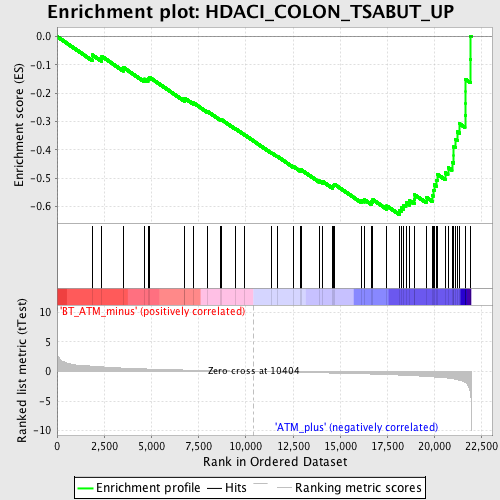
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | HDACI_COLON_TSABUT_UP |
| Enrichment Score (ES) | -0.627621 |
| Normalized Enrichment Score (NES) | -2.2110608 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0010798818 |
| FWER p-Value | 0.0050 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | SCYL3 | 1428918_at 1446655_at 1452960_at | 1863 | 0.887 | -0.0647 | No | ||
| 2 | AQP1 | 1416203_at | 2369 | 0.770 | -0.0700 | No | ||
| 3 | ROCK1 | 1423444_at 1423445_at 1441162_at 1446518_at 1450994_at 1460729_at | 3503 | 0.563 | -0.1088 | No | ||
| 4 | TMTC3 | 1436826_at 1457294_at | 4616 | 0.428 | -0.1497 | No | ||
| 5 | TSNAX | 1423395_at 1430045_at | 4815 | 0.409 | -0.1493 | No | ||
| 6 | BCKDK | 1443813_x_at 1460644_at | 4883 | 0.402 | -0.1431 | No | ||
| 7 | CPEB4 | 1420617_at 1420618_at 1449931_at | 6742 | 0.243 | -0.2224 | No | ||
| 8 | VEGF | 1420909_at 1451959_a_at | 6760 | 0.242 | -0.2176 | No | ||
| 9 | BIRC4 | 1421394_a_at 1426636_a_at 1437533_at 1440351_at 1450231_a_at 1450232_at 1456088_at | 7223 | 0.207 | -0.2340 | No | ||
| 10 | ZNF76 | 1434085_at 1442446_at | 7956 | 0.157 | -0.2638 | No | ||
| 11 | CRYL1 | 1416795_at 1430681_at 1447112_s_at 1459589_at | 8659 | 0.110 | -0.2934 | No | ||
| 12 | TMEM63C | 1456184_at | 8722 | 0.106 | -0.2938 | No | ||
| 13 | GRAMD1C | 1434725_at | 9457 | 0.061 | -0.3259 | No | ||
| 14 | MAP1LC3B | 1415928_a_at 1415929_at 1415930_a_at 1428024_at | 9924 | 0.031 | -0.3465 | No | ||
| 15 | SF3A2 | 1450576_a_at 1455546_s_at | 11374 | -0.061 | -0.4113 | No | ||
| 16 | HSPA1A | 1452388_at | 11655 | -0.078 | -0.4223 | No | ||
| 17 | NEUROD6 | 1418047_at | 12533 | -0.132 | -0.4594 | No | ||
| 18 | GPX3 | 1449106_at | 12875 | -0.154 | -0.4714 | No | ||
| 19 | CYB5B | 1417766_at 1417767_at 1443202_at 1448844_at | 12928 | -0.157 | -0.4701 | No | ||
| 20 | HSF1 | 1424622_at | 13898 | -0.221 | -0.5093 | No | ||
| 21 | FOSB | 1422134_at | 14066 | -0.232 | -0.5116 | No | ||
| 22 | NEU1 | 1416831_at 1447942_x_at 1448481_at | 14577 | -0.270 | -0.5287 | No | ||
| 23 | LRP8 | 1421459_a_at 1440882_at 1442347_at | 14618 | -0.273 | -0.5242 | No | ||
| 24 | SLC27A1 | 1422811_at 1435658_at | 14674 | -0.277 | -0.5204 | No | ||
| 25 | PLCB4 | 1425338_at 1425339_at 1430871_at 1435771_at 1441531_at | 16131 | -0.387 | -0.5780 | No | ||
| 26 | ALG12 | 1424818_at 1427557_at 1427558_s_at | 16257 | -0.398 | -0.5745 | No | ||
| 27 | GNAI2 | 1419449_a_at 1435652_a_at | 16638 | -0.433 | -0.5819 | No | ||
