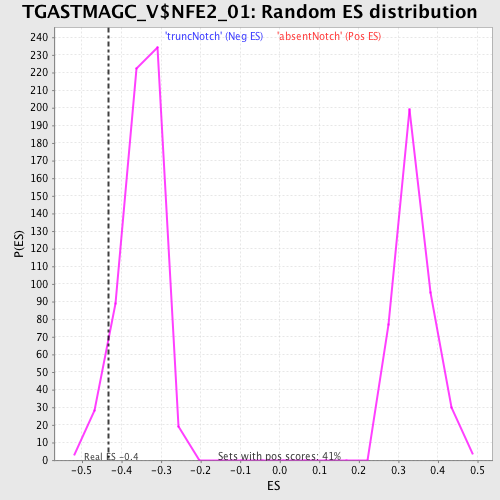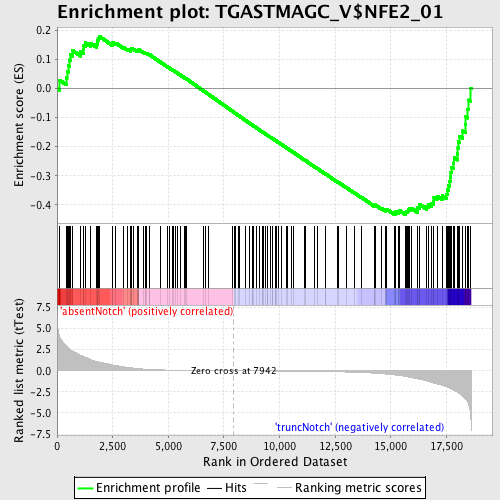
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #absentNotch_versus_truncNotch.phenotype_absentNotch_versus_truncNotch.cls #absentNotch_versus_truncNotch_repos |
| Phenotype | phenotype_absentNotch_versus_truncNotch.cls#absentNotch_versus_truncNotch_repos |
| Upregulated in class | truncNotch |
| GeneSet | TGASTMAGC_V$NFE2_01 |
| Enrichment Score (ES) | -0.43281916 |
| Normalized Enrichment Score (NES) | -1.2325232 |
| Nominal p-value | 0.06890757 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PPP2CA | 3990113 | 115 | 4.067 | 0.0280 | No | ||
| 2 | SKP1A | 2450102 | 407 | 2.919 | 0.0369 | No | ||
| 3 | TTC1 | 5890736 | 445 | 2.823 | 0.0587 | No | ||
| 4 | EIF4G1 | 4070446 | 496 | 2.717 | 0.0789 | No | ||
| 5 | XPOT | 7050184 | 553 | 2.580 | 0.0976 | No | ||
| 6 | NR1D1 | 2360471 770746 6590204 | 602 | 2.480 | 0.1159 | No | ||
| 7 | PSMD7 | 2030619 6220594 | 697 | 2.328 | 0.1304 | No | ||
| 8 | NEFH | 630239 | 1061 | 1.821 | 0.1261 | No | ||
| 9 | SNX10 | 3800035 | 1170 | 1.695 | 0.1345 | No | ||
| 10 | PSMD12 | 730044 | 1178 | 1.690 | 0.1484 | No | ||
| 11 | MDM2 | 3450053 5080138 | 1273 | 1.581 | 0.1566 | No | ||
| 12 | RBBP7 | 430113 450450 2370309 | 1507 | 1.334 | 0.1552 | No | ||
| 13 | GADD45G | 2510142 | 1766 | 1.097 | 0.1505 | No | ||
| 14 | NRD1 | 5900017 | 1797 | 1.077 | 0.1579 | No | ||
| 15 | BTK | 3130044 | 1814 | 1.068 | 0.1661 | No | ||
| 16 | SLC16A6 | 1690156 | 1865 | 1.035 | 0.1721 | No | ||
| 17 | ANGPTL4 | 2480195 6760593 | 1890 | 1.017 | 0.1794 | No | ||
| 18 | CMAS | 2190129 | 2480 | 0.688 | 0.1533 | No | ||
| 19 | MAP2K1 | 840739 | 2487 | 0.683 | 0.1587 | No | ||
| 20 | DLX1 | 360168 | 2642 | 0.614 | 0.1556 | No | ||
| 21 | SEC24D | 3060279 | 3003 | 0.462 | 0.1400 | No | ||
| 22 | MCART1 | 5270059 | 3180 | 0.397 | 0.1338 | No | ||
| 23 | LRRFIP2 | 670575 730152 6590438 | 3289 | 0.359 | 0.1310 | No | ||
| 24 | E2F3 | 50162 460180 | 3320 | 0.350 | 0.1323 | No | ||
| 25 | PDE4D | 2470528 6660014 | 3326 | 0.349 | 0.1350 | No | ||
| 26 | VDR | 130156 510438 | 3329 | 0.348 | 0.1378 | No | ||
| 27 | PEA15 | 4590685 | 3440 | 0.313 | 0.1345 | No | ||
| 28 | EIF4E | 1580403 70133 6380215 | 3604 | 0.264 | 0.1279 | No | ||
| 29 | APBA2BP | 5700341 | 3619 | 0.259 | 0.1293 | No | ||
| 30 | BACH1 | 290195 | 3662 | 0.249 | 0.1291 | No | ||
