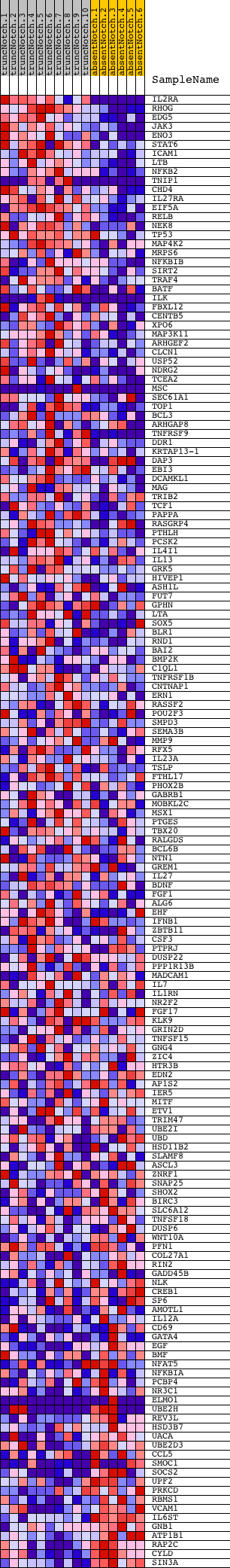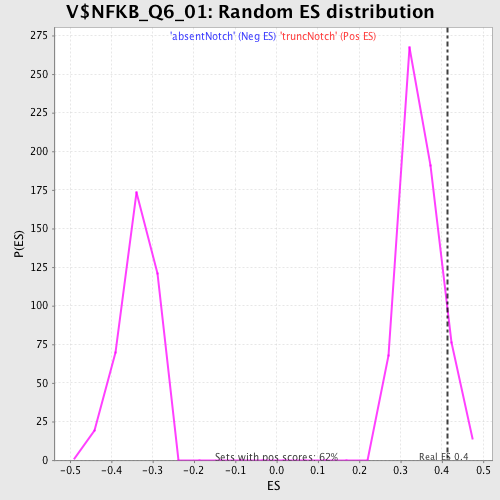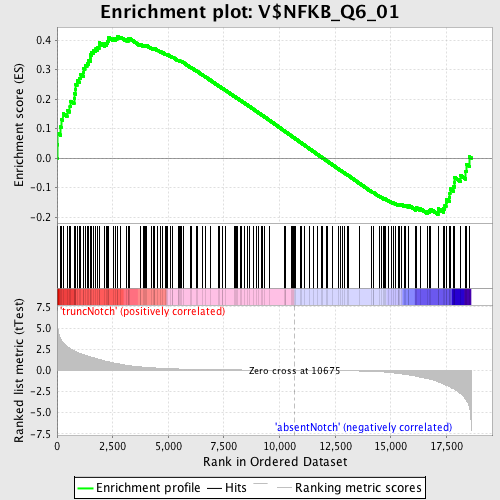
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #truncNotch_versus_absentNotch |
| Phenotype | phenotype_absentNotch_versus_truncNotch.cls#truncNotch_versus_absentNotch |
| Upregulated in class | truncNotch |
| GeneSet | V$NFKB_Q6_01 |
| Enrichment Score (ES) | 0.41339087 |
| Normalized Enrichment Score (NES) | 1.1919659 |
| Nominal p-value | 0.08766234 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | IL2RA | 6620450 | 8 | 6.074 | 0.0453 | Yes | ||
| 2 | RHOG | 6760575 | 27 | 5.267 | 0.0841 | Yes | ||
| 3 | EDG5 | 2640300 | 150 | 3.760 | 0.1058 | Yes | ||
| 4 | JAK3 | 70347 3290008 | 183 | 3.588 | 0.1311 | Yes | ||
| 5 | ENO3 | 5270136 | 264 | 3.305 | 0.1517 | Yes | ||
| 6 | STAT6 | 1190010 5720019 | 449 | 2.814 | 0.1629 | Yes | ||
| 7 | ICAM1 | 6980138 | 577 | 2.609 | 0.1757 | Yes | ||
| 8 | LTB | 3990672 | 608 | 2.549 | 0.1933 | Yes | ||
| 9 | NFKB2 | 2320670 | 763 | 2.310 | 0.2024 | Yes | ||
| 10 | TNIP1 | 2510279 5900041 6510435 | 786 | 2.282 | 0.2184 | Yes | ||
| 11 | CHD4 | 5420059 6130338 6380717 | 827 | 2.234 | 0.2331 | Yes | ||
| 12 | IL27RA | 940093 | 840 | 2.211 | 0.2491 | Yes | ||
| 13 | EIF5A | 1500059 5290022 6370026 | 896 | 2.143 | 0.2622 | Yes | ||
| 14 | RELB | 1400048 | 1011 | 2.010 | 0.2712 | Yes | ||
| 15 | NEK8 | 5700180 | 1053 | 1.962 | 0.2838 | Yes | ||
| 16 | TP53 | 6130707 | 1179 | 1.852 | 0.2910 | Yes | ||
| 17 | MAP4K2 | 4200037 | 1189 | 1.841 | 0.3044 | Yes | ||
| 18 | MRPS6 | 870164 | 1254 | 1.772 | 0.3142 | Yes | ||
| 19 | NFKBIB | 2260167 | 1345 | 1.698 | 0.3222 | Yes | ||
| 20 | SIRT2 | 630494 | 1423 | 1.629 | 0.3303 | Yes | ||
| 21 | TRAF4 | 3060041 4920528 6980286 | 1494 | 1.575 | 0.3384 | Yes | ||
| 22 | BATF | 6760390 | 1501 | 1.569 | 0.3499 | Yes | ||
| 23 | ILK | 870601 4480180 | 1538 | 1.541 | 0.3595 | Yes | ||
| 24 | FBXL12 | 2230040 3520008 | 1640 | 1.487 | 0.3653 | Yes | ||
| 25 | CENTB5 | 5690128 | 1743 | 1.416 | 0.3704 | Yes | ||
| 26 | XPO6 | 3940019 | 1827 | 1.328 | 0.3759 | Yes | ||
| 27 | MAP3K11 | 7000039 | 1887 | 1.287 | 0.3824 | Yes | ||