| 28 | SERPINE1 | 1419149_at | 16723 | -0.439 | -0.5756 | No | ||
| 29 | RHOB | 1449110_at | 17466 | -0.518 | -0.5975 | No | ||
| 30 | INPP5E | 1423229_at 1423230_at | 18125 | -0.601 | -0.6138 | Yes | ||
| 31 | ECHDC2 | 1425788_a_at | 18227 | -0.613 | -0.6042 | Yes | ||
| 32 | PIP5K2C | 1416387_at 1416388_at | 18345 | -0.630 | -0.5950 | Yes | ||
| 33 | PER1 | 1449851_at | 18478 | -0.648 | -0.5861 | Yes | ||
| 34 | RHBDF1 | 1424138_at | 18662 | -0.677 | -0.5789 | Yes | ||
| 35 | SEPT3 | 1417414_at 1439833_at 1444778_at | 18912 | -0.717 | -0.5737 | Yes | ||
| 36 | AVPR1A | 1418603_at 1418604_at | 18931 | -0.718 | -0.5580 | Yes | ||
| 37 | FKBP5 | 1416125_at 1447736_at 1448231_at 1458089_at | 19553 | -0.831 | -0.5672 | Yes | ||
| 38 | SAMD4A | 1429449_at 1454382_at | 19890 | -0.905 | -0.5616 | Yes | ||
| 39 | WIPI1 | 1424917_a_at 1427323_s_at 1438709_at 1439060_s_at 1452392_a_at 1456624_at | 19915 | -0.912 | -0.5417 | Yes | ||
| 40 | IL10RA | 1437808_x_at 1448731_at 1456173_at | 19971 | -0.926 | -0.5228 | Yes | ||
| 41 | CALCOCO1 | 1428513_at | 20092 | -0.956 | -0.5063 | Yes | ||
| 42 | PLAG1 | 1421745_at 1443943_at | 20147 | -0.968 | -0.4864 | Yes | ||
| 43 | FLNB | 1424828_a_at 1426750_at 1442107_at 1445534_at 1458226_at | 20553 | -1.065 | -0.4804 | Yes | ||
| 44 | ZUBR1 | 1454668_at | 20702 | -1.111 | -0.4615 | Yes | ||
| 45 | KCNA1 | 1417416_at 1437230_at 1442413_at 1455785_at | 20914 | -1.213 | -0.4431 | Yes | ||
| 46 | CREBBP | 1434633_at 1435224_at 1436983_at 1444856_at 1457641_at 1459804_at | 21006 | -1.257 | -0.4183 | Yes | ||
| 47 | GLI3 | 1444043_at 1450525_at 1455154_at 1456067_at | 21008 | -1.259 | -0.3893 | Yes | ||
| 48 | GLRX | 1416592_at 1416593_at | 21087 | -1.304 | -0.3628 | Yes | ||
| 49 | HTF9C | 1434244_x_at 1441920_x_at 1448114_a_at 1448115_at 1453995_a_at 1457620_at | 21203 | -1.375 | -0.3363 | Yes | ||
| 50 | ETV5 | 1420998_at 1428142_at 1450082_s_at | 21313 | -1.468 | -0.3074 | Yes | ||
| 51 | JARID1C | 1426497_at 1426498_at 1441449_at 1441450_s_at 1444157_a_at 1444158_at 1457930_at | 21602 | -1.819 | -0.2786 | Yes | ||
| 52 | TPM3 | 1427260_a_at 1427567_a_at 1436958_x_at 1449996_a_at 1449997_at | 21616 | -1.853 | -0.2364 | Yes | ||
| 53 | SAT1 | 1420502_at | 21621 | -1.860 | -0.1937 | Yes | ||
| 54 | SPARCL1 | 1416114_at | 21623 | -1.868 | -0.1506 | Yes | ||
| 55 | ID2 | 1422537_a_at 1435176_a_at 1453596_at | 21890 | -3.497 | -0.0821 | Yes | ||
| 56 | FOS | 1423100_at | 21901 | -3.644 | 0.0015 | Yes |

