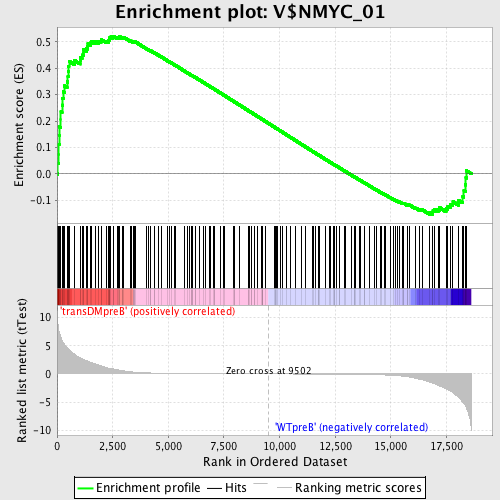
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | transDMpreB |
| GeneSet | V$NMYC_01 |
| Enrichment Score (ES) | 0.5218205 |
| Normalized Enrichment Score (NES) | 1.3455064 |
| Nominal p-value | 0.014652015 |
| FDR q-value | 0.23677205 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | FBL | 8955 | 36 | 8.936 | 0.0400 | Yes | ||
| 2 | PA2G4 | 5265 | 76 | 7.729 | 0.0743 | Yes | ||
| 3 | TIMM10 | 11390 2158 | 80 | 7.684 | 0.1102 | Yes | ||
| 4 | PPAT | 6081 | 93 | 7.493 | 0.1448 | Yes | ||
| 5 | ATP5G1 | 8636 | 101 | 7.285 | 0.1786 | Yes | ||
| 6 | HSPE1 | 9133 | 171 | 6.450 | 0.2052 | Yes | ||
| 7 | IPO4 | 7983 | 173 | 6.443 | 0.2354 | Yes | ||
| 8 | EIF4G1 | 22818 | 222 | 5.960 | 0.2608 | Yes | ||
| 9 | POLR3E | 18100 | 245 | 5.794 | 0.2869 | Yes | ||
| 10 | ATAD3A | 8440 | 275 | 5.592 | 0.3116 | Yes | ||
| 11 | FKBP5 | 23051 | 329 | 5.237 | 0.3333 | Yes | ||
| 12 | CEBPB | 8733 | 449 | 4.666 | 0.3488 | Yes | ||
| 13 | PPARGC1B | 9318 | 487 | 4.498 | 0.3679 | Yes | ||
| 14 | G6PC3 | 1221 20645 | 505 | 4.413 | 0.3877 | Yes | ||
| 15 | NOL5A | 12474 | 520 | 4.324 | 0.4073 | Yes | ||
| 16 | COPS7A | 11221 16997 | 535 | 4.263 | 0.4266 | Yes | ||
| 17 | LIG3 | 4998 | 783 | 3.486 | 0.4296 | Yes | ||
| 18 | MAPKAPK3 | 19003 | 1031 | 2.833 | 0.4295 | Yes | ||
| 19 | SLC20A1 | 14855 | 1056 | 2.794 | 0.4413 | Yes | ||
| 20 | ELOVL6 | 5009 | 1122 | 2.686 | 0.4504 | Yes | ||
| 21 | RAMP2 | 20660 | 1172 | 2.588 | 0.4599 | Yes | ||
| 22 | EIF4B | 13279 7979 | 1178 | 2.575 | 0.4717 | Yes | ||
| 23 | RBBP6 | 5365 | 1326 | 2.357 | 0.4748 | Yes | ||
| 24 | PANK3 | 20924 | 1368 | 2.298 | 0.4834 | Yes | ||
| 25 | PRDX4 | 24026 | 1386 | 2.261 | 0.4931 | Yes | ||
| 26 | VARS2 | 23265 1595 7588 | 1512 | 2.040 | 0.4959 | Yes | ||
| 27 | ZCCHC7 | 16225 | 1552 | 1.977 | 0.5031 | Yes | ||
| 28 | SRP72 | 16819 | 1714 | 1.747 | 0.5026 | Yes | ||
| 29 | HMOX1 | 18565 | 1867 | 1.582 | 0.5018 | Yes | ||
| 30 | BCL7C | 4442 | 1974 | 1.434 | 0.5028 | Yes | ||
| 31 | PES1 | 7236 | 1999 | 1.404 | 0.5081 | Yes | ||
| 32 | HSPD1 | 4078 9129 | 2210 | 1.146 | 0.5021 | Yes | ||
| 33 | PABPC4 | 10510 2418 2548 | 2319 | 1.021 | 0.5010 | Yes | ||