| 28 | ARHGEF2 | 3360577 | 1901 | 1.274 | 0.3913 | Yes | ||
| 29 | CLCN1 | 840161 | 2133 | 1.119 | 0.3872 | Yes | ||
| 30 | USP52 | 3710215 6380133 | 2231 | 1.057 | 0.3899 | Yes | ||
| 31 | NDRG2 | 450403 | 2275 | 1.032 | 0.3954 | Yes | ||
| 32 | TCEA2 | 6370546 | 2302 | 1.019 | 0.4017 | Yes | ||
| 33 | MSC | 5080139 | 2321 | 1.011 | 0.4083 | Yes | ||
| 34 | SEC61A1 | 3390437 | 2526 | 0.890 | 0.4040 | Yes | ||
| 35 | TOP1 | 770471 4060632 6650324 | 2641 | 0.831 | 0.4041 | Yes | ||
| 36 | BCL3 | 3990440 | 2693 | 0.803 | 0.4073 | Yes | ||
| 37 | ARHGAP8 | 380333 5390487 | 2694 | 0.803 | 0.4134 | Yes | ||
| 38 | TNFRSF9 | 2510400 6650484 | 2867 | 0.707 | 0.4094 | No | ||
| 39 | DDR1 | 2060044 5220180 | 3102 | 0.607 | 0.4013 | No | ||
| 40 | KRTAP13-1 | 2600242 | 3190 | 0.578 | 0.4009 | No | ||
| 41 | DAP3 | 1660528 2120039 5420593 | 3209 | 0.571 | 0.4043 | No | ||
| 42 | EBI3 | 5340300 | 3263 | 0.551 | 0.4055 | No | ||
| 43 | DCAMKL1 | 540095 2690092 | 3727 | 0.411 | 0.3835 | No | ||
| 44 | MAG | 2370037 | 3749 | 0.405 | 0.3855 | No | ||
| 45 | TRIB2 | 4120605 | 3902 | 0.373 | 0.3800 | No | ||
| 46 | TCF1 | 5390022 | 3919 | 0.371 | 0.3820 | No | ||
| 47 | PAPPA | 4230463 | 3954 | 0.361 | 0.3828 | No | ||
| 48 | RASGRP4 | 540102 | 4022 | 0.348 | 0.3818 | No | ||
| 49 | PTHLH | 5290739 | 4241 | 0.311 | 0.3724 | No | ||
| 50 | PCSK2 | 360017 430528 5900619 | 4347 | 0.292 | 0.3689 | No | ||
| 51 | IL4I1 | 3190161 | 4360 | 0.291 | 0.3704 | No | ||
| 52 | IL13 | 7040021 | 4372 | 0.287 | 0.3720 | No | ||
| 53 | GRK5 | 1940348 4670053 | 4533 | 0.262 | 0.3653 | No | ||
| 54 | HIVEP1 | 3850632 | 4659 | 0.246 | 0.3604 | No | ||
| 55 | ASH1L | 2510402 5360687 6650685 | 4742 | 0.236 | 0.3577 | No | ||
| 56 | FUT7 | 2900736 | 4873 | 0.222 | 0.3524 | No | ||
| 57 | GPHN | 3290195 3840121 | 4918 | 0.216 | 0.3516 | No | ||
| 58 | LTA | 1740088 | 4973 | 0.210 | 0.3502 | No | ||
| 59 | SOX5 | 2370576 2900167 3190128 5050528 | 5110 | 0.192 | 0.3443 | No | ||
| 60 | BLR1 | 1500300 5420017 | 5185 | 0.185 | 0.3417 | No | ||
| 61 | RND1 | 5080300 | 5199 | 0.183 | 0.3424 | No | ||
