 | 2842 | 3.914 | -0.0024 | No | ||
| 25 | MAPK12 | 2869 | 3.894 | -0.0007 | No | ||
| 26 | CDKN1A | 2944 | 3.835 | -0.0012 | No | ||
| 27 | ACVR1 | 3012 | 3.791 | -0.0015 | No | ||
| 28 | IL3 | 3065 | 3.756 | -0.0011 | No | ||
| 29 | IL31RA | 3090 | 3.740 | 0.0006 | No | ||
| 30 | TTN | 3353 | 3.533 | -0.0088 | No | ||
| 31 | MAP3K6 | 3427 | 3.476 | -0.0096 | No | ||
| 32 | MAST1 | 3467 | 3.451 | -0.0088 | No | ||
| 33 | TNKS | 3495 | 3.439 | -0.0075 | No | ||
| 34 | UQCRC2 | 3557 | 3.395 | -0.0077 | No | ||
| 35 | STK19 | 3697 | 3.311 | -0.0117 | No | ||
| 36 | LATS2 | 3716 | 3.300 | -0.0100 | No | ||
| 37 | NF2 | 3843 | 3.233 | -0.0134 | No | ||
| 38 | TNK1 | 3938 | 3.182 | -0.0154 | No | ||
| 39 | BTK | 3984 | 3.157 | -0.0151 | No | ||
| 40 | AURKC | 4018 | 3.134 | -0.0143 | No | ||
| 41 | VRK1 | 4077 | 3.102 | -0.0146 | No | ||
| 42 | UHMK1 | 4216 | 3.035 | -0.0187 | No | ||
| 43 | PRKCI | 4238 | 3.023 | -0.0174 | No | ||
| 44 | UQCRB | 4258 | 3.014 | -0.0160 | No | ||
| 45 | VRK2 | 4288 | 3.001 | -0.0151 | No | ||
| 46 | MYLK | 4466 | 2.925 | -0.0211 | No | ||
| 47 | GRK1 | 4525 | 2.902 | -0.0216 | No | ||
| 48 | BMPR1A | 4580 | 2.883 | -0.0219 | No | ||
| 49 | LTK | 4686 | 2.841 | -0.0246 | No | ||
| 50 | EIF2AK4 | 4802 | 2.792 | -0.0278 | No | ||
| 51 | BMX | 4945 | 2.739 | -0.0323 | No | ||
| 52 | BMPR1B | 5009 | 2.718 | -0.0332 | No | ||
| 53 | CBLC | 5155 | 2.667 | -0.0379 | No | ||
| 54 | RET | 5159 | 2.665 | -0.0360 | No | ||
| 55 | MAP3K9 | 5225 | 2.645 | -0.0371 | No | ||
| 56 | BRSK1 | 5357 | 2.606 | -0.0412 | No | ||
| 57 | GSG2 | 5396 | 2.594 | -0.0410 | No | ||
| 58 | CAMK2B | 5454 | 2.573 | -0.0417 | No | ||
| 59 | LIMK1 | 5462 | 2.569 | -0.0401 | No | ||
| 60 | INSR | 5561 | 2.539 | -0.0427 | No | ||
| 61 | MATK | 5604 | 2.524 | -0.0427 | No | ||
| 62 | STK3 | 5627 | 2.519 | -0.0419 | No | ||
| 63 | RAGE | 5709 | 2.492 | -0.0437 | No | ||
| 64 | CAMK1 | 5853 | 2.445 | -0.0485 | No | ||
| 65 | IRAK3 | 5862 | 2.443 | -0.0470 | No | ||
| 66 | INHBA | 6055 | 2.380 | -0.0541 | No | ||
| 67 | SRP72 | 6156 | 2.350 | -0.0570 | No | ||
| 68 | MYOD1 | 6186 | 2.340 | -0.0565 | No | ||
| 69 | PIK3R1 | 6230 | 2.327 | -0.0568 | No | ||
| 70 | KALRN | 6274 | 2.311 | -0.0570 | No | ||
| 71 | ADAM10 | 6405 | 2.277 | -0.0613 | No | ||
| 72 | IL12A | 6521 | 2.246 | -0.0649 | No | ||
| 73 | MAPK4 | 6559 | 2.234 | -0.0650 | No | ||
| 74 | HCK | 6566 | 2.233 | -0.0636 | No | ||
| 75 | IL20 | 6572 | 2.232 | -0.0621 | No | ||
| 76 | PGK2 | 6641 | 2.212 | -0.0636 | No | ||
| 77 | ACVRL1 | 6762 | 2.181 | -0.0675 | No | ||
| 78 | MOBKL1A | 6817 | 2.166 | -0.0684 | No | ||