| 34 | RBM3 | 5366 2625 | 2322 | 1.019 | 0.5057 | Yes | ||
| 35 | HRSP12 | 9123 9122 4870 9638 | 2353 | 1.001 | 0.5088 | Yes | ||
| 36 | HBP1 | 2048 2162 21086 | 2363 | 0.999 | 0.5130 | Yes | ||
| 37 | TOP1 | 5790 5789 10210 | 2370 | 0.996 | 0.5173 | Yes | ||
| 38 | SH3KBP1 | 2631 12206 7189 | 2421 | 0.965 | 0.5191 | Yes | ||
| 39 | HTF9C | 4886 1678 | 2513 | 0.889 | 0.5184 | Yes | ||
| 40 | FHOD1 | 18772 | 2527 | 0.882 | 0.5218 | Yes | ||
| 41 | RXRB | 23285 9768 | 2694 | 0.745 | 0.5163 | No | ||
| 42 | NR1D1 | 4650 20258 949 | 2741 | 0.711 | 0.5172 | No | ||
| 43 | HYAL2 | 4891 | 2797 | 0.662 | 0.5173 | No | ||
| 44 | UBE1 | 24370 2551 | 2800 | 0.660 | 0.5203 | No | ||
| 45 | LMNB1 | 23549 1943 | 2946 | 0.560 | 0.5151 | No | ||
| 46 | TIMM9 | 19686 | 2960 | 0.553 | 0.5169 | No | ||
| 47 | MYST3 | 18651 | 2998 | 0.523 | 0.5174 | No | ||
| 48 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 3317 | 0.365 | 0.5019 | No | ||
| 49 | CLCN2 | 22637 | 3321 | 0.364 | 0.5034 | No | ||
| 50 | GPD1 | 22361 | 3424 | 0.323 | 0.4994 | No | ||
| 51 | HOXB7 | 666 9110 | 3439 | 0.317 | 0.5001 | No | ||
| 52 | USP2 | 19480 3052 3043 | 3489 | 0.303 | 0.4989 | No | ||
| 53 | KRTCAP2 | 15541 | 3491 | 0.303 | 0.5003 | No | ||
| 54 | WEE1 | 18127 | 3500 | 0.297 | 0.5012 | No | ||
| 55 | MAX | 21034 | 4023 | 0.188 | 0.4738 | No | ||
| 56 | RPIA | 17124 | 4112 | 0.175 | 0.4699 | No | ||
| 57 | CACNA1D | 4464 8675 | 4198 | 0.163 | 0.4660 | No | ||
| 58 | HOXC13 | 22344 | 4212 | 0.162 | 0.4661 | No | ||
| 59 | BARHL1 | 14634 | 4355 | 0.146 | 0.4591 | No | ||
| 60 | CHST11 | 19921 | 4386 | 0.142 | 0.4581 | No | ||
| 61 | GTPBP1 | 4814 2255 | 4540 | 0.128 | 0.4504 | No | ||
| 62 | EIF3S1 | 905 8114 | 4709 | 0.115 | 0.4418 | No | ||
| 63 | XRN2 | 2936 6302 2905 | 4981 | 0.098 | 0.4276 | No | ||
| 64 | GPS1 | 1392 9943 5550 9944 | 5064 | 0.093 | 0.4236 | No | ||
| 65 | CUGBP1 | 2805 8819 4576 2924 | 5128 | 0.089 | 0.4206 | No | ||
| 66 | SEMA7A | 19426 9803 | 5281 | 0.082 | 0.4127 | No | ||
| 67 | AMPD2 | 1931 15197 | 5322 | 0.079 | 0.4109 | No | ||
| 68 | NPM1 | 1196 | 5708 | 0.064 | 0.3903 | No | ||
| 69 | GRK6 | 6449 | 5863 | 0.060 | 0.3823 | No | ||