| 31 | SLC22A18 | 7050168 | 3663 | 0.249 | 0.1312 | No | ||
| 32 | RABEP1 | 4760427 6200463 | 3673 | 0.247 | 0.1328 | No | ||
| 33 | PFN1 | 6130132 | 3881 | 0.201 | 0.1233 | No | ||
| 34 | LAMA3 | 1190131 5290239 | 3969 | 0.184 | 0.1201 | No | ||
| 35 | ADAM15 | 5890707 | 3980 | 0.182 | 0.1211 | No | ||
| 36 | DTNA | 1340600 1780731 2340278 2850132 | 4019 | 0.174 | 0.1205 | No | ||
| 37 | IDS | 360619 5720161 | 4151 | 0.153 | 0.1147 | No | ||
| 38 | MYO10 | 3830576 5570092 | 4154 | 0.153 | 0.1159 | No | ||
| 39 | AKT3 | 1580270 3290278 | 4634 | 0.102 | 0.0908 | No | ||
| 40 | IL6 | 380133 | 4953 | 0.080 | 0.0743 | No | ||
| 41 | SLC26A1 | 2190142 | 5043 | 0.075 | 0.0701 | No | ||
| 42 | CALB2 | 6130170 6590368 | 5176 | 0.068 | 0.0635 | No | ||
| 43 | SERPINB5 | 6940050 | 5210 | 0.066 | 0.0623 | No | ||
| 44 | SH3BGRL2 | 1780239 | 5313 | 0.062 | 0.0573 | No | ||
| 45 | SNCB | 3360133 | 5427 | 0.057 | 0.0517 | No | ||
| 46 | XLKD1 | 520441 | 5562 | 0.053 | 0.0449 | No | ||
| 47 | PTPRN | 5900577 | 5725 | 0.048 | 0.0365 | No | ||
| 48 | NUDT11 | 3840239 | 5770 | 0.046 | 0.0345 | No | ||
| 49 | PLCD1 | 2030484 | 5822 | 0.044 | 0.0321 | No | ||
| 50 | SLITRK6 | 7320739 | 6601 | 0.025 | -0.0098 | No | ||
| 51 | OMG | 6760066 | 6670 | 0.023 | -0.0133 | No | ||
| 52 | USP13 | 1450670 | 6819 | 0.020 | -0.0211 | No | ||
| 53 | PROCR | 4920687 | 7874 | 0.001 | -0.0782 | No | ||
| 54 | MAPRE3 | 7050504 | 7967 | -0.000 | -0.0831 | No | ||
| 55 | PHLDA2 | 4810494 | 8009 | -0.001 | -0.0853 | No | ||
| 56 | CRYBA2 | 5900138 | 8164 | -0.003 | -0.0936 | No | ||
| 57 | BNIP3 | 3140270 | 8199 | -0.004 | -0.0955 | No | ||
| 58 | PLS3 | 3290082 | 8484 | -0.009 | -0.1107 | No | ||
| 59 | SMPX | 6590440 | 8658 | -0.012 | -0.1200 | No | ||
| 60 | SYT2 | 60609 | 8773 | -0.014 | -0.1261 | No | ||
| 61 | LENEP | 4850369 | 8802 | -0.014 | -0.1275 | No | ||
| 62 | IL18RAP | 3850088 | 8826 | -0.015 | -0.1286 | No | ||
| 63 | TNXB | 630592 3830020 5360497 7000673 | 8948 | -0.017 | -0.1350 | No | ||
| 64 | PSMA3 | 5900047 7040161 | 9078 | -0.019 | -0.1418 | No | ||
| 65 | SFTPC | 3290133 | 9209 | -0.021 | -0.1487 | No | ||
| 66 | CPNE8 | 7040039 | 9242 | -0.022 | -0.1502 | No | ||
| 67 | MYBPH | 2190711 | 9266 | -0.022 | -0.1513 | No | ||
| 68 | TLX2 | 1090128 2900451 | 9352 | -0.024 | -0.1557 | No | ||
| 69 | PSMD11 | 2340538 6510053 | 9447 | -0.025 | -0.1606 | No | ||