| 79 | MAPK15 | 6907 | 2.138 | -0.0709 | No | ||
| 80 | BMP4 | 6968 | 2.126 | -0.0721 | No | ||
| 81 | ERG | 7011 | 2.116 | -0.0724 | No | ||
| 82 | EPHA5 | 7169 | 2.072 | -0.0781 | No | ||
| 83 | PSEN1 | 7252 | 2.051 | -0.0804 | No | ||
| 84 | SRPK1 | 7263 | 2.046 | -0.0793 | No | ||
| 85 | CCND1 | 7333 | 2.028 | -0.0810 | No | ||
| 86 | RPS6KA5 | 7337 | 2.027 | -0.0796 | No | ||
| 87 | LATS1 | 7482 | 1.992 | -0.0847 | No | ||
| 88 | BRSK2 | 7492 | 1.989 | -0.0836 | No | ||
| 89 | BCKDK | 7507 | 1.985 | -0.0828 | No | ||
| 90 | PMVK | 7525 | 1.981 | -0.0821 | No | ||
| 91 | WNK2 | 7648 | 1.947 | -0.0863 | No | ||
| 92 | PRKG2 | 7727 | 1.924 | -0.0884 | No | ||
| 93 | CCL11 | 7767 | 1.914 | -0.0888 | No | ||
| 94 | PRKAA2 | 7811 | 1.903 | -0.0894 | No | ||
| 95 | NEK4 | 7878 | 1.888 | -0.0910 | No | ||
| 96 | TSSK3 | 8327 | 1.782 | -0.1103 | No | ||
| 97 | FRK | 8587 | 1.722 | -0.1210 | No | ||
| 98 | MAP3K2 | 8636 | 1.709 | -0.1219 | No | ||
| 99 | UQCRH | 8656 | 1.702 | -0.1215 | No | ||
| 100 | CD24 | 8918 | 1.644 | -0.1323 | No | ||
| 101 | NEK11 | 8953 | 1.637 | -0.1327 | No | ||
| 102 | WNK3 | 9218 | 1.571 | -0.1437 | No | ||
| 103 | ERBB3 | 9243 | 1.565 | -0.1436 | No | ||
| 104 | ROCK2 | 9365 | 1.537 | -0.1481 | No | ||
| 105 | MARK1 | 9398 | 1.530 | -0.1484 | No | ||
| 106 | TRIB1 | 9436 | 1.520 | -0.1489 | No | ||
| 107 | PRKD1 | 9525 | 1.498 | -0.1519 | No | ||
| 108 | BCR | 9930 | 1.403 | -0.1695 | No | ||
| 109 | PTK6 | 10077 | 1.363 | -0.1752 | No | ||
| 110 | WNK4 | 10123 | 1.354 | -0.1763 | No | ||
| 111 | EREG | 10245 | 1.323 | -0.1809 | No | ||
| 112 | MERTK | 10414 | 1.284 | -0.1876 | No | ||
| 113 | IPPK | 10526 | 1.253 | -0.1918 | No | ||
| 114 | PRKCZ | 10572 | 1.242 | -0.1930 | No | ||
| 115 | DAPK2 | 10688 | 1.217 | -0.1974 | No | ||
| 116 | ABL2 | 10761 | 1.198 | -0.1998 | No | ||
| 117 | GLMN | 10882 | 1.165 | -0.2045 | No | ||
| 118 | STK36 | 11194 | 1.087 | -0.2180 | No | ||
| 119 | IGFBP3 | 11414 | 1.019 | -0.2273 | No | ||
| 120 | FASTK | 11446 | 1.008 | -0.2280 | No | ||
| 121 | NEK6 | 11571 | 0.974 | -0.2330 | No | ||
| 122 | BARD1 | 11636 | 0.955 | -0.2352 | No | ||
| 123 | ERBB2 | 11721 | 0.927 | -0.2384 | No | ||
| 124 | EGF | 12295 | 0.756 | -0.2643 | No | ||
| 125 | IL4 | 12402 | 0.724 | -0.2687 | No | ||
| 126 | TDGF1 | 12467 | 0.703 | -0.2711 | No | ||
| 127 | CARD14 | 12495 | 0.694 | -0.2718 | No | ||
| 128 | CDC2L1 | 12751 | 0.608 | -0.2831 | No | ||
| 129 | CD81 | 12940 | 0.537 | -0.2914 | No | ||
| 130 | MYO3A | 12986 | 0.518 | -0.2931 | No | ||
| 131 | PKN3 | 13131 | 0.464 | -0.2994 | No | ||