| 70 | SEPT3 | 6275 10775 | 5970 | 0.057 | 0.3768 | No | ||
| 71 | RTN4RL2 | 14544 | 6027 | 0.055 | 0.3740 | No | ||
| 72 | SCFD2 | 5607 | 6053 | 0.054 | 0.3729 | No | ||
| 73 | PCDHA10 | 8792 | 6065 | 0.054 | 0.3726 | No | ||
| 74 | STMN4 | 21977 | 6074 | 0.054 | 0.3724 | No | ||
| 75 | VPS16 | 2927 14845 2825 | 6203 | 0.051 | 0.3657 | No | ||
| 76 | TPM2 | 2367 2437 | 6219 | 0.050 | 0.3651 | No | ||
| 77 | NDUFS1 | 13940 | 6414 | 0.046 | 0.3548 | No | ||
| 78 | SC5DL | 6165 | 6563 | 0.043 | 0.3470 | No | ||
| 79 | B3GALT2 | 6498 | 6658 | 0.041 | 0.3421 | No | ||
| 80 | FGF11 | 20383 | 6871 | 0.037 | 0.3308 | No | ||
| 81 | LHX9 | 9276 | 6873 | 0.037 | 0.3309 | No | ||
| 82 | MTHFD1 | 2132 21238 | 7035 | 0.034 | 0.3223 | No | ||
| 83 | KIAA0427 | 11228 | 7075 | 0.033 | 0.3203 | No | ||
| 84 | NEUROD6 | 17138 | 7093 | 0.033 | 0.3196 | No | ||
| 85 | EN2 | 16898 | 7357 | 0.028 | 0.3055 | No | ||
| 86 | BAI2 | 2541 2379 16067 | 7497 | 0.027 | 0.2980 | No | ||
| 87 | FLT3 | 16288 | 7531 | 0.026 | 0.2964 | No | ||
| 88 | PABPC1 | 5219 9522 9523 23572 | 7921 | 0.021 | 0.2754 | No | ||
| 89 | PRKCE | 9575 | 7969 | 0.020 | 0.2729 | No | ||
| 90 | PLA2G4A | 13809 | 7985 | 0.020 | 0.2722 | No | ||
| 91 | NXPH1 | 17524 | 8202 | 0.017 | 0.2606 | No | ||
| 92 | RPL22 | 15979 | 8609 | 0.011 | 0.2386 | No | ||
| 93 | RUNX2 | 4480 8700 | 8624 | 0.011 | 0.2379 | No | ||
| 94 | LAP3 | 12442 | 8640 | 0.011 | 0.2371 | No | ||
| 95 | ZZZ3 | 15389 8445 4253 8444 1761 | 8722 | 0.010 | 0.2328 | No | ||
| 96 | ANGPT2 | 14885 3783 18923 | 8731 | 0.010 | 0.2324 | No | ||
| 97 | NEUROD2 | 20262 | 8856 | 0.008 | 0.2257 | No | ||
| 98 | POGK | 7739 | 8859 | 0.008 | 0.2256 | No | ||
| 99 | HRH3 | 14317 | 9008 | 0.006 | 0.2176 | No | ||
| 100 | TESK2 | 10504 | 9170 | 0.004 | 0.2089 | No | ||
| 101 | NKX2-2 | 5176 | 9246 | 0.003 | 0.2049 | No | ||
| 102 | MNT | 20784 | 9250 | 0.003 | 0.2047 | No | ||
| 103 | POU3F1 | 16093 | 9252 | 0.003 | 0.2047 | No | ||
| 104 | TIAL1 | 996 5753 | 9368 | 0.002 | 0.1985 | No | ||
| 105 | NUP153 | 21474 | 9748 | -0.003 | 0.1779 | No | ||
| 106 | SNCAIP | 12589 | 9831 | -0.005 | 0.1735 | No | ||
| 107 | HPCA | 9118 | 9835 | -0.005 | 0.1733 | No | ||