| 62 | BAI2 | 380632 630113 6650170 | 5433 | 0.165 | 0.3310 | No | ||
| 63 | BMP2K | 60541 5570504 | 5451 | 0.163 | 0.3313 | No | ||
| 64 | C1QL1 | 2030465 | 5521 | 0.158 | 0.3288 | No | ||
| 65 | TNFRSF1B | 3990035 5860372 | 5536 | 0.157 | 0.3292 | No | ||
| 66 | CNTNAP1 | 6650132 | 5593 | 0.153 | 0.3273 | No | ||
| 67 | ERN1 | 2360403 | 5686 | 0.146 | 0.3234 | No | ||
| 68 | RASSF2 | 2480079 5720546 | 5991 | 0.124 | 0.3079 | No | ||
| 69 | POU2F3 | 2030601 | 6032 | 0.122 | 0.3066 | No | ||
| 70 | SMPD3 | 520164 | 6251 | 0.110 | 0.2956 | No | ||
| 71 | SEMA3B | 5570563 | 6314 | 0.107 | 0.2931 | No | ||
| 72 | MMP9 | 580338 | 6541 | 0.096 | 0.2816 | No | ||
| 73 | RFX5 | 5420128 | 6661 | 0.091 | 0.2758 | No | ||
| 74 | IL23A | 6290044 | 6873 | 0.082 | 0.2650 | No | ||
| 75 | TSLP | 730408 1990500 | 7238 | 0.070 | 0.2458 | No | ||
| 76 | FTHL17 | 6550193 | 7290 | 0.069 | 0.2435 | No | ||
| 77 | PHOX2B | 5270075 | 7421 | 0.064 | 0.2370 | No | ||
| 78 | GABRB1 | 6760193 | 7422 | 0.064 | 0.2375 | No | ||
| 79 | MOBKL2C | 2640088 3830017 | 7577 | 0.060 | 0.2296 | No | ||
| 80 | MSX1 | 2650309 | 7991 | 0.050 | 0.2076 | No | ||
| 81 | PTGES | 1690075 | 8001 | 0.050 | 0.2075 | No | ||
| 82 | TBX20 | 2230279 4920102 6980020 | 8075 | 0.049 | 0.2039 | No | ||
| 83 | RALGDS | 430463 | 8126 | 0.047 | 0.2015 | No | ||
| 84 | BCL6B | 60047 | 8251 | 0.044 | 0.1951 | No | ||
| 85 | NTN1 | 5700600 | 8283 | 0.044 | 0.1938 | No | ||
| 86 | GREM1 | 3940180 | 8441 | 0.041 | 0.1856 | No | ||
| 87 | IL27 | 1990324 | 8541 | 0.039 | 0.1805 | No | ||
| 88 | BDNF | 2940128 3520368 | 8647 | 0.036 | 0.1751 | No | ||
| 89 | FGF1 | 4780435 5670601 | 8848 | 0.032 | 0.1645 | No | ||
| 90 | ALG6 | 4060131 | 8947 | 0.030 | 0.1594 | No | ||
| 91 | EHF | 4560088 | 9070 | 0.027 | 0.1530 | No | ||
| 92 | IFNB1 | 1400142 | 9172 | 0.025 | 0.1477 | No | ||
| 93 | ZBTB11 | 6520092 | 9239 | 0.024 | 0.1443 | No | ||
| 94 | CSF3 | 2230193 6660707 | 9341 | 0.022 | 0.1390 | No | ||
| 95 | PTPRJ | 4010707 | 9555 | 0.019 | 0.1276 | No | ||
| 96 | DUSP22 | 4280010 5550546 | 10209 | 0.008 | 0.0923 | No | ||
| 97 | PPP1R13B | 4010605 4760181 | 10254 | 0.007 | 0.0900 | No | ||
