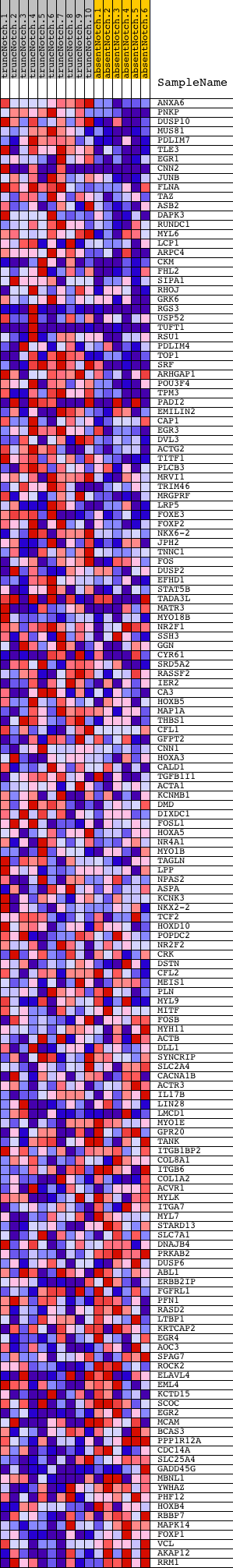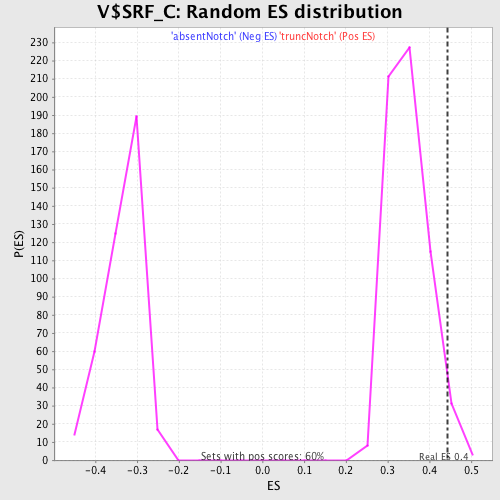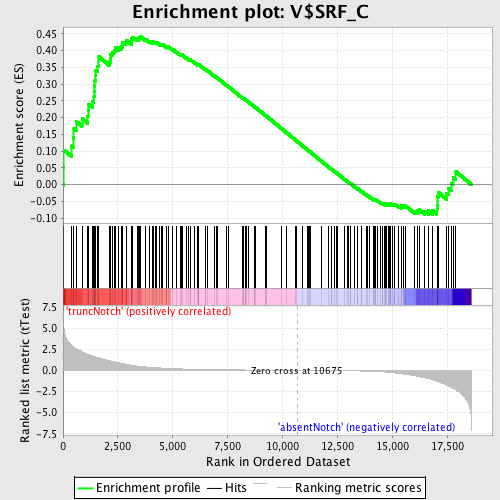
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #truncNotch_versus_absentNotch |
| Phenotype | phenotype_absentNotch_versus_truncNotch.cls#truncNotch_versus_absentNotch |
| Upregulated in class | truncNotch |
| GeneSet | V$SRF_C |
| Enrichment Score (ES) | 0.44304457 |
| Normalized Enrichment Score (NES) | 1.2704382 |
| Nominal p-value | 0.03697479 |
| FDR q-value | 0.9364467 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ANXA6 | 2190014 | 30 | 5.135 | 0.0521 | Yes | ||
| 2 | PNKP | 4610685 5720605 | 38 | 4.908 | 0.1032 | Yes | ||
| 3 | DUSP10 | 2850673 3360064 | 386 | 2.963 | 0.1154 | Yes | ||
| 4 | MUS81 | 6660184 | 461 | 2.781 | 0.1405 | Yes | ||
| 5 | PDLIM7 | 4780136 | 493 | 2.731 | 0.1674 | Yes | ||
| 6 | TLE3 | 580040 4730121 | 609 | 2.546 | 0.1879 | Yes | ||
| 7 | EGR1 | 4610347 | 861 | 2.187 | 0.1972 | Yes | ||
| 8 | CNN2 | 2230433 5270446 | 1121 | 1.897 | 0.2030 | Yes | ||
| 9 | JUNB | 4230048 | 1138 | 1.886 | 0.2219 | Yes | ||
| 10 | FLNA | 5390193 | 1171 | 1.860 | 0.2396 | Yes | ||
| 11 | TAZ | 7100193 | 1342 | 1.703 | 0.2483 | Yes | ||
| 12 | ASB2 | 4760168 | 1405 | 1.640 | 0.2621 | Yes | ||
| 13 | DAPK3 | 4590121 | 1414 | 1.635 | 0.2788 | Yes | ||
| 14 | RUNDC1 | 2450037 | 1439 | 1.612 | 0.2943 | Yes | ||
| 15 | MYL6 | 60563 6100152 | 1448 | 1.608 | 0.3108 | Yes | ||
| 16 | LCP1 | 4280397 | 1483 | 1.583 | 0.3255 | Yes | ||
| 17 | ARPC4 | 2900056 | 1492 | 1.577 | 0.3416 | Yes | ||
| 18 | CKM | 1450524 | 1583 | 1.513 | 0.3525 | Yes | ||
| 19 | FHL2 | 1400008 | 1612 | 1.500 | 0.3667 | Yes | ||
| 20 | SIPA1 | 5220687 | 1613 | 1.500 | 0.3824 | Yes | ||
| 21 | RHOJ | 6660176 | 2106 | 1.139 | 0.3677 | Yes | ||
| 22 | GRK6 | 1500053 | 2146 | 1.105 | 0.3772 | Yes | ||
| 23 | RGS3 | 60670 540736 1340180 1500369 3390735 4010131 4610402 6380114 | 2154 | 1.101 | 0.3883 | Yes | ||
| 24 | USP52 | 3710215 6380133 | 2231 | 1.057 | 0.3953 | Yes | ||
| 25 | TUFT1 | 540047 2370010 | 2320 | 1.011 | 0.4011 | Yes | ||