| 108 | ABCA1 | 15881 | 9838 | -0.005 | 0.1733 | No | ||
| 109 | ADCY3 | 21329 894 | 9900 | -0.005 | 0.1700 | No | ||
| 110 | SEMA3F | 18999 3097 | 10061 | -0.007 | 0.1613 | No | ||
| 111 | SLC12A5 | 14731 | 10118 | -0.008 | 0.1583 | No | ||
| 112 | NRAS | 5191 | 10314 | -0.010 | 0.1478 | No | ||
| 113 | DLC1 | 11931 3855 | 10504 | -0.013 | 0.1376 | No | ||
| 114 | COL2A1 | 22145 4542 | 10712 | -0.016 | 0.1265 | No | ||
| 115 | ADAM10 | 4336 | 10985 | -0.020 | 0.1118 | No | ||
| 116 | HS3ST3A1 | 20844 | 11181 | -0.022 | 0.1013 | No | ||
| 117 | SOX12 | 5478 | 11461 | -0.027 | 0.0863 | No | ||
| 118 | DNMT3A | 2167 21330 | 11522 | -0.028 | 0.0832 | No | ||
| 119 | SGTB | 21571 | 11595 | -0.029 | 0.0794 | No | ||
| 120 | HHIP | 4849 | 11746 | -0.032 | 0.0715 | No | ||
| 121 | PLAGL1 | 10372 3414 | 11764 | -0.032 | 0.0707 | No | ||
| 122 | ADAMTS17 | 598 | 11793 | -0.032 | 0.0693 | No | ||
| 123 | DLX1 | 14988 | 12048 | -0.037 | 0.0557 | No | ||
| 124 | GPC3 | 24151 | 12243 | -0.041 | 0.0454 | No | ||
| 125 | CTBP2 | 17591 1137 | 12277 | -0.042 | 0.0438 | No | ||
| 126 | CTSF | 23772 | 12419 | -0.045 | 0.0364 | No | ||
| 127 | DNAJB9 | 6565 | 12450 | -0.046 | 0.0350 | No | ||
| 128 | ALX4 | 4379 | 12457 | -0.046 | 0.0348 | No | ||
| 129 | SLC17A2 | 10163 5744 | 12567 | -0.049 | 0.0292 | No | ||
| 130 | CHRM1 | 23943 | 12677 | -0.052 | 0.0235 | No | ||
| 131 | PLAG1 | 7148 15949 12161 | 12683 | -0.053 | 0.0235 | No | ||
| 132 | IPO13 | 15784 | 12920 | -0.058 | 0.0110 | No | ||
| 133 | NRXN1 | 1581 1575 22875 | 12942 | -0.059 | 0.0101 | No | ||
| 134 | APOA5 | 19469 | 13222 | -0.069 | -0.0047 | No | ||
| 135 | ADPRHL1 | 18930 3863 | 13353 | -0.074 | -0.0114 | No | ||
| 136 | LIN28 | 15723 | 13371 | -0.075 | -0.0120 | No | ||
| 137 | FKBP10 | 20667 | 13416 | -0.077 | -0.0140 | No | ||
| 138 | GRIN2A | 22668 | 13577 | -0.084 | -0.0223 | No | ||
| 139 | KCNK5 | 4946 9212 | 13640 | -0.088 | -0.0252 | No | ||
| 140 | NR5A1 | 14597 | 13796 | -0.097 | -0.0332 | No | ||
| 141 | RNF128 | 24236 2641 | 13825 | -0.099 | -0.0342 | No | ||
| 142 | SOCS5 | 1619 23141 | 14043 | -0.114 | -0.0455 | No | ||
| 143 | ENPP6 | 6816 11583 | 14286 | -0.134 | -0.0580 | No | ||