| 70 | P2RXL1 | 4730390 5220446 | 9477 | -0.026 | -0.1619 | No | ||
| 71 | CLDN15 | 4150270 4730592 | 9605 | -0.029 | -0.1685 | No | ||
| 72 | PRDM1 | 3170347 3520301 | 9666 | -0.030 | -0.1715 | No | ||
| 73 | CRYGS | 2680398 | 9691 | -0.030 | -0.1726 | No | ||
| 74 | LRRC2 | 3130400 | 9808 | -0.033 | -0.1786 | No | ||
| 75 | EYA1 | 1450278 5220390 | 9835 | -0.033 | -0.1797 | No | ||
| 76 | WDFY3 | 3610041 4560333 4570273 | 9869 | -0.034 | -0.1812 | No | ||
| 77 | TIAL1 | 4150048 6510605 | 9953 | -0.036 | -0.1854 | No | ||
| 78 | SPATS2 | 2360066 | 10071 | -0.039 | -0.1914 | No | ||
| 79 | FOSL1 | 430021 | 10317 | -0.043 | -0.2043 | No | ||
| 80 | MT3 | 1450537 | 10349 | -0.044 | -0.2056 | No | ||
| 81 | SNCG | 3120725 | 10546 | -0.049 | -0.2158 | No | ||
| 82 | FGF11 | 840292 | 10619 | -0.050 | -0.2193 | No | ||
| 83 | MPP5 | 2100148 5220632 | 11117 | -0.062 | -0.2457 | No | ||
| 84 | DHRS3 | 360609 | 11160 | -0.063 | -0.2474 | No | ||
| 85 | FGF9 | 1050195 | 11581 | -0.077 | -0.2695 | No | ||
| 86 | TLL1 | 2680315 | 11696 | -0.081 | -0.2750 | No | ||
| 87 | LRP1B | 1450193 | 12044 | -0.094 | -0.2930 | No | ||
| 88 | MMP9 | 580338 | 12075 | -0.096 | -0.2938 | No | ||
| 89 | ELA1 | 3390167 | 12593 | -0.122 | -0.3207 | No | ||
| 90 | PVALB | 5570450 | 12646 | -0.126 | -0.3225 | No | ||
| 91 | DIRAS1 | 2640270 | 12992 | -0.150 | -0.3399 | No | ||
| 92 | FLT1 | 3830167 4920438 | 13370 | -0.179 | -0.3588 | No | ||
| 93 | RICS | 1940673 | 13677 | -0.213 | -0.3736 | No | ||
| 94 | PLEC1 | 6520292 50500 380075 1570088 2510692 2630497 2680017 2900746 4560497 4850376 6290398 7040471 | 14274 | -0.293 | -0.4034 | No | ||
| 95 | CD44 | 3990072 4850671 5860411 6860148 7050551 | 14275 | -0.293 | -0.4009 | No | ||
| 96 | RB1CC1 | 510494 7100072 | 14304 | -0.298 | -0.3999 | No | ||
| 97 | GADD45A | 2900717 | 14582 | -0.346 | -0.4120 | No | ||
| 98 | RAB13 | 3710309 | 14754 | -0.381 | -0.4180 | No | ||
| 99 | MMRN2 | 1740047 | 14773 | -0.386 | -0.4157 | No | ||
| 100 | TUBB4 | 5310681 | 14819 | -0.395 | -0.4148 | No | ||
| 101 | BLMH | 1410438 | 15152 | -0.483 | -0.4287 | Yes | ||
| 102 | ADRA1A | 1230446 2260390 | 15217 | -0.507 | -0.4279 | Yes | ||
| 103 | DUSP13 | 6400152 | 15219 | -0.508 | -0.4237 | Yes | ||
| 104 | MDFI | 6200041 | 15343 | -0.546 | -0.4258 | Yes | ||
| 105 | FBXO2 | 6510091 | 15347 | -0.549 | -0.4213 | Yes | ||