| 26 | RSU1 | 4850372 | 2377 | 0.982 | 0.4083 | Yes | ||
| 27 | PDLIM4 | 3990358 | 2518 | 0.898 | 0.4102 | Yes | ||
| 28 | TOP1 | 770471 4060632 6650324 | 2641 | 0.831 | 0.4123 | Yes | ||
| 29 | SRF | 70139 2320039 | 2690 | 0.804 | 0.4181 | Yes | ||
| 30 | ARHGAP1 | 2810010 5270064 | 2714 | 0.793 | 0.4251 | Yes | ||
| 31 | POU3F4 | 870274 | 2868 | 0.707 | 0.4243 | Yes | ||
| 32 | TPM3 | 670600 3170296 5670167 6650471 | 2876 | 0.702 | 0.4312 | Yes | ||
| 33 | PADI2 | 2940092 6420136 | 3101 | 0.607 | 0.4254 | Yes | ||
| 34 | EMILIN2 | 70253 | 3105 | 0.606 | 0.4316 | Yes | ||
| 35 | CAP1 | 2650278 | 3127 | 0.600 | 0.4368 | Yes | ||
| 36 | EGR3 | 6940128 | 3176 | 0.582 | 0.4403 | Yes | ||
| 37 | DVL3 | 360156 5390075 | 3371 | 0.515 | 0.4351 | Yes | ||
| 38 | ACTG2 | 4780180 | 3456 | 0.485 | 0.4357 | Yes | ||
| 39 | TITF1 | 3390554 4540292 | 3485 | 0.473 | 0.4391 | Yes | ||
| 40 | PLCB3 | 4670402 | 3504 | 0.469 | 0.4430 | Yes | ||
| 41 | MRVI1 | 4810338 4850601 5900441 | 3747 | 0.405 | 0.4342 | No | ||
| 42 | TRIM46 | 3520609 4590010 | 3932 | 0.367 | 0.4280 | No | ||
| 43 | MRGPRF | 4780746 | 4064 | 0.341 | 0.4245 | No | ||
| 44 | LRP5 | 2100397 3170484 | 4102 | 0.333 | 0.4260 | No | ||
| 45 | FOXE3 | 6900102 | 4221 | 0.314 | 0.4229 | No | ||
| 46 | FOXP2 | 3520561 4150372 4760524 | 4235 | 0.312 | 0.4255 | No | ||
| 47 | NKX6-2 | 2450332 3120333 | 4408 | 0.283 | 0.4191 | No | ||
| 48 | JPH2 | 940041 5860026 | 4473 | 0.272 | 0.4185 | No | ||
| 49 | TNNC1 | 1990575 | 4519 | 0.265 | 0.4188 | No | ||
| 50 | FOS | 1850315 | 4728 | 0.238 | 0.4101 | No | ||
| 51 | DUSP2 | 2370184 | 4730 | 0.237 | 0.4125 | No | ||
| 52 | EFHD1 | 4070358 | 4784 | 0.232 | 0.4120 | No | ||
| 53 | STAT5B | 6200026 | 4979 | 0.210 | 0.4037 | No | ||
| 54 | TADA3L | 4210070 5720088 6660333 | 5178 | 0.185 | 0.3949 | No | ||
| 55 | MATR3 | 1940170 5340278 | 5369 | 0.169 | 0.3864 | No | ||
| 56 | MYO18B | 7100670 | 5401 | 0.167 | 0.3865 | No | ||
| 57 | NR2F1 | 4120528 360402 | 5424 | 0.165 | 0.3870 | No | ||
| 58 | SSH3 | 6290110 6650497 | 5616 | 0.151 | 0.3783 | No | ||
