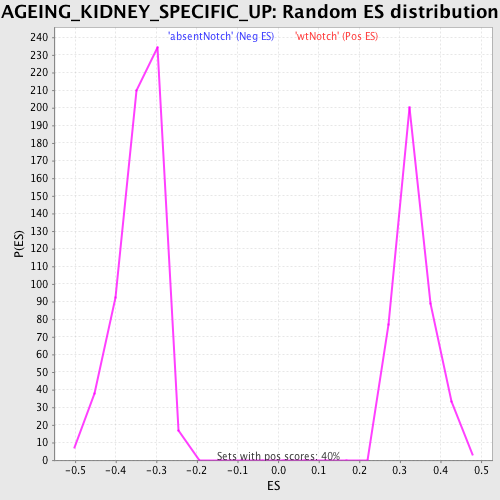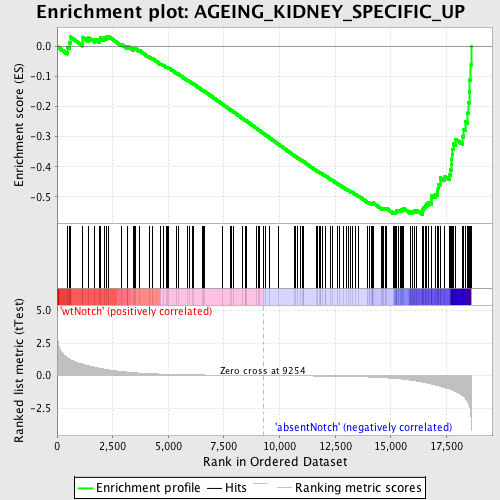
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_wtNotch.phenotype_absentNotch_versus_wtNotch.cls #wtNotch_versus_absentNotch |
| Phenotype | phenotype_absentNotch_versus_wtNotch.cls#wtNotch_versus_absentNotch |
| Upregulated in class | absentNotch |
| GeneSet | AGEING_KIDNEY_SPECIFIC_UP |
| Enrichment Score (ES) | -0.5589398 |
| Normalized Enrichment Score (NES) | -1.6350408 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.1841449 |
| FWER p-Value | 0.945 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | DEGS1 | 5910021 | 456 | 1.362 | -0.0019 | No | ||
| 2 | FEZ2 | 5910181 | 553 | 1.260 | 0.0139 | No | ||
| 3 | SOCS3 | 5550563 | 593 | 1.220 | 0.0322 | No | ||
| 4 | ITPR3 | 4010632 | 1153 | 0.873 | 0.0165 | No | ||
| 5 | GGTA1 | 1570292 | 1154 | 0.873 | 0.0311 | No | ||
| 6 | MRAS | 2030524 | 1402 | 0.754 | 0.0303 | No | ||
| 7 | ASNS | 110368 7100687 | 1696 | 0.630 | 0.0249 | No | ||
| 8 | B3GAT3 | 1580438 | 1907 | 0.562 | 0.0230 | No | ||
| 9 | MGLL | 2030446 4210520 | 1931 | 0.556 | 0.0310 | No | ||
| 10 | SMPD2 | 2370133 | 2117 | 0.491 | 0.0292 | No | ||
| 11 | COL15A1 | 6840113 | 2210 | 0.468 | 0.0320 | No | ||
| 12 | FBN1 | 3170181 | 2295 | 0.444 | 0.0349 | No | ||
| 13 | MMP11 | 1980619 | 2879 | 0.309 | 0.0085 | No | ||
| 14 | COL27A1 | 1740390 | 3152 | 0.261 | -0.0019 | No | ||
| 15 | DPP3 | 2680725 3780670 | 3184 | 0.255 | 0.0007 | No | ||
| 16 | GPNMB | 1780138 2340176 3390403 4560156 | 3431 | 0.221 | -0.0089 | No | ||
| 17 | PHLDA3 | 510398 | 3474 | 0.216 | -0.0076 | No | ||
| 18 | MFGE8 | 3440373 | 3536 | 0.208 | -0.0074 | No | ||
| 19 | RAB34 | 770411 | 3713 | 0.188 | -0.0138 | No | ||
| 20 | HOMER3 | 3830279 | 4144 | 0.148 | -0.0346 | No | ||
| 21 | STS | 630632 | 4297 | 0.137 | -0.0405 | No | ||
| 22 | TPM1 | 130673 | 4665 | 0.113 | -0.0585 | No | ||
| 23 | FGG | 4610717 | 4777 | 0.107 | -0.0627 | No | ||
| 24 | FN1 | 1170601 2970647 6220288 6940037 | 4923 | 0.099 | -0.0689 | No | ||
| 25 | VSNL1 | 3120017 | 4941 | 0.098 | -0.0682 | No | ||
| 26 | GABRP | 1940091 6650102 | 5007 | 0.095 | -0.0701 | No | ||
| 27 | ANXA1 | 2320053 | 5372 | 0.079 | -0.0885 | No | ||