| 106 | CDH23 | 4250242 5290397 6550397 | 15403 | -0.570 | -0.4195 | Yes | ||
| 107 | GSTP1 | 3170102 3710707 5080368 | 15649 | -0.665 | -0.4272 | Yes | ||
| 108 | GABARAPL1 | 2810458 | 15682 | -0.678 | -0.4232 | Yes | ||
| 109 | PSMD2 | 4670706 5050364 | 15757 | -0.711 | -0.4212 | Yes | ||
| 110 | MMP19 | 5080377 | 15774 | -0.722 | -0.4160 | Yes | ||
| 111 | TMEM24 | 5860039 | 15819 | -0.746 | -0.4121 | Yes | ||
| 112 | ARHGAP8 | 380333 5390487 | 15922 | -0.803 | -0.4108 | Yes | ||
| 113 | DTX2 | 4210041 7000008 | 16190 | -0.950 | -0.4173 | Yes | ||
| 114 | BAZ2A | 730184 | 16198 | -0.958 | -0.4096 | Yes | ||
| 115 | MFN2 | 2260195 6100164 | 16277 | -1.001 | -0.4054 | Yes | ||
| 116 | PADI4 | 1740075 4810441 | 16280 | -1.002 | -0.3970 | Yes | ||
| 117 | RIT1 | 5390338 | 16617 | -1.206 | -0.4051 | Yes | ||
| 118 | SYTL1 | 510487 | 16683 | -1.252 | -0.3980 | Yes | ||
| 119 | VASP | 7050500 | 16831 | -1.374 | -0.3944 | Yes | ||
| 120 | ABHD4 | 6760450 | 16929 | -1.462 | -0.3874 | Yes | ||
| 121 | CLIC1 | 110600 | 16937 | -1.464 | -0.3754 | Yes | ||
| 122 | BAG2 | 2900324 | 17077 | -1.540 | -0.3699 | Yes | ||
| 123 | VAT1 | 1050040 | 17308 | -1.732 | -0.3678 | Yes | ||
| 124 | NCDN | 50040 1980739 3520603 | 17519 | -1.916 | -0.3630 | Yes | ||
| 125 | FBXO44 | 4760372 | 17539 | -1.939 | -0.3477 | Yes | ||
| 126 | ADCK4 | 1660180 | 17583 | -1.985 | -0.3333 | Yes | ||
| 127 | ROM1 | 3130685 | 17638 | -2.044 | -0.3190 | Yes | ||
| 128 | ABCF3 | 7100100 | 17676 | -2.081 | -0.3035 | Yes | ||
| 129 | RAB3D | 4210253 | 17701 | -2.109 | -0.2870 | Yes | ||
| 130 | SQSTM1 | 6550056 | 17732 | -2.162 | -0.2704 | Yes | ||
| 131 | CAMKK1 | 6370324 | 17832 | -2.283 | -0.2565 | Yes | ||
| 132 | HDAC3 | 4060072 | 17839 | -2.296 | -0.2375 | Yes | ||
| 133 | ENO1 | 5340128 | 17995 | -2.530 | -0.2246 | Yes | ||
| 134 | PPP1R9B | 3130619 | 18017 | -2.558 | -0.2042 | Yes | ||
| 135 | ETV5 | 110017 | 18034 | -2.603 | -0.1831 | Yes | ||
| 136 | ITPKC | 5420021 | 18099 | -2.678 | -0.1640 | Yes | ||
| 137 | ABCA2 | 5050039 | 18231 | -2.965 | -0.1461 | Yes | ||
| 138 | ENO3 | 5270136 | 18352 | -3.305 | -0.1248 | Yes | ||
| 139 | ABCD1 | 5670373 | 18368 | -3.366 | -0.0972 | Yes | ||
| 140 | LIMK1 | 5080064 | 18452 | -3.660 | -0.0709 | Yes | ||
| 141 | ABCB6 | 6350138 | 18481 | -3.862 | -0.0399 | Yes | ||
| 142 | ALDOA | 6290672 | 18596 | -5.593 | 0.0011 | Yes |