| 59 | GGN | 1410528 4210082 5290170 | 5736 | 0.142 | 0.3733 | No | ||
| 60 | CYR61 | 1240408 5290026 4120452 6550008 | 5791 | 0.138 | 0.3718 | No | ||
| 61 | SRD5A2 | 6840433 | 5814 | 0.136 | 0.3721 | No | ||
| 62 | RASSF2 | 2480079 5720546 | 5991 | 0.124 | 0.3638 | No | ||
| 63 | IER2 | 2030008 | 6120 | 0.117 | 0.3581 | No | ||
| 64 | CA3 | 870687 5890390 | 6154 | 0.116 | 0.3575 | No | ||
| 65 | HOXB5 | 2900538 | 6179 | 0.114 | 0.3574 | No | ||
| 66 | MAP1A | 4920576 | 6187 | 0.114 | 0.3582 | No | ||
| 67 | THBS1 | 4560494 430288 | 6484 | 0.098 | 0.3432 | No | ||
| 68 | CFL1 | 2340735 | 6499 | 0.097 | 0.3435 | No | ||
| 69 | GFPT2 | 2370129 | 6584 | 0.094 | 0.3399 | No | ||
| 70 | CNN1 | 510707 | 6889 | 0.082 | 0.3243 | No | ||
| 71 | HOXA3 | 3610632 6020358 | 6985 | 0.078 | 0.3200 | No | ||
| 72 | CALD1 | 1770129 1940397 | 7033 | 0.077 | 0.3182 | No | ||
| 73 | TGFB1I1 | 2060288 6550450 | 7443 | 0.063 | 0.2968 | No | ||
| 74 | ACTA1 | 840538 | 7533 | 0.061 | 0.2926 | No | ||
| 75 | KCNMB1 | 4760139 | 8160 | 0.047 | 0.2591 | No | ||
| 76 | DMD | 1740041 3990332 | 8227 | 0.045 | 0.2560 | No | ||
| 77 | DIXDC1 | 6980435 | 8235 | 0.045 | 0.2561 | No | ||
| 78 | FOSL1 | 430021 | 8299 | 0.043 | 0.2532 | No | ||
| 79 | HOXA5 | 6840026 | 8304 | 0.043 | 0.2534 | No | ||
| 80 | NR4A1 | 6290161 | 8346 | 0.042 | 0.2516 | No | ||
| 81 | MYO1B | 770372 | 8443 | 0.041 | 0.2468 | No | ||
| 82 | TAGLN | 6660280 | 8722 | 0.034 | 0.2321 | No | ||
| 83 | LPP | 1660541 | 8726 | 0.034 | 0.2323 | No | ||
| 84 | NPAS2 | 770670 | 8767 | 0.033 | 0.2305 | No | ||
| 85 | ASPA | 3290400 | 9230 | 0.024 | 0.2058 | No | ||
| 86 | KCNK3 | 2690372 | 9234 | 0.024 | 0.2058 | No | ||
| 87 | NKX2-2 | 4150731 | 9251 | 0.024 | 0.2052 | No | ||
| 88 | TCF2 | 870338 5050632 | 9956 | 0.012 | 0.1672 | No | ||
| 89 | HOXD10 | 6100039 | 10161 | 0.009 | 0.1563 | No | ||
| 90 | POPDC2 | 2030056 | 10590 | 0.001 | 0.1331 | No | ||
| 91 | NR2F2 | 3170609 3310577 | 10615 | 0.001 | 0.1318 | No | ||
| 92 | CRK | 1230162 4780128 | 10922 | -0.004 | 0.1153 | No | ||
| 93 | DSTN | 3140592 | 11144 | -0.008 | 0.1034 | No | ||
