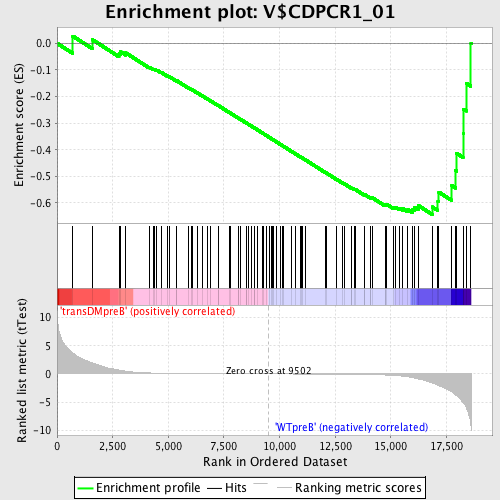
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMpreB_versus_WTpreB.phenotype_transDMpreB_versus_WTpreB.cls #transDMpreB_versus_WTpreB |
| Phenotype | phenotype_transDMpreB_versus_WTpreB.cls#transDMpreB_versus_WTpreB |
| Upregulated in class | WTpreB |
| GeneSet | V$CDPCR1_01 |
| Enrichment Score (ES) | -0.6431579 |
| Normalized Enrichment Score (NES) | -1.5583874 |
| Nominal p-value | 0.002016129 |
| FDR q-value | 0.07695113 |
| FWER p-Value | 0.482 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GART | 22543 1754 | 706 | 3.684 | 0.0270 | No | ||
| 2 | GRB7 | 20673 | 1581 | 1.934 | 0.0140 | No | ||
| 3 | CIB3 | 18840 | 2783 | 0.671 | -0.0390 | No | ||
| 4 | CREM | 2005 1965 23496 1951 | 2851 | 0.624 | -0.0316 | No | ||
| 5 | MYST2 | 20283 | 3088 | 0.470 | -0.0360 | No | ||
| 6 | IL15 | 18826 3801 | 4142 | 0.172 | -0.0898 | No | ||
| 7 | KLF15 | 17359 | 4332 | 0.148 | -0.0974 | No | ||
| 8 | KCNS1 | 14358 | 4384 | 0.142 | -0.0976 | No | ||
| 9 | PFN2 | 15337 1818 | 4480 | 0.134 | -0.1004 | No | ||
| 10 | BHLHB3 | 16935 | 4672 | 0.117 | -0.1086 | No | ||
| 11 | ST7 | 1054 17519 | 4963 | 0.099 | -0.1225 | No | ||
| 12 | MGAT5B | 20584 11197 | 5060 | 0.093 | -0.1261 | No | ||
| 13 | PAX8 | 2791 14671 | 5375 | 0.077 | -0.1417 | No | ||
| 14 | KCNA1 | 9202 | 5382 | 0.077 | -0.1406 | No | ||
| 15 | MAG | 17880 | 5920 | 0.058 | -0.1686 | No | ||
| 16 | RTN4RL2 | 14544 | 6027 | 0.055 | -0.1733 | No | ||
| 17 | EEF1A2 | 8880 14309 | 6081 | 0.054 | -0.1752 | No | ||
| 18 | CYP26B1 | 17093 | 6292 | 0.048 | -0.1857 | No | ||
| 19 | CSMD3 | 10735 7938 | 6532 | 0.044 | -0.1978 | No | ||
| 20 | HOXA10 | 17145 989 | 6757 | 0.039 | -0.2092 | No | ||
| 21 | NR3C2 | 18562 13798 | 6899 | 0.037 | -0.2162 | No | ||
| 22 | PLXNA3 | 24299 | 7256 | 0.030 | -0.2349 | No | ||
| 23 | HOXD10 | 14983 | 7258 | 0.030 | -0.2344 | No | ||
| 24 | VAX1 | 23636 | 7762 | 0.023 | -0.2611 | No | ||
| 25 | DIO2 | 21014 2159 | 7803 | 0.022 | -0.2629 | No | ||
| 26 | SERPING1 | 14546 | 8161 | 0.017 | -0.2819 | No | ||
| 27 | KDELR2 | 16634 12428 | 8174 | 0.017 | -0.2822 | No | ||
| 28 | DBC1 | 7147 | 8260 | 0.016 | -0.2865 | No | ||
| 29 | ITGA7 | 19835 | 8492 | 0.013 | -0.2988 | No | ||
| 30 | NTNG2 | 2731 9330 | 8513 | 0.013 | -0.2996 | No | ||
| 31 | GATA4 | 21792 4755 | 8608 | 0.011 | -0.3045 | No | ||
| 32 | NOG | 13663 | 8741 | 0.009 | -0.3114 | No | ||
| 33 | FGF12 | 1723 22621 | 8850 | 0.008 | -0.3171 | No | ||
| 34 | INSM2 | 19821 | 8854 | 0.008 | -0.3172 | No | ||
| 35 | BTBD3 | 10457 2975 6007 6006 | 9028 | 0.006 | -0.3264 | No | ||
| 36 | NKX2-2 | 5176 | 9246 | 0.003 | -0.3380 | No | ||
| 37 | KCND1 | 4940 | 9277 | 0.003 | -0.3396 | No | ||
| 38 | NFIB | 15855 | 9395 | 0.001 | -0.3459 | No | ||
| 39 | PPT2 | 1617 12034 | 9550 | -0.001 | -0.3542 | No | ||
| 40 | ERG | 1686 8915 | 9637 | -0.002 | -0.3588 | No | ||