| 28 | C1S | 840184 6840114 | 5442 | 0.077 | -0.0910 | No | ||
| 29 | MMP7 | 1780497 | 5852 | 0.062 | -0.1120 | No | ||
| 30 | MSR1 | 940538 | 5935 | 0.060 | -0.1155 | No | ||
| 31 | TSPAN1 | 2450519 | 6079 | 0.056 | -0.1223 | No | ||
| 32 | HGF | 3360593 | 6118 | 0.055 | -0.1234 | No | ||
| 33 | UBD | 5570632 | 6533 | 0.045 | -0.1451 | No | ||
| 34 | LIX1 | 1580324 3360520 4280195 4850079 | 6563 | 0.044 | -0.1459 | No | ||
| 35 | COL6A3 | 2640717 4070064 5390717 | 6627 | 0.043 | -0.1486 | No | ||
| 36 | RPN1 | 1240113 | 7420 | 0.027 | -0.1910 | No | ||
| 37 | SPARCL1 | 1990348 | 7809 | 0.021 | -0.2117 | No | ||
| 38 | BHLHB3 | 3450438 | 7842 | 0.020 | -0.2131 | No | ||
| 39 | PKIB | 780014 | 7941 | 0.019 | -0.2180 | No | ||
| 40 | LUM | 5420079 | 8343 | 0.013 | -0.2395 | No | ||
| 41 | COL14A1 | 1570398 | 8450 | 0.011 | -0.2451 | No | ||
| 42 | GPR56 | 1190494 | 8484 | 0.011 | -0.2467 | No | ||
| 43 | NOV | 2850746 | 8497 | 0.010 | -0.2472 | No | ||
| 44 | MOXD1 | 2450301 | 8980 | 0.004 | -0.2732 | No | ||
| 45 | ADRA2A | 5340520 | 9041 | 0.003 | -0.2764 | No | ||
| 46 | CLDN3 | 2570524 | 9092 | 0.002 | -0.2790 | No | ||
| 47 | PAM | 5290528 | 9287 | -0.000 | -0.2895 | No | ||
| 48 | VEGFC | 5910494 | 9361 | -0.001 | -0.2935 | No | ||
| 49 | CTHRC1 | 4150020 | 9546 | -0.004 | -0.3033 | No | ||
| 50 | CDH11 | 1230133 2100180 4610110 | 9946 | -0.010 | -0.3248 | No | ||
| 51 | CDC42EP1 | 6550279 6370215 4060450 4590066 | 10678 | -0.020 | -0.3640 | No | ||
| 52 | CDO1 | 2480279 | 10692 | -0.021 | -0.3643 | No | ||
| 53 | DCDC2 | 7000338 | 10819 | -0.023 | -0.3708 | No | ||
| 54 | CXCL2 | 610398 | 10919 | -0.024 | -0.3757 | No | ||
| 55 | GPR61 | 6980035 | 11044 | -0.026 | -0.3820 | No | ||
| 56 | PTRF | 2260204 | 11078 | -0.027 | -0.3833 | No | ||
| 57 | HEXB | 5860692 | 11092 | -0.027 | -0.3836 | No | ||
| 58 | PGM2L1 | 2350035 | 11653 | -0.037 | -0.4133 | No | ||
| 59 | LHFP | 3440368 | 11702 | -0.038 | -0.4152 | No | ||
| 60 | TNFRSF11B | 4610446 | 11784 | -0.040 | -0.4190 | No | ||
| 61 | ITGB6 | 2810068 4570332 | 11817 | -0.040 | -0.4200 | No | ||
| 62 | SYTL2 | 580097 5390576 6770603 | 11848 | -0.041 | -0.4210 | No | ||
| 63 | CKAP4 | 1050056 | 11914 | -0.042 | -0.4238 | No | ||
| 64 | PDGFC | 4200133 | 12060 | -0.045 | -0.4309 | No | ||