| 144 | U2AF2 | 5813 5812 5814 | 14350 | -0.140 | -0.0607 | No | ||
| 145 | TLE3 | 5767 19416 | 14526 | -0.159 | -0.0695 | No | ||
| 146 | MCM8 | 14834 | 14591 | -0.168 | -0.0722 | No | ||
| 147 | RAD9A | 913 3674 910 23958 | 14697 | -0.184 | -0.0770 | No | ||
| 148 | DUSP4 | 18632 3820 | 14743 | -0.190 | -0.0785 | No | ||
| 149 | BMP2K | 3657 16786 | 14982 | -0.233 | -0.0904 | No | ||
| 150 | SFXN2 | 3707 8255 3678 13588 | 15107 | -0.259 | -0.0959 | No | ||
| 151 | B4GALT2 | 11990 | 15204 | -0.281 | -0.0997 | No | ||
| 152 | RPL13A | 10226 79 | 15291 | -0.306 | -0.1030 | No | ||
| 153 | ETV1 | 4688 8925 8924 | 15380 | -0.333 | -0.1062 | No | ||
| 154 | RCL1 | 3728 3743 23894 | 15515 | -0.386 | -0.1116 | No | ||
| 155 | PDK2 | 20285 | 15551 | -0.404 | -0.1116 | No | ||
| 156 | JPH1 | 13994 | 15590 | -0.417 | -0.1117 | No | ||
| 157 | SLC38A5 | 24388 | 15749 | -0.489 | -0.1180 | No | ||
| 158 | SCYL1 | 23986 | 15759 | -0.492 | -0.1162 | No | ||
| 159 | ATP6V0B | 15786 | 15840 | -0.542 | -0.1180 | No | ||
| 160 | HMGN2 | 9095 | 16119 | -0.766 | -0.1294 | No | ||
| 161 | PFN1 | 9555 | 16293 | -0.962 | -0.1343 | No | ||
| 162 | AKAP10 | 20411 7145 | 16408 | -1.038 | -0.1356 | No | ||
| 163 | ANXA6 | 20449 | 16750 | -1.447 | -0.1473 | No | ||
| 164 | GADD45B | 19935 | 16872 | -1.620 | -0.1462 | No | ||
| 165 | CDC14A | 15184 | 16894 | -1.662 | -0.1396 | No | ||
| 166 | TEF | 2287 22414 2192 | 16957 | -1.731 | -0.1348 | No | ||
| 167 | PFKFB3 | 2748 | 17127 | -1.997 | -0.1346 | No | ||
| 168 | ARF6 | 21252 | 17192 | -2.126 | -0.1281 | No | ||
| 169 | GADD45G | 21637 | 17483 | -2.597 | -0.1316 | No | ||
| 170 | GNA13 | 20617 | 17564 | -2.763 | -0.1229 | No | ||
| 171 | PSEN2 | 13736 | 17674 | -2.999 | -0.1148 | No | ||
| 172 | MXD4 | 9346 | 17793 | -3.348 | -0.1054 | No | ||
| 173 | ZHX2 | 11842 | 18038 | -4.076 | -0.0995 | No | ||
| 174 | WBP2 | 20146 | 18237 | -5.059 | -0.0865 | No | ||
| 175 | ARRDC3 | 4191 | 18287 | -5.303 | -0.0642 | No | ||
| 176 | AP1GBP1 | 20725 | 18350 | -5.626 | -0.0411 | No | ||
| 177 | STAT3 | 5525 9906 | 18377 | -5.818 | -0.0152 | No | ||
| 178 | GJA1 | 20023 | 18396 | -5.981 | 0.0119 | No |

