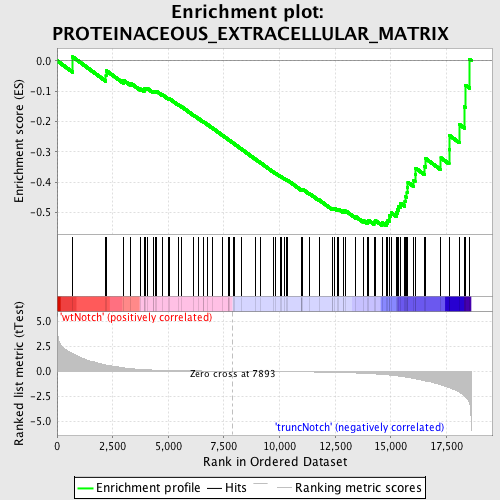
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_truncNotch_versus_wtNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch.phenotype_truncNotch_versus_wtNotch.cls #wtNotch_versus_truncNotch_repos |
| Phenotype | phenotype_truncNotch_versus_wtNotch.cls#wtNotch_versus_truncNotch_repos |
| Upregulated in class | truncNotch |
| GeneSet | PROTEINACEOUS_EXTRACELLULAR_MATRIX |
| Enrichment Score (ES) | -0.54435235 |
| Normalized Enrichment Score (NES) | -1.4856828 |
| Nominal p-value | 0.00309119 |
| FDR q-value | 0.5768066 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SGCB | 1240162 6370711 | 700 | 1.747 | 0.0128 | No | ||
| 2 | MMP11 | 1980619 | 2196 | 0.624 | -0.0498 | No | ||
| 3 | ERBB2IP | 580253 1090672 | 2205 | 0.621 | -0.0322 | No | ||
| 4 | COL5A1 | 2230050 | 2989 | 0.345 | -0.0645 | No | ||
| 5 | LAMC1 | 4730025 | 3306 | 0.264 | -0.0739 | No | ||
| 6 | LAMA3 | 1190131 5290239 | 3756 | 0.180 | -0.0929 | No | ||
| 7 | SMC3 | 870546 1400411 5700039 6450577 | 3910 | 0.158 | -0.0966 | No | ||
| 8 | COL13A1 | 4120079 | 3921 | 0.157 | -0.0926 | No | ||
| 9 | LTBP4 | 50164 1780301 | 3954 | 0.152 | -0.0899 | No | ||
| 10 | MEPE | 1690484 | 4042 | 0.142 | -0.0905 | No | ||
| 11 | SNTB2 | 2630014 5390035 | 4311 | 0.118 | -0.1015 | No | ||
| 12 | HAPLN1 | 580398 | 4347 | 0.114 | -0.1001 | No | ||
| 13 | TNXB | 630592 3830020 5360497 7000673 | 4400 | 0.109 | -0.0998 | No | ||
| 14 | COL4A4 | 1050541 | 4482 | 0.103 | -0.1012 | No | ||
| 15 | OPTC | 4850044 | 4716 | 0.087 | -0.1112 | No | ||
| 16 | SPG7 | 1940041 6220113 | 5011 | 0.072 | -0.1250 | No | ||
| 17 | FMOD | 540577 | 5053 | 0.070 | -0.1251 | No | ||
| 18 | COL5A2 | 780040 | 5457 | 0.054 | -0.1453 | No | ||
| 19 | SNTB1 | 730450 2680632 | 5601 | 0.049 | -0.1516 | No | ||
| 20 | COL4A5 | 1340161 | 6141 | 0.033 | -0.1798 | No | ||
| 21 | THBS4 | 3710348 | 6371 | 0.028 | -0.1913 | No | ||
| 22 | DSPP | 5890403 | 6585 | 0.023 | -0.2021 | No | ||
| 23 | COL4A3 | 5910075 | 6599 | 0.023 | -0.2022 | No | ||
| 24 | EFEMP1 | 5860324 | 6780 | 0.019 | -0.2113 | No | ||
| 25 | COL8A1 | 4590050 | 7002 | 0.015 | -0.2228 | No | ||
| 26 | SNTG1 | 6770373 | 7445 | 0.007 | -0.2464 | No | ||
| 27 | ADAMTS9 | 2760079 | 7710 | 0.003 | -0.2606 | No | ||
| 28 | IMPG1 | 5570025 | 7716 | 0.003 | -0.2608 | No | ||
| 29 | SGCE | 6840075 | 7728 | 0.003 | -0.2613 | No | ||
| 30 | CAV3 | 1770519 | 7944 | -0.001 | -0.2729 | No | ||
| 31 | ADAMTS5 | 5890592 | 7971 | -0.001 | -0.2742 | No | ||
| 32 | IMPG2 | 4730110 | 8277 | -0.006 | -0.2905 | No | ||
| 33 | DMP1 | 4760398 | 8898 | -0.016 | -0.3235 | No | ||
| 34 | LUM | 5420079 | 9141 | -0.021 | -0.3360 | No | ||
| 35 | SSPN | 4670091 | 9738 | -0.031 | -0.3673 | No | ||
| 36 | TINAG | 1570609 6510215 | 9837 | -0.033 | -0.3716 | No | ||
| 37 | TFPI2 | 3870324 | 10019 | -0.037 | -0.3803 | No | ||
| 38 | COL11A1 | 2100300 2810136 4200706 4810524 | 10088 | -0.038 | -0.3829 | No | ||
| 39 | LAMA2 | 7040020 | 10206 | -0.040 | -0.3880 | No | ||
| 40 | COL5A3 | 1940471 | 10328 | -0.043 | -0.3933 | No | ||