| 65 | C1R | 2340025 3290152 4850452 | 12068 | -0.045 | -0.4305 | No | ||
| 66 | RBMS3 | 6110722 | 12291 | -0.050 | -0.4417 | No | ||
| 67 | SLC1A3 | 1500070 | 12390 | -0.053 | -0.4461 | No | ||
| 68 | SPON1 | 1660301 | 12621 | -0.059 | -0.4576 | No | ||
| 69 | PROM1 | 3170537 | 12698 | -0.061 | -0.4607 | No | ||
| 70 | CHRNA3 | 6760100 | 12850 | -0.065 | -0.4677 | No | ||
| 71 | SETD7 | 130450 6420739 | 13019 | -0.071 | -0.4757 | No | ||
| 72 | OCLN | 2570671 | 13077 | -0.072 | -0.4775 | No | ||
| 73 | ISLR | 110273 6450368 | 13202 | -0.077 | -0.4830 | No | ||
| 74 | BNC2 | 4810603 | 13271 | -0.079 | -0.4853 | No | ||
| 75 | NNMT | 450471 | 13415 | -0.085 | -0.4916 | No | ||
| 76 | EBF1 | 3130397 | 13551 | -0.091 | -0.4974 | No | ||
| 77 | RPL35A | 2370706 | 13956 | -0.110 | -0.5174 | No | ||
| 78 | CCL2 | 4760019 | 14048 | -0.116 | -0.5204 | No | ||
| 79 | COL3A1 | 6420273 6620044 | 14149 | -0.122 | -0.5238 | No | ||
| 80 | COL1A1 | 730020 | 14172 | -0.124 | -0.5229 | No | ||
| 81 | TNC | 670053 1780039 1980020 3060411 4780091 6860433 | 14184 | -0.125 | -0.5214 | No | ||
| 82 | ITM2C | 780647 | 14199 | -0.125 | -0.5201 | No | ||
| 83 | ARHGAP1 | 2810010 5270064 | 14597 | -0.156 | -0.5390 | No | ||
| 84 | RARRES1 | 770139 5860390 | 14631 | -0.160 | -0.5381 | No | ||
| 85 | FCAMR | 50132 | 14684 | -0.164 | -0.5382 | No | ||
| 86 | FHL1 | 60278 | 14759 | -0.170 | -0.5393 | No | ||
| 87 | C3 | 1740372 | 14801 | -0.174 | -0.5386 | No | ||
| 88 | STAB1 | 5390707 | 15135 | -0.213 | -0.5531 | Yes | ||
| 89 | NR2F1 | 4120528 360402 | 15166 | -0.219 | -0.5511 | Yes | ||
| 90 | RGS1 | 4060347 4540181 | 15231 | -0.228 | -0.5507 | Yes | ||
| 91 | GGTLA1 | 3140273 4920168 | 15267 | -0.233 | -0.5487 | Yes | ||
| 92 | CLDN1 | 5670746 | 15270 | -0.233 | -0.5449 | Yes | ||
| 93 | LOXL1 | 3520537 | 15350 | -0.245 | -0.5451 | Yes | ||
| 94 | PLK2 | 6450152 | 15429 | -0.259 | -0.5450 | Yes | ||
| 95 | COL5A1 | 2230050 | 15484 | -0.268 | -0.5435 | Yes | ||
| 96 | RPS27L | 50577 | 15530 | -0.277 | -0.5413 | Yes | ||
| 97 | TRIM47 | 5080717 | 15574 | -0.285 | -0.5389 | Yes | ||
| 98 | TIMP1 | 1010326 | 15882 | -0.351 | -0.5496 | Yes | ||
| 99 | COL1A2 | 380364 | 15969 | -0.373 | -0.5480 | Yes | ||