| 98 | MADCAM1 | 1580010 | 10260 | 0.007 | 0.0897 | No | ||
| 99 | IL7 | 5360440 | 10545 | 0.002 | 0.0744 | No | ||
| 100 | IL1RN | 2370333 | 10595 | 0.001 | 0.0717 | No | ||
| 101 | NR2F2 | 3170609 3310577 | 10615 | 0.001 | 0.0707 | No | ||
| 102 | FGF17 | 3130022 | 10652 | 0.000 | 0.0687 | No | ||
| 103 | KLK9 | 4120164 | 10721 | -0.001 | 0.0651 | No | ||
| 104 | GRIN2D | 6620372 | 10950 | -0.005 | 0.0527 | No | ||
| 105 | TNFSF15 | 360341 | 10993 | -0.005 | 0.0505 | No | ||
| 106 | GNG4 | 2630600 | 11135 | -0.008 | 0.0429 | No | ||
| 107 | ZIC4 | 1500082 | 11366 | -0.012 | 0.0306 | No | ||
| 108 | HTR3B | 430438 520301 | 11512 | -0.015 | 0.0228 | No | ||
| 109 | EDN2 | 6760647 | 11703 | -0.018 | 0.0127 | No | ||
| 110 | AP1S2 | 780279 4280707 | 11872 | -0.022 | 0.0037 | No | ||
| 111 | IER5 | 2350575 | 11920 | -0.023 | 0.0013 | No | ||
| 112 | MITF | 380056 | 12093 | -0.026 | -0.0078 | No | ||
| 113 | ETV1 | 70735 2940603 5080463 | 12133 | -0.027 | -0.0097 | No | ||
| 114 | TRIM47 | 5080717 | 12388 | -0.033 | -0.0232 | No | ||
| 115 | UBE2I | 2680056 6350446 | 12636 | -0.040 | -0.0363 | No | ||
| 116 | UBD | 5570632 | 12731 | -0.043 | -0.0411 | No | ||
| 117 | HSD11B2 | 5900053 | 12830 | -0.045 | -0.0460 | No | ||
| 118 | SLAMF8 | 1230048 | 12932 | -0.049 | -0.0511 | No | ||
| 119 | ASCL3 | 4570484 | 13052 | -0.053 | -0.0572 | No | ||
| 120 | ZNRF1 | 70592 | 13103 | -0.054 | -0.0595 | No | ||
| 121 | SNAP25 | 360520 | 13606 | -0.076 | -0.0861 | No | ||
| 122 | SHOX2 | 3190438 6450059 | 14135 | -0.115 | -0.1139 | No | ||
| 123 | BIRC3 | 6550411 | 14201 | -0.122 | -0.1165 | No | ||
| 124 | SLC6A12 | 3170685 | 14224 | -0.124 | -0.1167 | No | ||
| 125 | TNFSF18 | 1990458 2850520 | 14469 | -0.154 | -0.1288 | No | ||
| 126 | DUSP6 | 5910286 7100070 | 14573 | -0.170 | -0.1331 | No | ||
| 127 | WNT10A | 2100706 | 14686 | -0.191 | -0.1377 | No | ||
| 128 | PFN1 | 6130132 | 14735 | -0.201 | -0.1388 | No | ||
| 129 | COL27A1 | 1740390 | 14752 | -0.205 | -0.1381 | No | ||
| 130 | RIN2 | 2450184 | 14893 | -0.234 | -0.1439 | No | ||
| 131 | GADD45B | 2350408 | 15040 | -0.272 | -0.1498 | No | ||