| 132 | ERN2 | 13142 | 0.460 | -0.2995 | No | ||
| 133 | NUAK2 | 13262 | 0.412 | -0.3047 | No | ||
| 134 | TSSK2 | 13314 | 0.389 | -0.3067 | No | ||
| 135 | SNF1LK | 13371 | 0.369 | -0.3091 | No | ||
| 136 | PIM1 | 13451 | 0.338 | -0.3124 | No | ||
| 137 | INHA | 14268 | -0.072 | -0.3501 | No | ||
| 138 | PNKP | 14482 | -0.211 | -0.3597 | No | ||
| 139 | CDKN2B | 14499 | -0.220 | -0.3603 | No | ||
| 140 | IHPK3 | 14545 | -0.244 | -0.3622 | No | ||
| 141 | PRKAA1 | 14731 | -0.364 | -0.3705 | No | ||
| 142 | ICK | 14903 | -0.477 | -0.3780 | No | ||
| 143 | STK38L | 14967 | -0.508 | -0.3805 | No | ||
| 144 | PLK3 | 15685 | -1.005 | -0.4129 | No | ||
| 145 | MAP4K3 | 15752 | -1.048 | -0.4151 | No | ||
| 146 | STK11 | 15968 | -1.215 | -0.4241 | No | ||
| 147 | CDK9 | 15989 | -1.229 | -0.4241 | No | ||
| 148 | CDC42BPB | 16031 | -1.265 | -0.4251 | No | ||
| 149 | CAMKK2 | 16123 | -1.344 | -0.4283 | No | ||
| 150 | DAPK1 | 16250 | -1.448 | -0.4330 | No | ||
| 151 | DYRK1B | 16393 | -1.569 | -0.4384 | No | ||
| 152 | TAF1 | 16532 | -1.684 | -0.4435 | No | ||
| 153 | F2R | 16595 | -1.749 | -0.4450 | No | ||
| 154 | ABI2 | 16841 | -1.966 | -0.4549 | No | ||
| 155 | RPS6KA4 | 16863 | -1.997 | -0.4543 | No | ||
| 156 | CDKL1 | 16876 | -2.009 | -0.4534 | No | ||
| 157 | PRKCH | 16959 | -2.083 | -0.4556 | No | ||
| 158 | DMPK | 16977 | -2.101 | -0.4548 | No | ||
| 159 | HCLS1 | 17013 | -2.142 | -0.4548 | No | ||
| 160 | AKT1 | 17066 | -2.193 | -0.4556 | No | ||
| 161 | DAPK3 | 17080 | -2.200 | -0.4545 | No | ||
| 162 | TYK2 | 17095 | -2.213 | -0.4535 | No | ||
| 163 | PRKG1 | 17162 | -2.273 | -0.4549 | No | ||
| 164 | PICK1 | 17237 | -2.332 | -0.4565 | No | ||
| 165 | ERC1 | 17266 | -2.356 | -0.4561 | No | ||
| 166 | CDC42BPG | 17294 | -2.382 | -0.4555 | No | ||
| 167 | IRAK2 | 17297 | -2.389 | -0.4538 | No | ||
| 168 | MAST2 | 17317 | -2.404 | -0.4529 | No | ||
| 169 | LIPE | 17401 | -2.487 | -0.4549 | No | ||
| 170 | MAP2K6 | 17632 | -2.723 | -0.4634 | No | ||
| 171 | CHUK | 17643 | -2.730 | -0.4618 | No | ||
| 172 | CTBP1 | 17721 | -2.801 | -0.4633 | No | ||
| 173 | MARK4 | 17722 | -2.802 | -0.4612 | No | ||
| 174 | PAK1 | 17746 | -2.826 | -0.4601 | No | ||
| 175 | MAP3K8 | 17860 | -2.954 | -0.4631 | No | ||
| 176 | JAK3 | 17929 | -3.030 | -0.4640 | No | ||
| 177 | CD80 | 17952 | -3.058 | -0.4627 | No | ||
| 178 | MAPK3 | 17962 | -3.067 | -0.4608 | No | ||
| 179 | MKNK2 | 18089 | -3.199 | -0.4643 | No | ||
| 180 | NDUFS4 | 18111 | -3.233 | -0.4628 | No | ||
| 181 | AKT3 | 18133 | -3.248 | -0.4613 | No | ||
| 182 | BRAF | 18334 | -3.456 | -0.4680 | Yes | ||