| 41 | NRXN3 | 2153 5196 | 9658 | -0.002 | -0.3598 | No | ||
| 42 | TCF7L2 | 10048 5646 | 9716 | -0.003 | -0.3629 | No | ||
| 43 | DPP10 | 13850 | 9881 | -0.005 | -0.3716 | No | ||
| 44 | STAT5B | 20222 | 10037 | -0.007 | -0.3799 | No | ||
| 45 | SLC12A5 | 14731 | 10118 | -0.008 | -0.3840 | No | ||
| 46 | SFTPC | 21759 | 10160 | -0.009 | -0.3861 | No | ||
| 47 | GRIK4 | 19157 | 10555 | -0.014 | -0.4071 | No | ||
| 48 | COL2A1 | 22145 4542 | 10712 | -0.016 | -0.4152 | No | ||
| 49 | RPH3A | 13628 | 10948 | -0.019 | -0.4276 | No | ||
| 50 | ADAM10 | 4336 | 10985 | -0.020 | -0.4292 | No | ||
| 51 | DGKB | 5722 | 11035 | -0.020 | -0.4315 | No | ||
| 52 | HOXB2 | 8346 | 11145 | -0.022 | -0.4370 | No | ||
| 53 | NTRK1 | 15299 | 12045 | -0.037 | -0.4848 | No | ||
| 54 | FCHSD1 | 6661 11435 | 12102 | -0.038 | -0.4872 | No | ||
| 55 | SON | 5473 1657 1684 | 12559 | -0.049 | -0.5109 | No | ||
| 56 | TBX6 | 1993 18075 | 12826 | -0.056 | -0.5243 | No | ||
| 57 | MAOB | 24182 2556 | 12918 | -0.058 | -0.5282 | No | ||
| 58 | RFX4 | 7721 3410 | 13240 | -0.070 | -0.5443 | No | ||
| 59 | HOXC5 | 22340 | 13250 | -0.070 | -0.5435 | No | ||
| 60 | LIN28 | 15723 | 13371 | -0.075 | -0.5487 | No | ||
| 61 | STMN2 | 15638 | 13406 | -0.076 | -0.5492 | No | ||
| 62 | NHLH2 | 9462 | 13811 | -0.098 | -0.5692 | No | ||
| 63 | NOL4 | 2028 11432 | 13828 | -0.099 | -0.5684 | No | ||
| 64 | CUL2 | 23630 | 14078 | -0.116 | -0.5797 | No | ||
| 65 | CPNE6 | 22010 | 14095 | -0.118 | -0.5785 | No | ||
| 66 | INSRR | 15557 | 14172 | -0.124 | -0.5804 | No | ||
| 67 | THRA | 1447 10171 1406 | 14741 | -0.189 | -0.6077 | No | ||
| 68 | DUSP4 | 18632 3820 | 14743 | -0.190 | -0.6044 | No | ||
| 69 | HOXC6 | 22339 2302 | 14803 | -0.199 | -0.6041 | No | ||
| 70 | TBX3 | 16723 | 15109 | -0.260 | -0.6160 | No | ||
| 71 | PIGN | 13865 | 15187 | -0.277 | -0.6152 | No | ||
| 72 | ETV1 | 4688 8925 8924 | 15380 | -0.333 | -0.6197 | No | ||
| 73 | BRUNOL4 | 2010 4229 | 15546 | -0.403 | -0.6215 | No | ||
| 74 | NR2F6 | 4651 8876 18848 8912 | 15765 | -0.496 | -0.6245 | No | ||
| 75 | AP3M2 | 11 | 15986 | -0.648 | -0.6249 | Yes | ||
| 76 | SERPINI1 | 340 | 16072 | -0.724 | -0.6167 | Yes | ||
| 77 | STAG2 | 5521 | 16249 | -0.911 | -0.6101 | Yes | ||
| 78 | TLE4 | 23720 3697 | 16862 | -1.610 | -0.6147 | Yes | ||
| 79 | PDCD10 | 15314 | 17116 | -1.984 | -0.5933 | Yes | ||
| 80 | EXT1 | 22297 | 17159 | -2.061 | -0.5592 | Yes | ||
| 81 | MEIS1 | 20524 | 17740 | -3.157 | -0.5348 | Yes | ||
| 82 | MBNL1 | 1921 15582 | 17912 | -3.707 | -0.4785 | Yes | ||
| 83 | HOXB4 | 20686 | 17960 | -3.831 | -0.4134 | Yes | ||
| 84 | EMP1 | 17260 | 18249 | -5.131 | -0.3383 | Yes | ||
| 85 | TRIB2 | 21102 | 18257 | -5.165 | -0.2474 | Yes | ||
| 86 | LYPLA3 | 18482 | 18390 | -5.904 | -0.1503 | Yes | ||
| 87 | EGR1 | 23598 | 18600 | -9.195 | 0.0009 | Yes |