| 132 | NLK | 2030010 2450041 | 15117 | -0.295 | -0.1517 | No | ||
| 133 | CREB1 | 1500717 2230358 3610600 6550601 | 15202 | -0.323 | -0.1538 | No | ||
| 134 | SP6 | 60484 510452 2690333 | 15341 | -0.366 | -0.1585 | No | ||
| 135 | AMOTL1 | 380279 | 15349 | -0.370 | -0.1561 | No | ||
| 136 | IL12A | 7100551 | 15410 | -0.389 | -0.1565 | No | ||
| 137 | CD69 | 380167 4730088 | 15476 | -0.411 | -0.1569 | No | ||
| 138 | GATA4 | 1410215 2690609 | 15615 | -0.463 | -0.1609 | No | ||
| 139 | EGF | 5220154 | 15653 | -0.474 | -0.1593 | No | ||
| 140 | BMF | 610102 | 15810 | -0.537 | -0.1637 | No | ||
| 141 | NFAT5 | 2510411 5890195 6550152 | 15815 | -0.540 | -0.1598 | No | ||
| 142 | NFKBIA | 1570152 | 16124 | -0.682 | -0.1714 | No | ||
| 143 | PCBP4 | 6350722 | 16142 | -0.693 | -0.1671 | No | ||
| 144 | NR3C1 | 630563 | 16337 | -0.804 | -0.1715 | No | ||
| 145 | ELMO1 | 110338 1190451 1580148 1850609 2350706 2370369 2450129 3840113 4150184 4280079 6290066 6380435 6770494 7100180 | 16641 | -0.978 | -0.1806 | No | ||
| 146 | UBE2H | 1980142 2970079 | 16734 | -1.024 | -0.1779 | No | ||
| 147 | REV3L | 1090717 | 16793 | -1.065 | -0.1730 | No | ||
| 148 | HSD3B7 | 1410129 | 17126 | -1.356 | -0.1807 | No | ||
| 149 | UACA | 1190725 | 17146 | -1.374 | -0.1714 | No | ||
| 150 | UBE2D3 | 3190452 | 17364 | -1.609 | -0.1710 | No | ||
| 151 | CCL5 | 3710397 | 17429 | -1.684 | -0.1618 | No | ||
| 152 | SMOC1 | 840039 | 17484 | -1.741 | -0.1516 | No | ||
| 153 | SOCS2 | 4760692 | 17492 | -1.751 | -0.1388 | No | ||
| 154 | UPF2 | 4810471 | 17631 | -1.906 | -0.1319 | No | ||
| 155 | PRKCD | 770592 | 17650 | -1.931 | -0.1183 | No | ||
| 156 | RBMS1 | 6400014 | 17662 | -1.950 | -0.1042 | No | ||
| 157 | VCAM1 | 2900450 | 17830 | -2.190 | -0.0968 | No | ||
| 158 | IL6ST | 5890047 | 17857 | -2.230 | -0.0814 | No | ||
| 159 | GNB1 | 2120397 | 17866 | -2.237 | -0.0649 | No | ||
| 160 | ATP1B1 | 3130594 | 18138 | -2.762 | -0.0588 | No | ||
| 161 | RAP2C | 1690132 | 18365 | -3.418 | -0.0453 | No | ||
| 162 | CYLD | 6590079 | 18393 | -3.506 | -0.0203 | No | ||
| 163 | SIN3A | 2190121 | 18531 | -4.292 | 0.0046 | No |