| 183 | CAMK2A | 18342 | -3.465 | -0.4657 | Yes | ||
| 184 | MKNK1 | 18365 | -3.498 | -0.4641 | Yes | ||
| 185 | OXSR1 | 18380 | -3.517 | -0.4621 | Yes | ||
| 186 | STK16 | 18441 | -3.592 | -0.4622 | Yes | ||
| 187 | EIF2AK1 | 18481 | -3.633 | -0.4613 | Yes | ||
| 188 | IHPK1 | 18482 | -3.633 | -0.4585 | Yes | ||
| 189 | ILK | 18512 | -3.670 | -0.4571 | Yes | ||
| 190 | JAK2 | 18520 | -3.677 | -0.4547 | Yes | ||
| 191 | PRKCD | 18528 | -3.688 | -0.4522 | Yes | ||
| 192 | CCNT1 | 18554 | -3.710 | -0.4506 | Yes | ||
| 193 | FYB | 18658 | -3.819 | -0.4525 | Yes | ||
| 194 | MAP3K10 | 18760 | -3.957 | -0.4542 | Yes | ||
| 195 | PRPF4B | 18784 | -3.983 | -0.4523 | Yes | ||
| 196 | ERN1 | 18848 | -4.074 | -0.4521 | Yes | ||
| 197 | PAK2 | 18865 | -4.102 | -0.4498 | Yes | ||
| 198 | CSNK1G2 | 18900 | -4.144 | -0.4483 | Yes | ||
| 199 | SGK3 | 18946 | -4.207 | -0.4472 | Yes | ||
| 200 | EIF2AK3 | 18978 | -4.241 | -0.4454 | Yes | ||
| 201 | RAF1 | 19074 | -4.383 | -0.4465 | Yes | ||
| 202 | ALS2CR2 | 19089 | -4.405 | -0.4439 | Yes | ||
| 203 | PKN2 | 19121 | -4.460 | -0.4420 | Yes | ||
| 204 | TESK2 | 19259 | -4.659 | -0.4448 | Yes | ||
| 205 | PINK1 | 19279 | -4.705 | -0.4421 | Yes | ||
| 206 | PRKAG1 | 19286 | -4.710 | -0.4389 | Yes | ||
| 207 | MAP3K12 | 19326 | -4.778 | -0.4371 | Yes | ||
| 208 | MAP3K11 | 19341 | -4.796 | -0.4341 | Yes | ||
| 209 | PXK | 19343 | -4.802 | -0.4306 | Yes | ||
| 210 | PRKD3 | 19453 | -4.974 | -0.4319 | Yes | ||
| 211 | ULK1 | 19480 | -5.004 | -0.4293 | Yes | ||
| 212 | CSNK1D | 19547 | -5.118 | -0.4285 | Yes | ||
| 213 | CSNK1A1 | 19558 | -5.138 | -0.4252 | Yes | ||
| 214 | PRKX | 19622 | -5.230 | -0.4241 | Yes | ||
| 215 | RAD17 | 19642 | -5.256 | -0.4211 | Yes | ||
| 216 | CLCF1 | 19735 | -5.380 | -0.4213 | Yes | ||
| 217 | ABL1 | 19742 | -5.391 | -0.4175 | Yes | ||
| 218 | CAMK4 | 19772 | -5.438 | -0.4148 | Yes | ||
| 219 | COL4A3BP | 19878 | -5.631 | -0.4154 | Yes | ||
| 220 | PRKACB | 19908 | -5.677 | -0.4125 | Yes | ||
| 221 | ACVR1B | 19911 | -5.677 | -0.4083 | Yes | ||
| 222 | TNK2 | 19917 | -5.699 | -0.4043 | Yes | ||
| 223 | ROCK1 | 20004 | -5.882 | -0.4038 | Yes | ||
| 224 | TLK1 | 20203 | -6.276 | -0.4083 | Yes | ||
| 225 | BCL10 | 20231 | -6.352 | -0.4048 | Yes | ||
| 226 | TRIB2 | 20241 | -6.377 | -0.4004 | Yes | ||
| 227 | SNF1LK2 | 20268 | -6.428 | -0.3968 | Yes | ||
| 228 | ITGB2 | 20306 | -6.502 | -0.3936 | Yes | ||
| 229 | TXK | 20361 | -6.619 | -0.3911 | Yes | ||
| 230 | STK10 | 20418 | -6.774 | -0.3886 | Yes | ||
| 231 | MAP3K3 | 20422 | -6.791 | -0.3837 | Yes | ||
| 232 | PIM2 | 20446 | -6.848 | -0.3796 | Yes | ||