| 41 | MUC2 | 1570025 | 10357 | -0.043 | -0.3936 | No | ||
| 42 | CD248 | 2480605 | 10964 | -0.058 | -0.4246 | No | ||
| 43 | MATN3 | 2570292 | 10996 | -0.059 | -0.4246 | No | ||
| 44 | VCAN | 3290017 5910053 6940014 | 11000 | -0.059 | -0.4231 | No | ||
| 45 | COL9A2 | 940670 | 11046 | -0.060 | -0.4237 | No | ||
| 46 | SGCA | 4560164 6380577 | 11346 | -0.068 | -0.4379 | No | ||
| 47 | COL9A3 | 4050541 | 11771 | -0.082 | -0.4584 | No | ||
| 48 | DMD | 1740041 3990332 | 12380 | -0.107 | -0.4881 | No | ||
| 49 | LTBP2 | 4850039 | 12393 | -0.108 | -0.4856 | No | ||
| 50 | COL9A1 | 4570369 7100446 | 12464 | -0.111 | -0.4862 | No | ||
| 51 | COL15A1 | 6840113 | 12604 | -0.119 | -0.4903 | No | ||
| 52 | FBLN5 | 6550010 | 12651 | -0.121 | -0.4892 | No | ||
| 53 | DST | 430026 1090035 2340577 3170068 3870112 4780519 6400167 6450358 7040347 | 12867 | -0.133 | -0.4970 | No | ||
| 54 | CTGF | 4540577 | 12887 | -0.135 | -0.4941 | No | ||
| 55 | POSTN | 450411 6040451 | 12977 | -0.141 | -0.4948 | No | ||
| 56 | SGCG | 4920161 | 13426 | -0.174 | -0.5140 | No | ||
| 57 | NID2 | 2320059 | 13752 | -0.205 | -0.5256 | No | ||
| 58 | CHAD | 3060100 | 13972 | -0.229 | -0.5308 | No | ||
| 59 | COL10A1 | 2360307 | 13988 | -0.230 | -0.5249 | No | ||
| 60 | FBLN2 | 730601 3440215 | 14245 | -0.263 | -0.5311 | No | ||
| 61 | COL18A1 | 610301 4570338 | 14307 | -0.270 | -0.5266 | No | ||
| 62 | COL1A2 | 380364 | 14606 | -0.319 | -0.5334 | No | ||
| 63 | COL7A1 | 1230168 2190072 | 14810 | -0.353 | -0.5341 | Yes | ||
| 64 | COL3A1 | 6420273 6620044 | 14868 | -0.363 | -0.5267 | Yes | ||
| 65 | COL6A3 | 2640717 4070064 5390717 | 14929 | -0.376 | -0.5190 | Yes | ||
| 66 | ECM2 | 1470561 | 14933 | -0.377 | -0.5083 | Yes | ||
| 67 | SNTG2 | 2710441 | 15007 | -0.394 | -0.5008 | Yes | ||
| 68 | MMP10 | 7100707 | 15243 | -0.445 | -0.5006 | Yes | ||
| 69 | FBN1 | 3170181 | 15304 | -0.461 | -0.4905 | Yes | ||
| 70 | MATN1 | 2850075 | 15349 | -0.476 | -0.4791 | Yes | ||
| 71 | EFEMP2 | 5550086 | 15438 | -0.500 | -0.4693 | Yes | ||
| 72 | PRELP | 5900390 | 15634 | -0.571 | -0.4633 | Yes | ||
| 73 | SGCD | 2940685 5080753 | 15670 | -0.584 | -0.4483 | Yes | ||
| 74 | FBN2 | 50435 | 15726 | -0.603 | -0.4338 | Yes | ||
| 75 | TGFB1 | 1940162 | 15749 | -0.610 | -0.4173 | Yes | ||
| 76 | MAGEE1 | 1660138 | 15769 | -0.616 | -0.4005 | Yes | ||
| 77 | APLP1 | 2680154 | 16032 | -0.728 | -0.3935 | Yes | ||
| 78 | COMP | 3800273 | 16091 | -0.755 | -0.3748 | Yes | ||
| 79 | ECM1 | 940068 5390309 | 16113 | -0.765 | -0.3537 | Yes | ||
| 80 | LAMB2 | 4920184 | 16520 | -0.948 | -0.3482 | Yes | ||
| 81 | COL16A1 | 1780520 | 16565 | -0.973 | -0.3224 | Yes | ||
| 82 | CHI3L1 | 1170039 | 17254 | -1.360 | -0.3201 | Yes | ||
| 83 | COL4A2 | 2350619 | 17617 | -1.618 | -0.2928 | Yes | ||
| 84 | MGP | 6900736 | 17642 | -1.640 | -0.2465 | Yes | ||
| 85 | AGRN | 6770152 | 18074 | -2.084 | -0.2094 | Yes | ||
| 86 | LAMA4 | 3170647 | 18310 | -2.437 | -0.1515 | Yes | ||
| 87 | FBLN1 | 2480059 5670735 | 18359 | -2.537 | -0.0806 | Yes | ||
| 88 | DGCR6 | 5890129 | 18546 | -3.258 | 0.0038 | Yes |