| 94 | CFL2 | 380239 6200368 | 11203 | -0.009 | 0.1003 | No | ||
| 95 | MEIS1 | 1400575 | 11236 | -0.010 | 0.0987 | No | ||
| 96 | PLN | 3940132 | 11256 | -0.010 | 0.0978 | No | ||
| 97 | MYL9 | 4210750 7050138 | 11759 | -0.019 | 0.0708 | No | ||
| 98 | MITF | 380056 | 12093 | -0.026 | 0.0530 | No | ||
| 99 | FOSB | 1940142 | 12252 | -0.030 | 0.0448 | No | ||
| 100 | MYH11 | 7100273 | 12254 | -0.030 | 0.0450 | No | ||
| 101 | ACTB | 1660736 2030204 2060215 5050047 | 12348 | -0.032 | 0.0403 | No | ||
| 102 | DLL1 | 1770377 | 12441 | -0.035 | 0.0357 | No | ||
| 103 | SYNCRIP | 1690195 3140113 4670279 | 12501 | -0.036 | 0.0329 | No | ||
| 104 | SLC2A4 | 540441 | 12819 | -0.045 | 0.0162 | No | ||
| 105 | CACNA1B | 1740010 5360593 | 12941 | -0.049 | 0.0102 | No | ||
| 106 | ACTR3 | 1400497 | 13022 | -0.052 | 0.0064 | No | ||
| 107 | IL17B | 6940711 | 13088 | -0.054 | 0.0034 | No | ||
| 108 | LIN28 | 6590672 | 13101 | -0.054 | 0.0033 | No | ||
| 109 | LMCD1 | 1740484 2190097 | 13299 | -0.062 | -0.0067 | No | ||
| 110 | MYO1E | 3140504 | 13408 | -0.066 | -0.0119 | No | ||
| 111 | GPR20 | 6620601 | 13437 | -0.067 | -0.0127 | No | ||
| 112 | TANK | 1090020 2060458 2370400 | 13601 | -0.076 | -0.0207 | No | ||
| 113 | ITGB1BP2 | 6200129 | 13607 | -0.076 | -0.0202 | No | ||
| 114 | COL8A1 | 4590050 | 13810 | -0.089 | -0.0302 | No | ||
| 115 | ITGB6 | 2810068 4570332 | 13881 | -0.094 | -0.0330 | No | ||
| 116 | COL1A2 | 380364 | 13981 | -0.102 | -0.0373 | No | ||
| 117 | ACVR1 | 6840671 | 14141 | -0.115 | -0.0447 | No | ||
| 118 | MYLK | 4010600 7000364 | 14162 | -0.118 | -0.0445 | No | ||
| 119 | ITGA7 | 1230739 | 14183 | -0.120 | -0.0444 | No | ||
| 120 | MYL7 | 2480541 | 14228 | -0.124 | -0.0455 | No | ||
| 121 | STARD13 | 540722 | 14252 | -0.128 | -0.0454 | No | ||
| 122 | SLC7A1 | 2450193 2710241 | 14336 | -0.137 | -0.0484 | No | ||
| 123 | DNAJB4 | 380195 2760372 3940735 | 14483 | -0.157 | -0.0547 | No | ||
| 124 | PRKAB2 | 6980300 | 14557 | -0.167 | -0.0569 | No | ||
| 125 | DUSP6 | 5910286 7100070 | 14573 | -0.170 | -0.0559 | No | ||