| 233 | MAP3K13 | 20449 | -6.853 | -0.3746 | Yes | ||
| 234 | PKN1 | 20455 | -6.883 | -0.3696 | Yes | ||
| 235 | DYRK1A | 20638 | -7.321 | -0.3725 | Yes | ||
| 236 | MINK1 | 20643 | -7.336 | -0.3672 | Yes | ||
| 237 | PRKCE | 20653 | -7.365 | -0.3621 | Yes | ||
| 238 | STK4 | 20820 | -7.881 | -0.3639 | Yes | ||
| 239 | TGFB1 | 20897 | -8.163 | -0.3612 | Yes | ||
| 240 | CSNK2A1 | 20926 | -8.269 | -0.3563 | Yes | ||
| 241 | CRKRS | 20965 | -8.430 | -0.3518 | Yes | ||
| 242 | WNK1 | 20997 | -8.567 | -0.3468 | Yes | ||
| 243 | ABI1 | 21003 | -8.586 | -0.3406 | Yes | ||
| 244 | MAPK6 | 21006 | -8.592 | -0.3342 | Yes | ||
| 245 | MAP4K4 | 21016 | -8.644 | -0.3282 | Yes | ||
| 246 | TAOK3 | 21049 | -8.758 | -0.3231 | Yes | ||
| 247 | CCND2 | 21139 | -9.105 | -0.3203 | Yes | ||
| 248 | GMFG | 21143 | -9.118 | -0.3136 | Yes | ||
| 249 | PCTK1 | 21155 | -9.160 | -0.3073 | Yes | ||
| 250 | LMTK2 | 21182 | -9.262 | -0.3015 | Yes | ||
| 251 | MAPK8 | 21208 | -9.408 | -0.2956 | Yes | ||
| 252 | TOLLIP | 21244 | -9.606 | -0.2900 | Yes | ||
| 253 | CDKN1B | 21294 | -9.870 | -0.2849 | Yes | ||
| 254 | SOCS1 | 21356 | -10.225 | -0.2801 | Yes | ||
| 255 | TGFBR1 | 21368 | -10.287 | -0.2728 | Yes | ||
| 256 | FRAP1 | 21383 | -10.384 | -0.2657 | Yes | ||
| 257 | CSK | 21416 | -10.654 | -0.2592 | Yes | ||
| 258 | MAP4K5 | 21442 | -10.831 | -0.2522 | Yes | ||
| 259 | GSK3B | 21455 | -10.935 | -0.2446 | Yes | ||
| 260 | SRPK2 | 21476 | -11.046 | -0.2372 | Yes | ||
| 261 | SNRK | 21532 | -11.545 | -0.2311 | Yes | ||
| 262 | ABI3 | 21533 | -11.552 | -0.2224 | Yes | ||
| 263 | TLK2 | 21642 | -12.867 | -0.2178 | Yes | ||
| 264 | DYRK2 | 21648 | -12.985 | -0.2083 | Yes | ||
| 265 | STAT1 | 21656 | -13.091 | -0.1988 | Yes | ||
| 266 | HIPK3 | 21676 | -13.309 | -0.1897 | Yes | ||
| 267 | PTK2B | 21693 | -13.678 | -0.1802 | Yes | ||
| 268 | TEC | 21697 | -13.757 | -0.1700 | Yes | ||
| 269 | IHPK2 | 21750 | -14.578 | -0.1614 | Yes | ||
| 270 | STK38 | 21758 | -14.684 | -0.1508 | Yes | ||
| 271 | LCK | 21778 | -15.166 | -0.1403 | Yes | ||
| 272 | IGF1R | 21784 | -15.295 | -0.1290 | Yes | ||
| 273 | CSNK1E | 21807 | -15.890 | -0.1181 | Yes | ||
| 274 | IKBKE | 21809 | -15.920 | -0.1062 | Yes | ||
| 275 | MAP4K2 | 21842 | -17.169 | -0.0948 | Yes | ||
| 276 | PCTK2 | 21857 | -17.748 | -0.0822 | Yes | ||
| 277 | MARK2 | 21858 | -17.748 | -0.0689 | Yes | ||
| 278 | STK17B | 21860 | -17.835 | -0.0555 | Yes | ||
| 279 | PRKCB1 | 21866 | -17.985 | -0.0423 | Yes | ||
| 280 | PDPK1 | 21879 | -18.696 | -0.0288 | Yes | ||
| 281 | CDKN2D | 21895 | -19.642 | -0.0148 | Yes | ||
| 282 | CCND3 | 21924 | -22.833 | 0.0011 | Yes |