| 100 | CALU | 2680403 3520056 4610348 5720176 6130121 | 16074 | -0.400 | -0.5470 | Yes | ||
| 101 | SERPING1 | 5550440 | 16170 | -0.424 | -0.5450 | Yes | ||
| 102 | ANXA2 | 3140402 | 16428 | -0.502 | -0.5506 | Yes | ||
| 103 | AATF | 5220575 | 16436 | -0.505 | -0.5425 | Yes | ||
| 104 | SERPINE2 | 2810390 | 16471 | -0.515 | -0.5357 | Yes | ||
| 105 | CXCL14 | 840114 6450324 | 16540 | -0.542 | -0.5304 | Yes | ||
| 106 | RPS25 | 520139 | 16593 | -0.559 | -0.5239 | Yes | ||
| 107 | PIGR | 6520441 | 16693 | -0.594 | -0.5193 | Yes | ||
| 108 | TFPI | 1400288 5550008 | 16840 | -0.652 | -0.5163 | Yes | ||
| 109 | ABCC3 | 4050242 | 16843 | -0.652 | -0.5055 | Yes | ||
| 110 | RPL35 | 6940070 | 16847 | -0.655 | -0.4948 | Yes | ||
| 111 | CFB | 730577 | 17015 | -0.729 | -0.4916 | Yes | ||
| 112 | AEBP1 | 6450707 | 17102 | -0.769 | -0.4835 | Yes | ||
| 113 | PHCA | 5890215 6130113 | 17118 | -0.776 | -0.4713 | Yes | ||
| 114 | TPBG | 630161 | 17156 | -0.794 | -0.4601 | Yes | ||
| 115 | NFKBIZ | 2970450 | 17212 | -0.823 | -0.4493 | Yes | ||
| 116 | LTF | 6770364 | 17213 | -0.824 | -0.4355 | Yes | ||
| 117 | POLE4 | 6860097 | 17428 | -0.944 | -0.4313 | Yes | ||
| 118 | RBP1 | 1690072 | 17652 | -1.039 | -0.4261 | Yes | ||
| 119 | GGA2 | 520451 1240082 2650332 | 17688 | -1.065 | -0.4102 | Yes | ||
| 120 | BICC1 | 4670647 | 17709 | -1.083 | -0.3932 | Yes | ||
| 121 | C1QB | 5910292 | 17725 | -1.091 | -0.3758 | Yes | ||
| 122 | SSR4 | 1340619 4150132 | 17749 | -1.106 | -0.3586 | Yes | ||
| 123 | NFIX | 2450152 | 17781 | -1.136 | -0.3413 | Yes | ||
| 124 | PLOD3 | 5080152 | 17821 | -1.164 | -0.3240 | Yes | ||
| 125 | RAD54L | 2970041 | 17899 | -1.222 | -0.3077 | Yes | ||
| 126 | P4HB | 6110056 | 18215 | -1.537 | -0.2991 | Yes | ||
| 127 | C1QC | 5700131 | 18283 | -1.626 | -0.2756 | Yes | ||
| 128 | ANTXR1 | 1990593 2850056 | 18364 | -1.788 | -0.2501 | Yes | ||
| 129 | SCN5A | 4070017 | 18462 | -2.053 | -0.2210 | Yes | ||
| 130 | PRICKLE1 | 450671 | 18486 | -2.159 | -0.1862 | Yes | ||
| 131 | CYFIP1 | 5690082 | 18521 | -2.260 | -0.1504 | Yes | ||
| 132 | AXL | 1770095 | 18548 | -2.436 | -0.1111 | Yes | ||
| 133 | VCAM1 | 2900450 | 18602 | -3.170 | -0.0610 | Yes | ||
| 134 | SLC40A1 | 2370286 | 18611 | -3.698 | 0.0003 | Yes |