| 126 | ABL1 | 1050593 2030050 4010114 | 14663 | -0.186 | -0.0588 | No | ||
| 127 | ERBB2IP | 580253 1090672 | 14707 | -0.196 | -0.0591 | No | ||
| 128 | FGFRL1 | 6370047 | 14709 | -0.197 | -0.0571 | No | ||
| 129 | PFN1 | 6130132 | 14735 | -0.201 | -0.0563 | No | ||
| 130 | RASD2 | 3360010 | 14823 | -0.218 | -0.0588 | No | ||
| 131 | LTBP1 | 780035 2100746 2760402 3780551 6550184 6860193 | 14894 | -0.234 | -0.0601 | No | ||
| 132 | KRTCAP2 | 5220040 | 14908 | -0.238 | -0.0583 | No | ||
| 133 | EGR4 | 3120750 | 14913 | -0.239 | -0.0560 | No | ||
| 134 | AOC3 | 6840129 | 15020 | -0.266 | -0.0590 | No | ||
| 135 | SPAG7 | 4560524 | 15099 | -0.290 | -0.0602 | No | ||
| 136 | ROCK2 | 1780035 5720025 | 15118 | -0.295 | -0.0580 | No | ||
| 137 | ELAVL4 | 50735 3360086 5220167 | 15306 | -0.352 | -0.0645 | No | ||
| 138 | EML4 | 5700373 | 15414 | -0.390 | -0.0662 | No | ||
| 139 | KCTD15 | 2810735 | 15425 | -0.395 | -0.0626 | No | ||
| 140 | SCOC | 610048 2230053 | 15517 | -0.427 | -0.0631 | No | ||
| 141 | EGR2 | 3800403 | 15588 | -0.452 | -0.0621 | No | ||
| 142 | MCAM | 5130270 6620577 | 16019 | -0.635 | -0.0788 | No | ||
| 143 | BCAS3 | 4050647 | 16135 | -0.687 | -0.0778 | No | ||
| 144 | PPP1R12A | 4070066 4590162 | 16223 | -0.748 | -0.0747 | No | ||
| 145 | CDC14A | 4050132 | 16473 | -0.881 | -0.0789 | No | ||
| 146 | SLC25A4 | 2360519 | 16645 | -0.981 | -0.0779 | No | ||
| 147 | GADD45G | 2510142 | 16850 | -1.097 | -0.0775 | No | ||
| 148 | MBNL1 | 2640762 7100048 | 17040 | -1.263 | -0.0745 | No | ||
| 149 | YWHAZ | 1230717 | 17058 | -1.283 | -0.0620 | No | ||
| 150 | PHF12 | 870400 3130687 | 17070 | -1.295 | -0.0490 | No | ||
| 151 | HOXB4 | 540131 | 17071 | -1.296 | -0.0355 | No | ||
| 152 | RBBP7 | 430113 450450 2370309 | 17109 | -1.334 | -0.0235 | No | ||
| 153 | MAPK14 | 5290731 | 17476 | -1.736 | -0.0252 | No | ||
| 154 | FOXP1 | 6510433 | 17556 | -1.822 | -0.0104 | No | ||
| 155 | VCL | 4120487 | 17715 | -2.044 | 0.0025 | No | ||
| 156 | AKAP12 | 1450739 | 17771 | -2.116 | 0.0217 | No | ||
| 157 | RRM1 | 4150433 | 17904 | -2.297 | 0.0386 | No |